Submitted:
14 November 2023
Posted:
15 November 2023
You are already at the latest version
Abstract
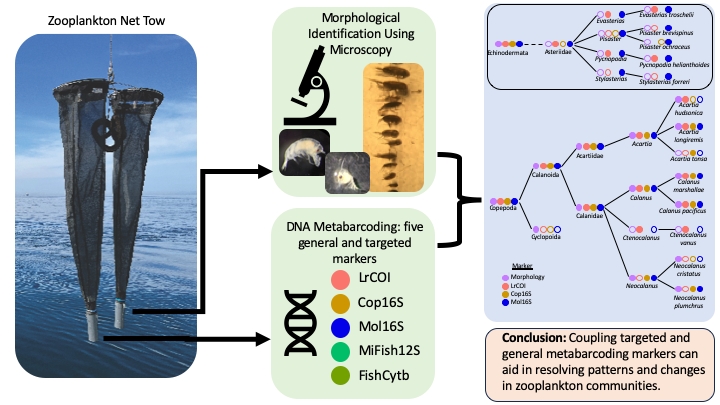
Keywords:
1. Introduction
1.1. Zooplankton community and approach
1.2. Metabarcoding markers evaluated for the zooplankton and ichthyoplankton communities
1.3. Project aim and scientific question
2. Materials and Methods
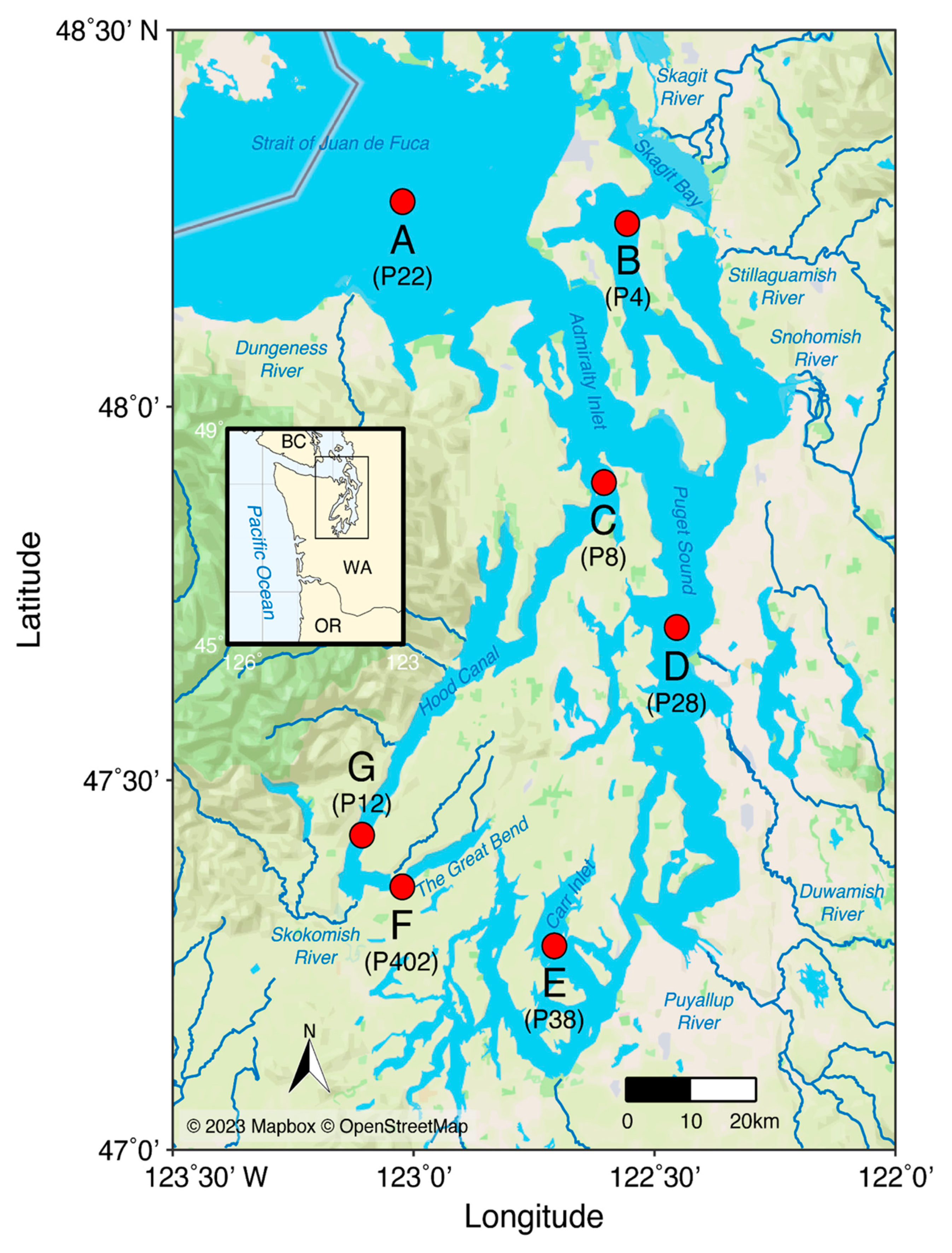
2.1. Sampling, metadata, and morphological identifications
2.2. DNA extraction, library preparation, and metabarcoding
2.3. Bioinformatic processing
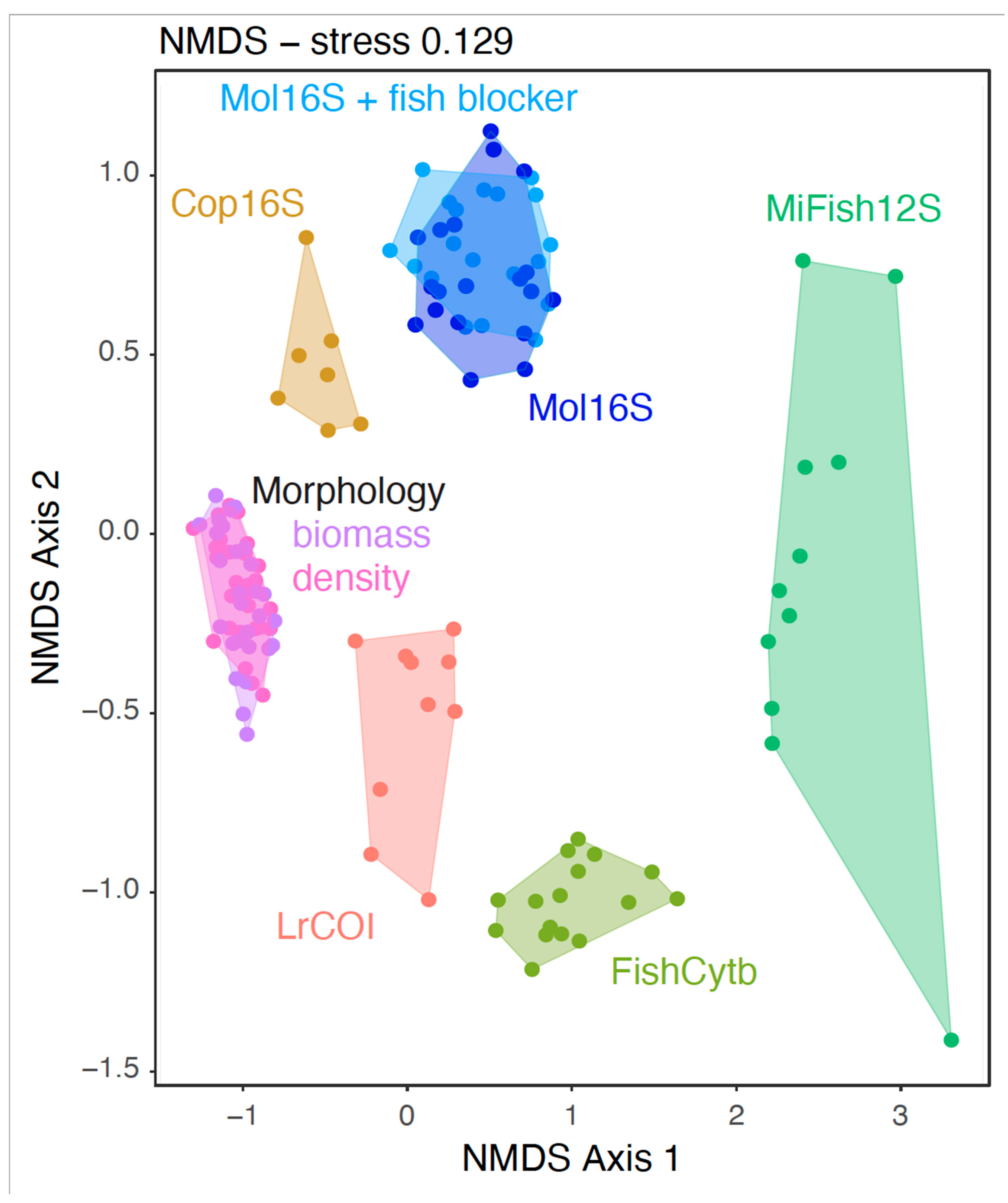
2.4. Statistical analyses
3. Results
3.1. Marker strengths and limitations
3.1.1. LrCOI
3.1.2. Cop16S
3.1.3. Mol16S
3.1.4. MiFish12S
3.1.5. FishCytb
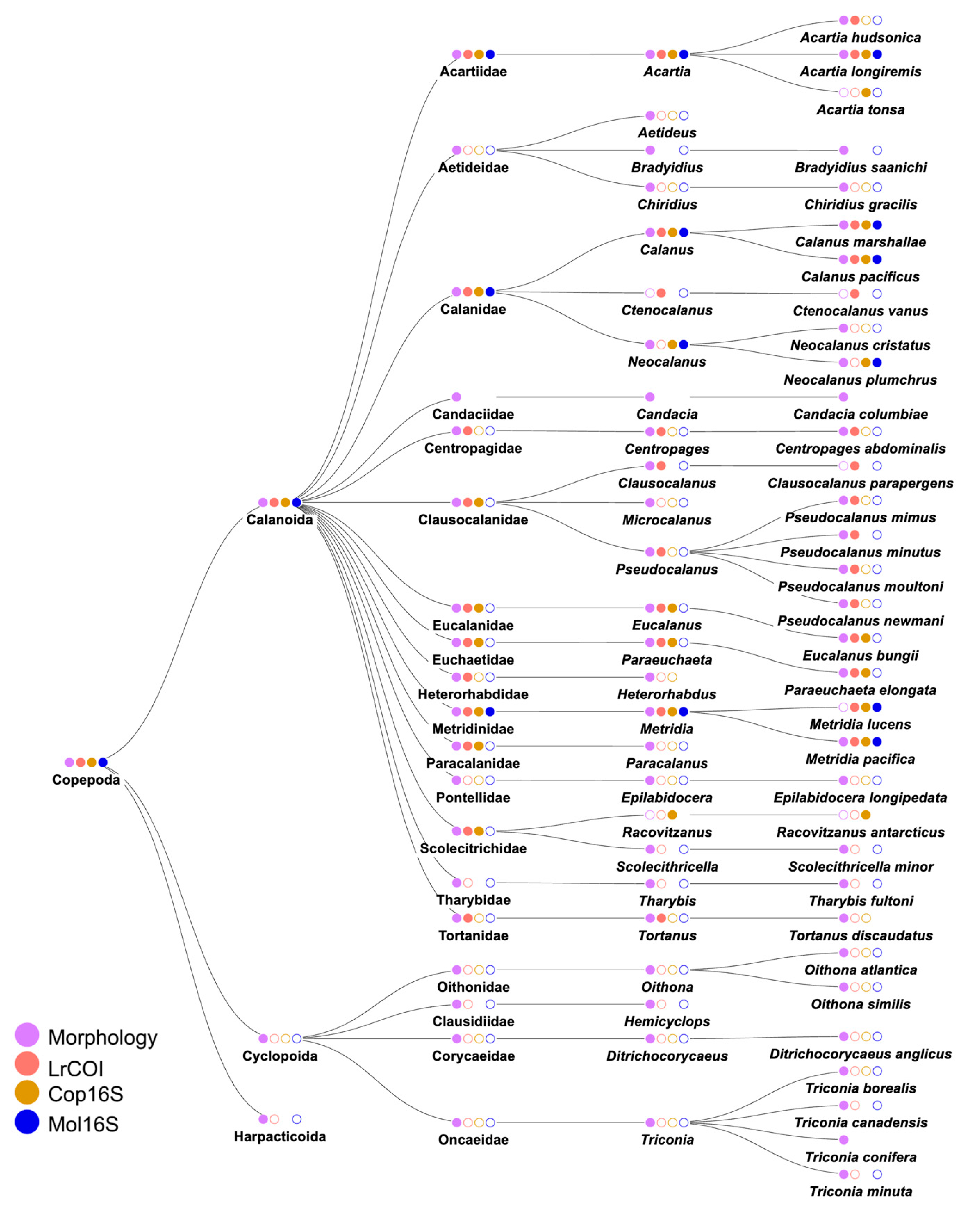
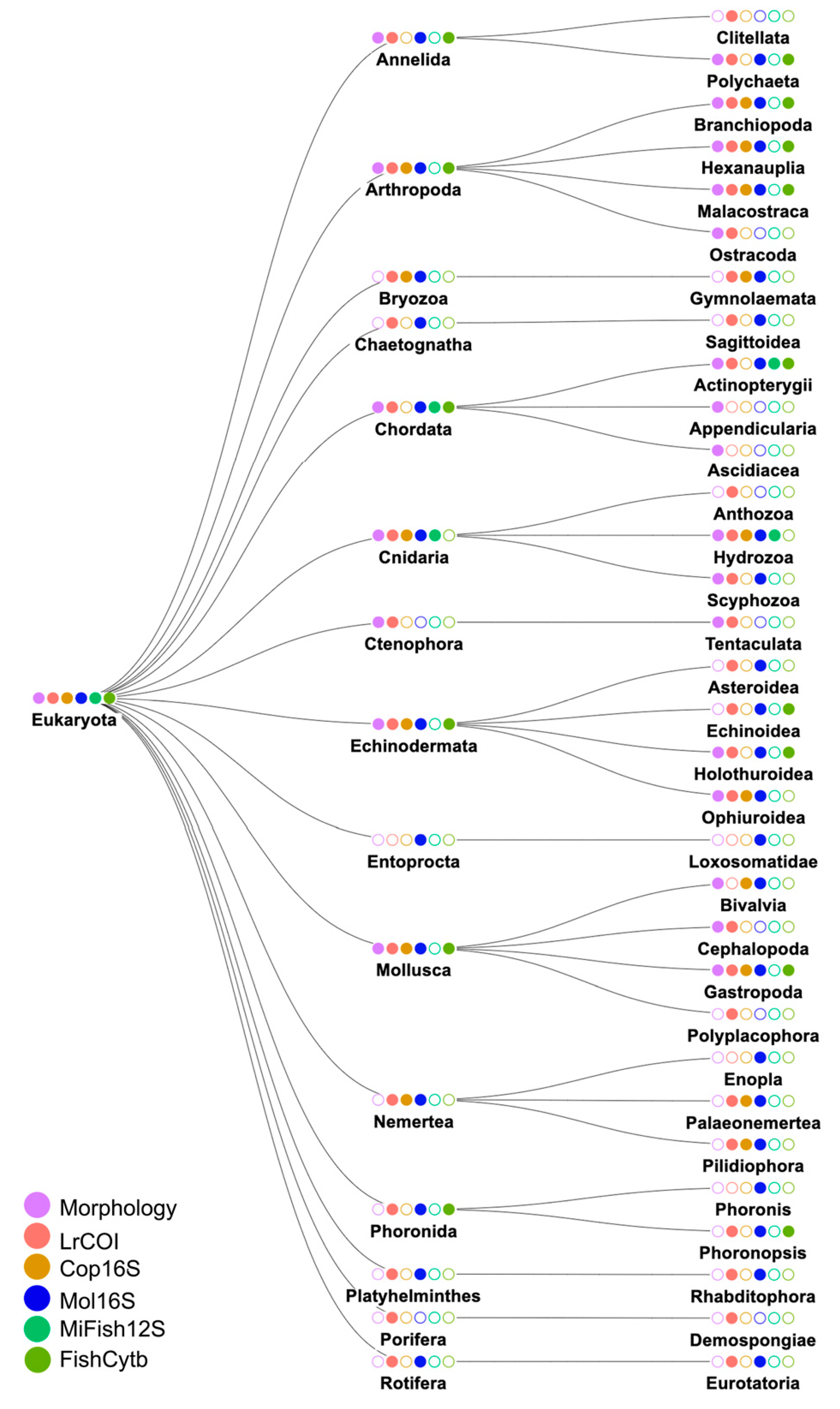
3.2. Species identities and distributions of major taxa
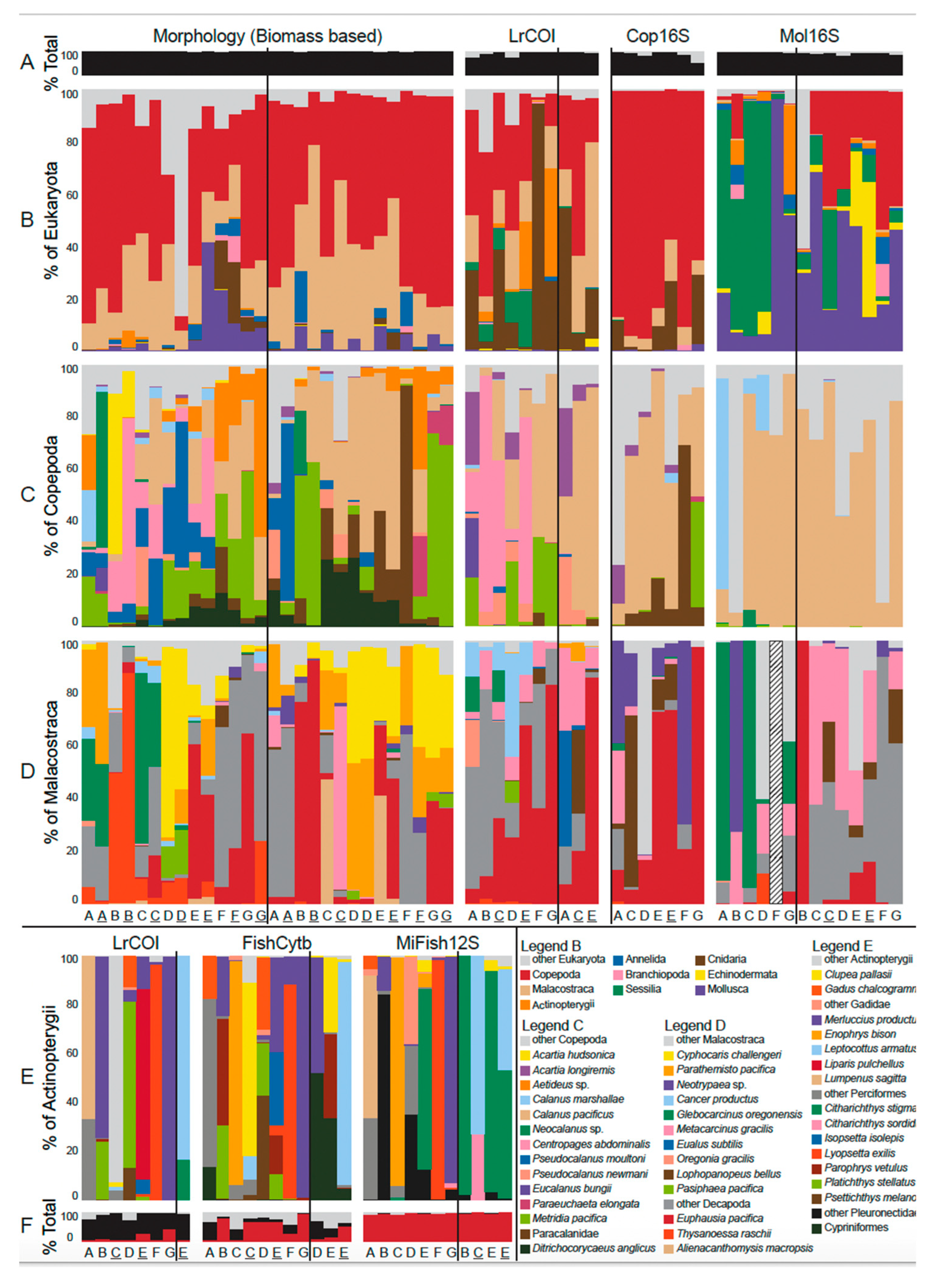
3.2.1. Phylum Annelida; Class Polychaeta (bristleworms, fanworms, clamworms) (Figure 6B, Supplementary Table S4)
3.2.2. Phylum Arthropoda: Class Crustacea (Supplementary Table S4)
3.2.3. Phylum Cnidaria: Class Hydrozoa (hydroids, colonial jellies, siphonophores), Class Anthozoa (sea anemones, corals, sea fans), and Class Scyphozoa (true jellyfish) (Supplementary Figure S2B, Supplementary Table S4)
3.2.4. Phylum Echinodermata: Class Holothuroidea (sea cucumbers), Class Asteroidea (sea stars), Class Ophiuroidea (serpent and brittle stars), and Class Echinoidea (sea urchins, cake urchins) (Supplementary Figure S2C, Supplementary Table S4).
3.2.5. Phylum Mollusca (Supplementary Table S4)
3.2.6. Phylum Chordata: Class Osteichthyes (Bony Fishes), Order Actinopterygii (Ray-finned fishes) (Figure 6E, Supplementary Figure S2E)
4. Discussion
4.1. Metabarcoding marker specificities
4.2. Species detected by morphology and missed by metabarcoding
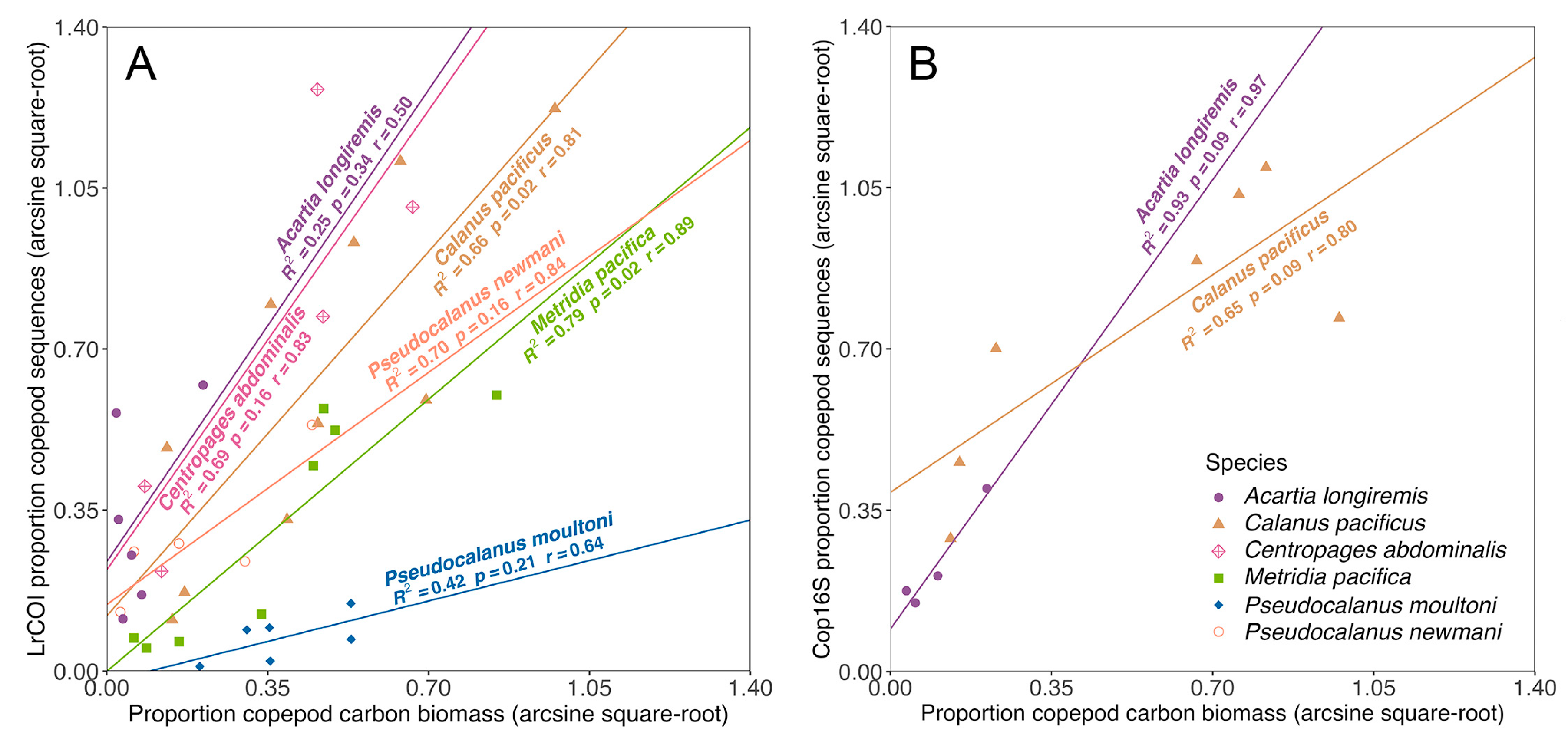
4.3. Sequence read proportion relationships and potential biases
4.4. Threatened or vulnerable fishery species
4.5. Non-indigenous species detections
4.6. Species identifications and reference libraries
4.7. Ecological patterns and trends
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Coyle, K.O.; Pinchuk, A.K.; Eisner, L.B.; Napp J.M. Zooplankton species composition, abundance and biomass on the eastern Bering Sea shelf during summer: the potential role of water-column stability and nutrients in structuring the zooplankton community. Deep Sea Res. II. 2008; 55, 1755–1791. [CrossRef]
- Pomerleau, C.; Sastri, A.R.; Beisner, B.E. Evaluation of functional trait diversity for marine zooplankton communities in the Northeast subarctic Pacific Ocean. J Plankton Res. 2015; 37(4): 726. [CrossRef]
