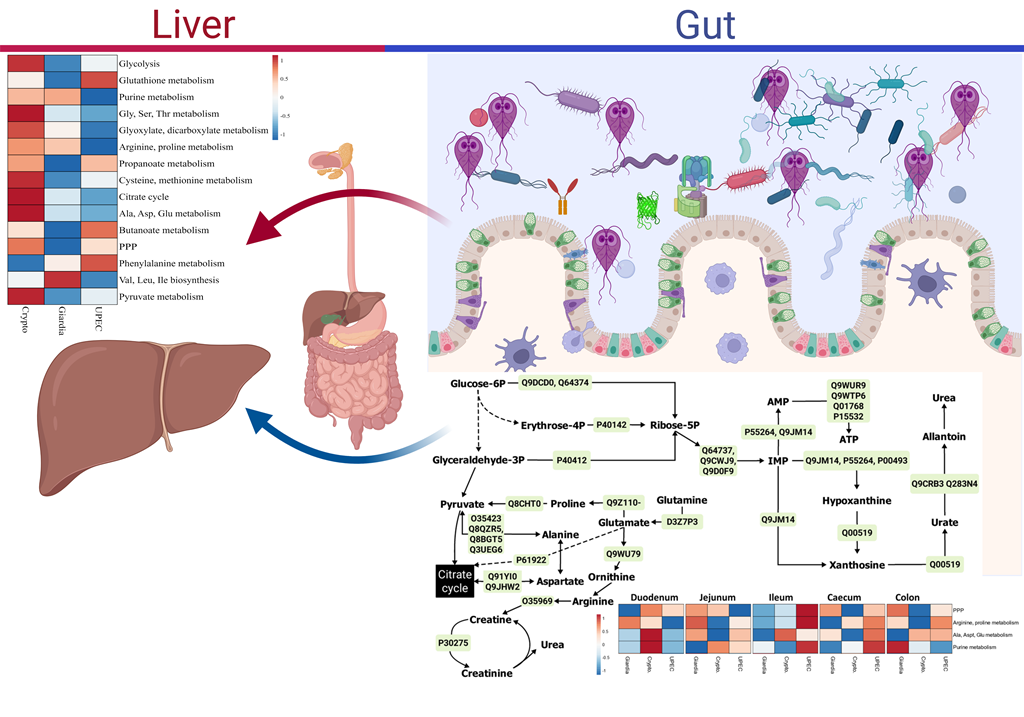
Apicomplexan infections, such as giardiasis and cryptosporidiosis, negatively impact a considerable proportion of human and commercial livestock populations. Despite this, the molecular mecha-nisms of disease, particularly the effect on the body beyond the gastrointestinal tract, are still poorly understood. To highlight host-parasite-microbiome biochemical interactions, we utilised integrated metabolomics-16S rRNA genomics and metabolomics-proteomics approaches in a C57BL/6J mouse model of giardiasis and compared these to Cryptosporidium and uropathogenic Escherichia coli (UPEC) infections. Comprehensive samples (faeces, blood, liver, and luminal contents from duo-denum, jejunum, ileum, caecum and colon) were collected 10 days post infection and subjected to proteome and metabolome analysis by liquid and gas chromatographic mass-spectrometry, re-spectively. Microbial populations in faeces and luminal washes were examined using 16S rRNA metagenomics. Proteome-metabolome analyses indicated that 12 and 16 key pathways were sig-nificantly altered in the gut and liver, respectively, during giardiasis with respect to other infections. Energy pathways including glycolysis and supporting pathways of glyoxylate and dicarboxylate metabolism, and redox pathway of glutathione metabolism, were upregulated in small intestinal luminal contents and the liver during giardiasis. Metabolomics-16S rRNA genetics integration in-dicated that populations of three bacterial families –Autopobiaceae (Up), Desulfovibrionaceae (Up) and Akkermanasiaceae (Down) – were most significantly affected across the gut during giardiasis, causing upregulated glycolysis and short-chained fatty acid (SCFA) metabolism. In particular, the perturbed Akkermanasiaceae population seemed to cause oxidative stress responses along the gut-liver axis. Overall, the systems biology approach applied in this study highlighted that the effects of host-parasite-microbiome biochemical interactions extended beyond the gut ecosystem to the gut-liver axis. These findings form the first steps in a comprehensive comparison to ascertain the major molecular and biochemical contributors of host-parasite interactions and contribute towards the development of biomarker discovery and precision health solutions for apicomplexan infections.