- Hirai, J.; Katakura, S.; Kasai, H.; Nagai, S. Cryptic zooplankton diversity revealed by a metagenetic approach to monitoring metazoan communities in the coastal waters of the Okhotsk Sea, Northeastern Hokkaido. Front Mar Sci. 2017. [CrossRef]
- Gobler, C.J.; Baumann, H. Hypoxia and acidification in ocean ecosystems: coupled dynamics and effects on marine life. Biol Lett. 2016; 5, 20150976. [CrossRef]
- Breitburg, D. Effects of hypoxia, and the balance between hypoxia and enrichment, on coastal fishes and fisheries. Estuaries. 2002; 25, 767–781. 25. [CrossRef]
- Keil, K.E.; Klinger, T.; Keister, J.E.; McLaskey, A.K. Comparative sensitivities of zooplankton to ocean acidification conditions in experimental and natural settings. Front Mar Sci. 2021; 86, 613778. [CrossRef]
- Yasuhara, M.; Hunt, G.; Dowsett, H.J.; Robinson, M.M.; Stoll, D.K. Latitudinal species diversity gradient of marine zooplankton for the last three million years. Ecol Lett. 2012; 15, 1174–1179. [CrossRef]
- Patti, B.; Marco, T.; Angela, C. Interannual summer biodiversity changes in ichthyoplankton assemblages of the Strait of Sicily (Central Mediterranean) over the period 2001–2016. Front Mar Sci. 2022; 9, 960929. [CrossRef]
- Trebitz, A.; Hoffman, J.; Darling, J.; Pilgrim, E.; Kelly, J.; Brown, E.; et al. Early detection monitoring for aquatic non-indigenous species: Optimizing surveillance, incorporating advanced technologies, and identifying research needs. J Environ Manage. 2017; 202(1), 299–310. [CrossRef]
- Ershova, E.A.; Descoteaux, R.; Wangensteen, O.S.; Iken, K.; Hopcroft, R.R.; Smoot, C.; Grebmeier, J.M.; et al. Diversity and distribution of meroplanktonic larvae in the Pacific Arctic and connectivity with adult benthic invertebrate communities. Front Mar Sci. 2019; 6, 490. [CrossRef]
- Ratcliffe, F.C.; Uren Webster, T.M.; Rodriguez-Barreto, D.; O'Rorke, R.; Garcia de Leaniz, C.; Consuegra, S. Quantitative assessment of fish larvae community composition in spawning areas using metabarcoding of bulk samples. Ecol Appl. 2021; 31(3), e02284. [CrossRef]
- Canonico, G.; Buttigieg, P.L.; Montes, E.; Muller-Karger, F.E.; Stepien, C.A.; Wright, D.; et al. Global observational needs and resources for marine biodiversity. Front Mar Sci. 2019; 6, 367. [CrossRef]
- Bucklin, A.; Batta-Lona, P.G.; Questel, J.M.; Wiebe, P.H.; Richardson, D.E.; Copley, N.J.; et al. COI metabarcoding of zooplankton species diversity for time-series monitoring of the NW Atlantic continental shelf. Front Mar Sci. 2022; 9. [CrossRef]
- Shu, L.; Ludwig, A.; Peng, Z. Standards for methods utilizing environmental DNA for detection of fish species: Review. Genes (Basel); 2020 Mar 11; 11(3), 296. [CrossRef]
- Pietsch, T.; Orr, J.W. Fishes of the Salish Sea: Puget Sound and the Straits of Georgia and Juan de Fuca. 2019; University of Washington Press, Seattle, WA. 1074 pp. ISBN: 9780295743745.
- Keister, J.E.; Winans, A.K.; Herrmann, B. Zooplankton community response to seasonal hypoxia: A test of three hypotheses. Diversity. 2020; 12, 21. 12. [CrossRef]
- Caldeira, K.; Wickett, M.E. Anthropogenic carbon and ocean pH. Nature. 2003; 425. 365. [CrossRef]
- Barton, A.; Hales, B.; Waldbusser, G.C.; Langdon, C.; Feely, R.A. The Pacific oyster, Crassostrea gigas, shows negative correlation to naturally elevated carbon dioxide levels: Implications for near-term ocean acidification effects. Limnol Oceanogr. 2012; 57, 698–710. [CrossRef]
- Feely, R.A.; Doney, S.C.; Cooley, S.R. Ocean acidification: Present conditions and future changes in a high-CO2 world. Oceanography. 2009; 22, 36–47. Available from: https://denning.atmos.colostate.edu/ats760/Readings/15.Acidification.Feeley.pdf. 22.
- Feely, R.A.; Alin, S.R.; Newton, J.; Sabine, C.L.; Warner M.; Krembs, C.; et al. The combined effects of ocean acidification, mixing, and respiration on pH and carbonate saturation in an urbanized estuary. Estuar Coast Shelf Sci. 2010; 88, 442−449. [CrossRef]
- Busch, D.S.; Alin, S.; Feely, R.A.; McElhany, P.; Poe M.; Carter, B.; et al. West Coast region acidification research. 2020; Chapter 5 in NOAA Ocean, Coastal, and Great Lakes Acidification Research Plan: 2020–2029, Jewett, E.B.; Osborne, E.B.; Arzayus, K.M.; Osgood. K.; DeAngelo, B.J.; Mintz, J.M., Eds. https://oceanacidification.noaa.gov/ResearchPlan2020 (accessed 29 September 2023).
- Zark, M.; Broda, N.K.; Hornick, T.; Grossart, H-P.; Riebesell, U.; Dittmar, T. Ocean acidification experiments in large-scale mesocosms reveal similar dynamics of dissolved organic matter production and viotransformation. Front Mar Sci. 2017; 4, 271. 4. [CrossRef]
- Byrne, M.; Fitzer, S. The impact of environmental acidification on the microstructure and mechanical integrity of marine invertebrate skeletons. Conserv Physiol. 2019; 7(1), coz062. [CrossRef]
- McLaskey, A.K.; Keister, J.E.; McElhany, P.; Olson, M.B.; Busch, D.S.; Maher, M.; et al. (2016). Development of Euphausia pacifica (krill) larvae is impaired under pCO2 levels currently observed in the Northeast Pacific. Mar Ecol Progr Ser. [CrossRef]
- Daewe, U.; Hjøllo, S.S.; Huret, M.; Ji, R.; Maar M..; Niiranen, S.; et al. Predation control of zooplankton dynamics: a review of observations and models, ICES J Mar Sci. 2014; 71(2), 254–271. [CrossRef]
- Brewer, G.D.; Kleppel, G.S.; Dempsey, M. Apparent predation on ichthyoplankton by zooplankton and fishes in nearshore waters of southern California. Mar Biol. 1984; 80, 17–28. 80. [CrossRef]
- Frederiksen, M.; Edwards, M.; Richardson, A.J.; Halliday, N.C.; Wanless, S. From plankton to top predators: bottom-up control of a marine food web across four trophic levels. J Anim Ecol. 2006; 75(6), 1259–1268. [CrossRef]
- Lindeque, P.K.; Parry, H.E.; Harmer, R.A.; Somerfield, P.J.; Atkinson A. Next generation sequencing reveals the hidden diversity of zooplankton assemblages. PLoS One. 2013; 8(11), e81327. [CrossRef]
- van der Loos, L.M.; Nijand, R. Biases in bulk: DNA metabarcoding of marine communities and the methods involved. Mol Ecol Res. 2021; 30, 3270–3288. [CrossRef]
- Song, C.; Choi, H.; Jeon, M.S.; Kim, E.-J.; Jeong, H.G.; Kim, S.; et al. Zooplankton diversity monitoring strategy for the urban coastal region using metabarcoding analysis. Sci Rep. 2021; 11, 24339. [CrossRef]
- Marshall, N.T.; Stepien, C.A. Macroinvertebrate community diversity and habitat quality relationships along a large river from targeted eDNA metabarcoding assays. Environ DNA. 2020; 2(4), 572–586. https:/iro/onlinelibrary.wiley.com/doi/full/10.1002/edn3.90.
- Claver, C.; Canals, O.; de Amézaga, L.G.; Mendibil, I.; Rodriguez-Ezpeleta, N. An automated workflow to assess completeness and curate GenBank for environmental DNA metabarcoding: The marine fish assemblage as case study. Environ DNA. 2023; 00, 1–15. 00. [CrossRef]
- Elbrecht, V.; Leese, F. Validation and development of COI metabarcoding primers for freshwater macroinvertebrate bioassessment. Front Environ Sci. 2017; 5, 11. [CrossRef]
- Zhang, G.K.; Chain, F.J.J.; Abbot., C.L.; Cristescu, M.E. Metabarcoding using multiplexed markers increases species detection in complex zooplankton communities. Evol Appl. 2018; 11, 1901–1914. [CrossRef]
- Takeuchi, A.; Sado, T.; Gotoh, R.O.; et al. New PCR primers for metabarcoding environmental DNA from freshwater eels, genus Anguilla. Sci Rep. 2019; 9, 7977. [CrossRef]
- Klymus, K.E.; Marshall, N.T.; Stepien, C.A. Environmental DNA (eDNA) metabarcoding assays to detect invasive invertebrate species in the Great Lakes. PLoS One. 2017; 12(5), e0177643. [CrossRef]
- Kim, A.R.; Yoon, T.H.; Lee, C.I.; Kang, C.-K.; Kim, H.-W. Metabarcoding analysis of ichthyoplankton in the East/Japan Sea using the novel fish-specific universal primer set, Front Mar Sci. 2012; 8. [CrossRef]
- Kocher, T.D.; Thomas, W.K.; Meyer, A.; Edwards, V.; Paabo, S.; Villablanca, F.X.; Wilson, A.C. Dynamics of mitochondrial DNA evolution in animals: Amplification and sequencing with conserved primers. Proc Natl Aca. Sci, USA. 1989; 86, 6196–6200. [CrossRef]
- Palumbi, S. Nucleic Acids II: The polymerase chain reaction. Ch. 7, pp. 205–248. In: Molecular Systematics, 2nd ed. 1996. Hillis, D.M.; Moritz, C.; Mable, B.K., Eds. Sinauer Assoc., Sunderland, MA. 655 pp. ISBN-10: 0878932828, ISBN-13: 978-0878932825.
- Stepien, C.A.; Kocher, T.D. Molecules and morphology in studies of fish evolution. Ch.1, pp. 1–11 in: Kocher, TD.; Stepien CA., eds. Molecular Systematics of Fishes. 1997; Academic Press, 313 pp. [CrossRef]
- Lavrov, D.V. Mitochondrial genomes in invertebrate animals. In: Bell, E., ed. Molecular Life Sciences. 2014; Springer, New York, NY. [CrossRef]
- Cole, L.W. The evolution of per-cell organelle number. Front Cell Dev Biol. 2016; Aug 18, 4:85. 18. [CrossRef]
- Hebert, P.D.; Cywinska, A.; Ball, S.L.; deWaard, J.R. Biological identifications through DNA barcodes. Proc Biol Sci. 2003; Feb 2. 270(1512):313–321. [CrossRef]
- Bucklin, A.; Steinke, D.; Blanco-Bercial, L. DNA barcoding of marine Metazoa. Annu Rev Mar Sci. 2011; 3, 471–508. [CrossRef]
- Leray, M.; Knowlton, N. DNA barcoding and metabarcoding of standardized samples reveal patterns of marine benthic diversity. Proc Natl Acad Sci USA. 2015; 112(7), 2076–2081. [CrossRef]
- Günther, B.; Knebelsberger, T.; Neumann, H.; Laakmann, S.; Arbizu, PM. Metabarcoding of marine environmental DNA based on mitochondrial and nuclear genes. Sci Rep. 2018; 8: 14822. [CrossRef]
- Radulovici, A.E.; Vieira, P.E.; Duarte, S.; Teixeira, M.A.L.; Borges, L.M.S.; Deagle, B.E.; et al. Revision and annotation of DNA barcode records for marine invertebrates: report of the 8th iBOL conference hackathon. Metabarcoding Metagenom. 2021; 5, e67862. [CrossRef]
- Kim, H.; Yoon, M.; Kim, H.J. The complete mitochondrial genome of rockfish Sebastes oculatus Valenciennes, 1833 from southwest Atlantic Ocean. Mitochondrial DNA B Resour. 2019; Oct 9; 4(2), 3407–3408. PMID: 33366015; PMCID: PMC7707287. [CrossRef]
- Leray, M.; Yang, J.Y.; Meyer, C.P.; Mills, S.C.; Agudelo, N.; Ranwez, V.; et al. A new versatile primer set targeting a short fragment of the mitochondrial COI region for metabarcoding metazoan diversity: application for characterizing coral reef fish gut contents. Frontiers in Zoology, 2013; 10, 34. [CrossRef]
- Geller, J.; Meyer, C.; Hawk, H. Redesign of PCR primers for mitochondrial cytochrome c oxidase subunit I for marine invertebrates and application in all-taxa biotic surveys. Molecular Ecology Resources. 2013. [CrossRef] [PubMed]
- Casey, J.M.; Ransome, E.; Collins, A.G.; Mahardini, A.; Kurniasih, E.M.; Sembiring, A.; et al. DNA metabarcoding marker choice skews perception of marine eukaryotic biodiversity. Environment DNA, 2021; 3, 1229–1246. [CrossRef]
- Pappalardo, P.; Collins, A.G.; Pagenkopp Lohan, K.M.; Hanson, K.M.; Truskey, S.B.; Jaeckle, W.; et al. The role of taxonomic expertise in interpretation of metabarcoding studies. ICES J Mar Sci. 2021; 78, 3397–3410. [CrossRef]
- Yang, L.; Tan, Z.; Wang, D.; Xue, L.; Guan, M.; Huang, T.; Li, R. Species identification through mitochondrial rRNA genetic analysis. Sci Reports, 2014; 4, 4089. [CrossRef]
- Wang, H-Y.; Lee, S-C. Secondary structure of mitochondrial 12S rRNA among fish and its phylogenetic applications. Mol Biol Evol. 2022; 19(2), 138–148. [CrossRef]
- Clarke, L.J.; Beard, J.M.; Swadling, K.M.; Deagle, B.E. Effect of marker choice and thermal cycling protocol on zooplankton DNA metabarcoding studies. Ecol Evol. 2017; 7, 873–883. [CrossRef]
- Stepien, C.A.; Andrews, K.; Elz, A.; Marshall, N.T.; Snyder, M.R. Understanding marine community responses in species composition, diversity, and population genetics: targeted metagenomics from environmental DNA and plankton samples. Symposium 6647: Application and Innovation in the use of Environmental DNA (eDNA) for Use with Aquatic Species. American Fisheries Society Annual Meeting. 2018; 32989, Atlantic City, New Jersey, USA. 2018; Available from: https://afs.confex.com/afs/2018/oral/papers/viewonly.cgi?password=299439&username=32989 (Accessed 30 September 2023). 30 September.
- Strong, E.E.; Gargominy, O.; Ponder, W.F.; Bouchet, P. 2008. Global diversity of gastropods (Gastropoda; Mollusca) in freshwater. Hydrobiologia. 2008; 595, 149–166. [CrossRef]
- Dobrzycka-Krahel, A.; Stepien, C.A.; Nuc, Z. Neocosmopolitan distributions of invertebrate aquatic invasive species due to euryhaline geographic history and human-mediated dispersal: Ponto-Caspian versus other geographic origins. Ecol Process. 2023; 12, 2. [CrossRef]
- Miya, M.; Sato, Y.; Fukunaga, T.; Sado, T.; Poulsen, JY.; Sato, K.; Minamoto, T.; et al. MiFish, a set of universal PCR primers for metabarcoding environmental DNA from fishes: detection of more than 230 subtropical marine species. Royal Society Open Science. 2015; 2(7). [CrossRef]
- Stoeckle, M.Y.; Adolf, J.; Charlop-Powers, Z.; Dunton, K.J.; Hinks, G.; VanMorter, S.M. Trawl and eDNA assessment of marine fish diversity, seasonality, and relative abundance in coastal New Jersey, USA. ICES J Mar Sci. 2021; 78(1), 293–304. [CrossRef]
- Stoeckle, MY.; Adolf, J.; Ausubel, J.H.; Charlop-Powers, Z.; Dunton, K.J.; Hinks, G. Current laboratory protocols for detecting fish species with environmental DNA optimize sensitivity and reproducibility, especially for more abundant populations, ICES J Mar Sci. 2022; 79(2), 403–412. [CrossRef]
- Snyder, M.R.; Stepien, C.A. Increasing confidence for discerning species and population compositions from metabarcoding assays of environmental samples. Metabarcod Metagenomics. 2020; 4, 47–63. [CrossRef]
- Snyder, M.R.; Stepien, C.A.; Marshall, N.T.; Scheppler, H.; Black, C.; Czajkowski, K. Detecting aquatic invasive species in bait and pond stores with targeted environmental DNA high-throughput sequencing metabarcode assays: Angler, retailer, and manager implications. Biolog Conserv. 2020; 245, 108430. 1084. [Google Scholar] [CrossRef]
- Carrete, V.G.; Wiens, J.J. Why are there so few fish in the sea? Proc R Soc B. 2012; 279, 2323–2329. [CrossRef]
- Northwest Association of Networked Ocean Observing Systems (NANOOS), Pacific Northwest Regional Association of the Integrated Ocean Observing System (IOOS), NVS Salish Cruises Reports, http://nvs.nanoos.org/CruiseSalish (accessed 29 September 2023).
- Winans, A.K.; Herrmann, B.; Keister, J.E. Spatio-temporal variation in zooplankton community composition in the southern Salish Sea: changes during the 2015–2016 Pacific marine heatwave. 2023; Prog Oceanog. Accepted, Forthcoming.
- Bucklin, A. Methods for population genetic analysis of zooplankton. In: Harris, R.; Wiebe, P.; Lenz, J.; Skjoldal, H.R.; Huntley, M. ICES Zooplankton Methodology Manual. 2000; 533–570. Academic Press, ISBN 9780123276452. [CrossRef]
- Gardner, G.A.; Szabo, I. British Columbia Pelagic Marine Copepoda: An Identification Manual and Annotated Bibliography, Ottawa: Dept. of Fisheries and Oceans. 1982; Available from [internet] http://publications.gc.ca/collections/collection_2016/mpo-dfo/Fs41-31-62-eng.pdf (accessed 30 September 2023).
- Shanks, A.L. An Identification Guide to the Larval Marine Invertebrates of the Pacific Northwest, First ed., Corvallis, Oregon: Oregon State University Press, Corvallis, OR; 2001. ISBN 13: 9780870715310.
- Light, S.F.; Carlton, J.T. The Light and Smith Manual: Intertidal Invertebrates from Central California to Oregon, 4th ed., completely rev. and expanded., Berkeley, Calif.: University of California Press; 2007. ISBN 13: 978-0520239395.
- Wrobel, D.; Mills, C. Pacific Coast Pelagic Invertebrates: a Guide to the Common Gelatinous Animals, Monterey, CA: Sea Challengers. Monterey Bay Aquarium, Monterey, CA; 1998. 9: ISBN 13, 9780. [Google Scholar]
- Lough, R.G. Dynamics of Crab Larvae (Anomura, Brachyura) off the Central Oregon Coast, 1969–1971. Ph.D. Thesis. Published by Oregon State University, School of Oceanography, Corvallis, OR, pp. 28–83; 1975. https://ir.library.oregonstate.edu/concern/parent/j098zc90v/file_sets/rx913s09f (accessed 30 September 2023).
- Sorochan, K.A.; Duguid, W.D.P.; Quijón, P.A. Diagnostic morphological characteristics of laboratory-reared Cancer oregonensis (Brachyura: Cancridae) with recommendations for identifying cancrid zoeae in the Salish Sea. Mar Biol Res, 2015; 11(6), 1–9. [CrossRef]
- Brinton, E.; Ohman, M.D.; Townsend, A.W.; Knight, M.D.; Bridgeman, A.L. Euphausiids of the World Ocean, 2015; v. 1.1. https:///euphausiids.linnaeus.naturalis.nl/linnaeus_ng/app/views/introduction/topic.php?id=11.
- Thorp, J.H.; Covich, A.P. (eds) Ecology and Classification of North American Freshwater Invertebrates. 1991; Academic Press, Inc., Boston, MA. 911pp. ISBN 9780080530673.
- Marsh, T.; Wood, T.S. Results of a freshwater bryozoan survey in the Pacific Northwest. In: Jackson, W.; Jones, B; Jones S., eds. Bryozoan Studies. 2001; Swets and Zeitlinger, Lisse., ISBN 90-5809-388-3.
- Van Guelpen, L.; Markle, D.F.; Duggan, D.J. An evaluation of accuracy, precision, and speed of several zooplankton sub-sampling techniques. J Cons int Expl Mer. 1982; 40, 226–236. https://dokumen.tips/documents/an-evaluation-of-accuracy-precision-and-speed-of-several-zooplanktonsubsampling. html?page=1.
- Hillis, D.M.; Mable, B.K.; Larson, A.; Davis, S.K.; Zimmer, E.A. Nucleic Acids IV: Sequencing and cloning, Ch. 9, pp 321–384. In: Hillis, D.M.; Moritz, C.; Mable, B.K., eds. Molecular Systematics. 1996; 2nd Ed., Sinauer Assoc., Sunderland, MA. 655 pp. ISBN-10: 0878932828, ISBN-13: 978-087893282578.
- Marshall, N.T.; Stepien, C.A. Invasion genetics from eDNA and thousands of larvae: A targeted metabarcoding assay that distinguishes species and population variation of zebra and quagga mussels. Ecol Evol. 2019; 9, 1–24. [CrossRef]
- Fadrosh, D.W.; Ma, B.; Gajer, P.; Sengamalay, N.; Ott, S.; Brotman, R.M.; et al. An improved dual-indexing approach for multiplexed 16S rRNA gene sequencing on the Illumina MiSeqplatform. Microbiome 2014; 2,6. [CrossRef]
- Wu, L.; Wen, C.; Qin, Y.; Wu, L.; Wen, Ch.; Yin, H.; et al. Phasing amplicon sequencing on Illumina Miseq for robust environmental microbial community analysis. BMC Microbiol. 2015; 15, 125. [CrossRef]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal, 2011; [S.l.], v. 17, n. 1, p. pp. 10–12, May 2011. ISSN 2226-6089. doi: 10.14806/ej.17.1.200. Available from: https://journal.embnet.org/index.php/embnetjournal/article/view/200. Date accessed: 30 September 2023. [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods, 2016; 13. 581–583. [CrossRef]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: architecture and applications. BMC Bioinformatics. 2009; Dec 15;10:421. PMID: 20003500; PMCID: PMC2803857. [CrossRef]
- National Center for Biotechnology Information (NCBI), National Institutes of Health (NIH), National Library of Medicine, USA. 2021; Available from: https://blast.ncbi.nlm.nih.gov/blast/Blast.cgi?CMD=Web&PAGE_TYPE=BlastHome (accessed 30 September 2023).
- Menzel, P. Subtree program in: Taxonomy Tools. 2021; Available from: https://github.com/pmenzel/taxonomy-tools (accessed 30 September 2023).
- Shen, W.; Ren, H. TaxonKit: a practical and efficient NCBI taxonomy toolkit, J Genet Genomics. 2021; 48(9), 844–850. [CrossRef]
- Ondov, B.D.; Bergman, N.H.; Phillippy, A.M. Interactive metagenomic visualization in a Web browser. BMC Bioinformatics, 2011; 12. 385. [CrossRef]
- McMurdie, P.J.; Holmes, S. Phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE, 2013; 8(4). Program available from: https://www.bioconductor.org/packages/release/bioc/html/phyloseq.html) Accessed 30 September 2023. [CrossRef]
- Smith, S. NMDS Phyloseq. Classification clustering genetic variability metagenomics microbiome multiple comparison sequencing software visualization. 2022; Available from [internet]: https://rdrr.io/github/schuyler-smith/phyloschuyler/man/nmds_phyloseq.html (accessed 30 September 2023).
- Chen, H.; Boutros, P.C. VennDiagram: a package for the generation of highly-customizable Venn and Euler diagrams in R. BMC Bioinformatics, 2011 Dec;12(1):1–7. [CrossRef]
- visNetwork, an R package for interactive network visualization, r. 2.1.1. 2022; Available from: https://datastormopen.github.io/visNetwork/ (accessed 30 September 2023).
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Statist. Soc. B. 1995; 57, 289–300.
- Morisette, J.; Burgiel, S.; Brantley, K.; Daniel, W.M.; Darling, J.; Davis, J.; et al. Strategic considerations for invasive species managers in the utilization of environmental DNA (eDNA): steps for incorporating this powerful surveillance tool. Manage Biol Invasion, 2021; 12(3). 747–775. [CrossRef]
- Rabinowitz, C.N.; Brown, S.D.; McAllister, S.M.; Winans, A.K.; Keister, J.E.; and Galaska, M.P. The complete mitochondrial genome of Cyphocaris challengeri (Amphipoda: Cyphocarididae). Mitochondrial DNA Part B. Forthcoming.
- 96. Shih, C-T.; Hendrycks, E.A. A new species and new records of the genus Vibilia Milne Edwards, 1830 (Amphipoda: Hyperiidea: Vibiliidae) occurring in the eastern Pacific Ocean. J Nat Hist. 2003; 37(3), 253–296. [CrossRef]
- Andrikou, C.; Passamaneck, Y.J.; Lowe, C.J.; Martindale, M.Q.; Hejnol, A. Molecular patterning during the development of Phoronopsis harmeri reveals similarities to rhynchonelliform brachiopods. EvoDevo, 2019; 10, 33. [CrossRef]
- Blake, J.A.; Ruff, R. pp. 309-410, In: Carleton J.T., ed. The Light and Smith Manual: Intertidal Invertebrates from Central California to Oregon (4th edition), 2007; University of California, Berkeley, CA. ISBN-10: 0520239393, ISBN-13: 978-0520239395.
- Ruiz, G. Proceraea okadai: Annelids-Polychaetes. Smithsonian Environmental Research Center’s National Estuarine and Marine Exotic Species Information System (NEMESIS). Available from: https://invasions.si.edu/nemesis/species_summary/-683 (Accessed 29 September 2023).
- Nishizawa, R.; Sato, M.; Furota, T. et al. Cryptic invasion of northeast Pacific estuaries by the Asian polychaete, Hediste diadroma (Nereididae). Mar Biol. 2014; 161. 187–194. [CrossRef]
- 101. Snyder, M,R. Environmental DNA Detection and Population Genetic Patterns of Native and Invasive Great Lakes Fishes. /: of Toledo, Toledo, Ohio. 2019. Available from: http, 29 September 2019.
- Ershova, E.A.; Wangensteen, O.S.; Descoteaux, R.; Barth-Jensen, C.; Præbel, K. Metabarcoding as a quantitative tool for estimating biodiversity and relative biomass of marine zooplankton, ICES J Mar Sci. 2021; 78(9), 3342–3355. [CrossRef]
- Matthews, S.A.; Goetze, E.; Ohman, M.D. Recommendations for interpreting zooplankton metabarcoding and integrating molecular methods with morphological analyses. ICES J Mar Sci, 2021; 78, 3387–3396. 3387; 78. [Google Scholar] [CrossRef]
- Hänfling, B.; Handley, L.L.; Read, D.S.; Hahn, C.; Li, J.; Nichols, P.; Blackman, R.C.; Oliver, A.; Winfield, I.J. Environmental DNA metabarcoding of lake fish communities reflects long-term data from established survey methods. Mol Ecol, 2016; 25, 3101–3119. [CrossRef]
- Thomsen, P.F.; Møller, P.R.; Sigsgaard, E.E.; Knudsen, S.W.; Jørgensen, O.A.; Willerslev, E. Environmental DNA from seawater samples correlate with trawl catches of subarctic, deepwater fishes. PLoS ONE, 2016; 11, e0165252. [CrossRef]
- Parsons, K.M.; Everett, M.; Dahlheim, M.; Park, L. Water, water everywhere: Environmental DNA can unlock population structure in elusive marine species. R Soc Open Sci, 2018; 5, 180537. [CrossRef]
- Shaw, J.L.A.; Clarke, L.J.; Wedderburn, S.D.; Barnes, T.C.; Weyrich, L.S.; Cooper, A. Comparison of environmental DNA metabarcoding and conventional fish survey methods in a river system. Biol Conserv. 2016; 197, 131–138. [CrossRef]
- Gillet, B.; Cottet, M.; Destanque, T.; Kue, K.; Descloux, S.; Chanudet, V.; Hughes, S. Direct fishing and eDNA metabarcoding for biomonitoring during a 3-year survey significantly improves number of fish detected around a South East Asian reservoir. PLoS ONE, 2018; 13, e0208592. [CrossRef]
- Lamb, P.D.; Hunter, E.; Pinnegar, J.K.; Creer, S.; Davies, R.G.; Taylor, M.I. How quantitative is metabarcoding: A meta-analytical approach. Mol Ecol, 4: Jan (2). [CrossRef]
- Shelton, A.O.; Gold, Z.J.; Jensen, A.J.; D′Agnese, E.; Andruszkiewicz, A.; Van Cise, A.; Gallego, R.; et al. Toward quantitative metabarcoding. Ecology. 2023; 104( 2), e3906. [CrossRef]
- Xiong, W.; Li, H.; Zhan, A. Early detection of invasive species in marine ecosystems using high-throughput sequencing: technical challenges and possible solutions. Mar Biol, 2022; 163, 1–139. [CrossRef]
- Alberdi, A.; Aizpurua, O.; Gilbert, M.T.P.; Bohmann, K. Scrutinizing key steps for reliable metabarcoding of environmental samples. Methods Ecol Evol. 2017; 9, 134–147. [CrossRef]
- Piñol, J.; Mir, G.; Gomez-Polo, P.; Agustí, N. Universal and blocking primer mismatches limit the use of high-throughput DNA sequencing for the quantitative metabarcoding of arthropods. Mol Ecol Res, 2015; 15(4), 819–830. [CrossRef]
- Silverman, J.D.; Bloom, R.J.; Jiang, S.; Durand, H.K.; Dallow, E.; Mukherjee, S.; David, L.A. Measuring and mitigating PCR bias in microbiota datasets. PLoS Comput Biol, 2021; 17(7), e1009113. [CrossRef]
- Hoshino, T.; Nakao, R.; Doi, H.; Minamoto, T. Simultaneous absolute quantification and sequencing of fish environmental DNA in a mesocosm by quantitative sequencing technique. Sci Rep, 2021; 11, 4372. [CrossRef]
- Gold, Z.; Kelly, R.P.; Shelton, A.O.; Thompson, A.R.; Goodwin, K.D.; Gallego, R.; Parsons, K.M.; Thompson, LR.; Kacev, D.; Barber, P.H. Archived DNA reveals marine heatwave-associated shifts in fish assemblages. Environm DNA, 2023; 00:1–14. [CrossRef]
- McCarthy, A.; Rajabi, H.; McClenaghan, B.; Fahner, NA.; Porter, E.; Singer, G.A.C.; Hajibabaei, M. Comparative analysis of fish eDNA reveals higher sensitivity achieved through targeted sequence-based metabarcoding. Mol Ecol Res. 2022. [CrossRef]
- Jiang, R.; Lusana, J.L.; Chen, Y. High-throughput DNA metabarcoding as an approach for ichthyoplankton survey in Oujiang River Estuary, China. Diversity, 2022; 14(12):1111. [CrossRef]
- Nobile, A.B.; Freitas-Souza, D.; Ruiz-Ruano, F.J.; Nobile, M.L.M.O.; Costa, G.O.; de Lima, F.P.; et al. DNA metabarcoding of Neotropical ichthyoplankton: Enabling high accuracy with lower cost. Metabarcoding Metagenom, 2019; 3, e35060. [CrossRef]
- Mariac, C.; Renno, J.; Garcia-davila, C.; Vigouroux, Y.; Mejia, E.; Angulo, C.; et al. Species-level ichthyoplankton dynamics for 97 fishes in two major river basins of the Amazon using quantitative metabarcoding. Mol Ecol, 2022; 31(6), 1627–1648. [CrossRef]
- NOAA Fisheries; 2022. Chum Salmon-Protected. Available from: https://www.fisheries.noaa.gov/species/chum-salmonprotected Accessed 29 September 2023.
- Washington Department of Fish and Wildlife. Chum salmon, (Hood Canal Summer ESU) (Oncorhynchus keta pop. 2) [Internet] https://wdfw.wa.gov/species-habitats/species/oncorhynchus-keta-pop-2. Accessed 22 September 2023.
- Washington Department of Fish and Wildlife. Walleye pollock (South Puget Sound) (Gadus chalcogrammus) Available from: https://wdfw.wa.gov/species-habitats/species/gadus-chalcogrammus#desc-range Accessed 29 September 2023.
- Small, M.P.; Loxterman, J.L.; Frye, A.E.; Vonbargen, J.F.; Bowman, C.; Young, S.F. Temporal and spatial genetic structure among some Pacific herring populations in Puget Sound and the Southern Strait of Georgia. Trans Amer Fish Soc, 2005; 134(5). 1329–1341. [CrossRef]
- NOAA Fisheries. Species Directory. Pacific herring. Available from: https://www.fisheries.noaa.gov/species/pacific-herring Accessed 22 September 2023.
- Washington Department of Fish and Wildlife. Pacific herring (Clupea pallasii). https://wdfw.wa.gov/specieshabitats/species/clupea-pallasii#conservation Accessed 29 September 2023.
- Washington Department of Fish and Wildlife. Pacific hake (Merluccius productus). Available from: https://wdfw.wa.gov/species-habitats/species/merluccius-productus Accessed 29 September 2023.
- Chittaro, P.; Grandin, C.; Pacunski, R.; Zabel, R. Five decades of change in somatic growth of Pacific hake from Puget Sound and Strait of Georgia. Peer Peer J, 2022; 10, e13577. [CrossRef]
- Draheim, R.W. Annual Progress Report to Pacific States Fisheries Commission. Lower Columbia River Aquatic Nonindigenous Species Survey 2001–2003. 2002; Phase I: Literature Review and Sampling Plan. https://citeseerx.ist.psu.edu/document?repid=rep1&type.
- Emmett, R.L.; Stone, S.L.; Hinton, S.A.; Monaco, M.E. Distribution and abundance of fishes and invertebrates in west coast estuaries, 1991; Volume II: Species life history summaries. ELMR Rep. No. 8. NOAA/NOS Strategic Environmental Assessments Division, Rockville, MD, 329 p. Available from: https://repository.library.noaa.gov/view/noaa/2871 (accessed 25 September 2023).
- Hasselman, D.J.; Hinrichsen, R.A.; Shields, B.A.; Ebbesmeyer, C.C. The rapid establishment, dispersal, and increased abundance of invasive American shad in the Pacific Northwest. Fisheries, 2012; 37(3), 103–113. [CrossRef]
- Grason, E.W.; McDonald, P.S.; Adams, J.W.; Litle, K.; Apple, J.K.; Pleus, A. Citizen science program detects range expansion of the globally invasive European green crab in Washington State (USA). Manage Biolog Invas. 2018; 9, 39–47. [CrossRef]
- Behrens Yamada, S.; Thomson, R.E.; Gillespie, G.E.; Norgard, T.C. Lifting barriers to range expansion: the European green crab Carcinus maenas (Linnaeus, 1758) enters the Salish Sea. Journal of Shellfish Research, 2017; 36, 201–208. [CrossRef]
- Washington Department of Fish and Wildlife. European green crab (Carcinus maenas). 2023; Available from: https://wdfw.wa.gov/species-habitats/invasive/carcinus-maenas#desc-range (Accessed 22 September 2023).
- Westfall, K.M.; Therriault, T.W.; Abbott, C.L. Targeted next-generation sequencing of environmental DNA improves detection of invasive European green crab (Carcinus maenas). Environ DNA, 2022; 4(2), 440–452. [CrossRef]
- Fernández-Álvarez, F.Á.; Machordom, A. DNA barcoding reveals a cryptic nemertean invasion in Atlantic and Mediterranean waters. Helgol Mar Res, 2013; 67, 599–605. 67. [CrossRef]
- USGS (US Geological Survey). Invasive Species Program: Early Detection and Rapid Response. 2023; Available from: https://www.usgs.gov/tools/national-early-detection-and-rapid-response-edrr-framework (Accessed 22 September 2023).
- Bianchi, FM.; Gonçalves, L.T. Getting science priorities straight: how to increase the reliability of specimen identification? Biol Lett. 2021; 17, 20200874. [CrossRef]
- Orr, J.W.; Spies, I.; Stevenson, D.E.; Longo, G.C.; Kai, Y.; Ghods, S.; Hollowed, M. Molecular phylogenetics of snailfishes (Cottoidei: Liparidae) based on mtDNA and RADseq genomic analyses, with comments on selected morphological characters. Zootaxa. 2019; 4642 (1), 001–079. [CrossRef]
- Hoban, M.L.; Whitney, J.; Collins, A.G.; Meyer, C.; Murphy, K.R.; Reft, A.J.; Bemis, K.E. 2022. Skimming for barcodes: rapid production of mitochondrial genome and nuclear ribosomal repeat reference markers through shallow shotgun sequencing. PeerJ, 10:e13790. [CrossRef]
- Smithsonian/NOAA/BOEM and Partners Genome Skimming Project. 2023; Available from: https://geomedb.org/workbench/project-overview?projectId=446 (Accessed 17 September 2023).
- Bucklin, A.; Peijnenburg, K.T.C.A.; Kosobokova, K.N.; O’Brien, T.D.; Blanco-Bercial, L.; Cornils, A.; et al. Toward a global reference database of COI barcodes for marine zooplankton. Mar Biol, 2021; 168. 78.
- Meyer, C.; Duffy, E.; Collins, A.; Paulay, G.; Wetzer, R. The US Ocean Biocode. Marine Technology Society Journal, 2021; 55(3), 140–141. [CrossRef]
- NOAA Genome Skimming of Marine Animals Inhabiting the US Exclusive Economic Zone. NIH (National Institutes of Health) https://www.ncbi.nlm.nih.gov/bioproject/720393 (Accessed 26 September 2023).
- Gold, Z.; Wall, A.R.; Schweizer, T.M.; Pentcheff, N.D.; Curd, E.E.; Barber, P.H.; et al. A manager’s guide to using eDNA metabarcoding in marine ecosystems. PeerJ, 2022; 10:e14071. [CrossRef]
- Stiassny, M.H.; Mittermayer, F.H.; Sswat, M.; Voss, R.; Jutfelt, F.; Chierici, M.; et al. Ocean acidification effects on Atlantic cod larval survival and recruitment to the fished population. PLoS ONE, 2016; 11(8). e0155448. [CrossRef]
- Sommer, S.A.; Van Woudenberg, L.; Lenz, P.H.; Cepeda, G.; Goetze, E. Vertical gradients in species richness and community composition across the twilight zone in the North Pacific Subtropical Gyre. Mol Ecol, 2017; 26, 6136–6156. [CrossRef]
- Casas, L.; Pearman, J.K.; Irigoien, X. Metabarcoding reveals seasonal and temperature-dependent succession of zooplankton communities in the Red Sea. Front Mar Sci, 2017; 4, 241. [CrossRef]
- Hirai, J.; Tachibana, A.; Tsuda, A. Large-scale metabarcoding analysis of epipelagic and mesopelagic copepods in the Pacific. PLoS One, 2020; 15(5), e0233189. [CrossRef]
- Kobari, T.; Tokumo, Y.; Sato, I.; Kume, G.; Hirai, J. Metabarcoding analysis of trophic sources and linkages in the plankton community of the Kuroshio and neighboring waters. Sci Rep, 2021; 11, 23265. [CrossRef]
- USGS (US Geological Survey). Nonindigenous Aquatic Species. Website: https://nas.er.usgs.gov/queries/SpeciesList.aspx?group=&state=WA&Sortby=2 (Accessed 29 September 2023).

| Site and season |
Latitude | Longitude | Sampling date |
Tow start time |
Salinity (PSU) range (and mean) |
Fluorescence (mg/m3) range (and mean) |
pH range (and mean) |
Temp. (°C) range (and mean) |
Diss.O2 (mg/L) range (and mean) |
|---|---|---|---|---|---|---|---|---|---|
| A (P22) Spring |
48.2723 | -123.0212 | 04/08/18 | 11:46 | 30.49–32.91 (31.62) |
0.15–0.45 (0.30) |
7.70–7.80 (7.75) |
8.15–8.57 (8.28) |
4.96–8.10 (6.64) |
| B (P4) Spring |
48.2437 | -122.5562 | 04/07/18 | 20:20 | 24.69-29.41 (27.97) |
0.24–1.20 (0.51) |
7.56–7.84 (7.63) |
8.70–9.57 (9.07) |
3.72–9.29 (6.23) |
| C (P8) Spring |
47.8989 | -122.6054 | 04/09/18 | 07:29 | 29.10–30.62 (30.08) |
0.32–0.89 (0.46) |
7.73–7.85 (7.79) |
8.44–9.19 (8.61) |
7.81–9.09 (8.21) |
| D (P28) Spring |
47.7047 | -122.4544 | 04/07/18 | 12:39 | 28.97–29.65 (29.47) |
0.20–0.77 (0.33) |
7.74–7.76 (7.75) |
8.32–8.73 (8.38) |
3.34–3.69 (3.42) |
| E (P38) Spring |
47.2776 | -122.7085 | 04/11/18 | 10:55 | 28.42–28.91 (28.78) |
0.41–21.20 (3.06) |
7.72–8.19 (7.81) |
8.43–9.47 (8.64) |
8.12–12.64 (9.06) |
| F (P402) Spring |
47.3558 | -123.0239 | 04/09/18 | 17:10 | 25.94–30.07 (29.65) |
0.21–6.61 (0.68) |
7.43–7.79 (7.48) |
10.15–10.97 (10.26) |
3.51–9.21 (4.20) |
| G (P12) Spring |
47.4256 | -123.1074 | 04/09/18 | 14:59 | 26.92–30.39 (30.01) |
0.12–1.32 (0.24) |
7.41–7.55 (7.48) |
9.54–10.58 (10.11) |
3.14–7.43 (4.24) |
| A (P22) Autumn |
48.27268 | -123.02076 | 09/12/18 | 08:49 | 31.43–32.99 (32.26) |
0.12–0.46 (0.24) |
7.62–7.75 (7.67) |
8.81–11.01 (9.85) |
4.38–5.37 (4.91) |
| B (P4) Autumn |
48.2428 | -122.55274 | 09/11/18 | 18:17 | 26.91–30.18 (29.62) |
0.17–12.51 (1.87) |
7.42–8.27 (7.53) |
10.95–14.50 (11.51) |
5.64–7.03 (5.76) |
| C (P8) Autumn |
47.89546 | -122.60412 | 09/12/18 | 15:36 | 29.81–31.01 (30.83) |
0.44–3.80 (0.78) |
7.72–7.83 (7.74) |
11.84–12.93 (12.03) |
5.79–6.55 (6.11) |
| D (P28) Autumn |
47.70974 | -122.45264 | 09/11/18 | 11:44 | 30.05–30.69 (30.52) |
0.13–0.87 (0.26) |
7.68–7.79 (7.73) |
12.30–13.59 (12.56) |
5.61–6.91 (5.82) |
| E (P38) Autumn |
47.2767 | -122.7086 | 09/15/18 | 11:30 | 29.62–30.12 (30.00) |
0.09–6.02 (0.59) |
7.71–7.95 (7.75) |
13.38–14.75 (13.60) |
6.24–7.45 (6.57) |
| F (P402) Autumn |
47.35682 | -123.02252 | 09/13/18 | 16:26 | 28.55–30.42 (29.88) |
0.12–5.76 (0.74) |
7.24–8.33 (7.30) |
9.86–17.01 (10.39) |
1.10–10.68 (2.02) |
| G (P12) Autumn |
47.42576 | -123.10922 | 09/13/18 | 14:00 | 27.89–30.30 (29.92) |
0.05–5.24 (0.51) |
7.32–7.65 (7.38) |
9.82–13.01 (10.29) |
2.50–5.72 (3.05) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





