1. Introduction
Infection with the influenza virus is one of the most common diseases in the world, as it might infect 20% of the world's population each season, which results in a death toll of 650 thousand people each year [
1]. The main problem is that the influenza virus genome mutates at a high rate because its polymerase complex lacks a proofreading mechanism. This character leads to the constant emergence of strains that are resistant to previously effective drugs and are not recognizable by the immune system that has been exposed to previous vaccine strains [
2,
3]. Thus, a mutation in the M2 gene, leading to the S31N substitution in the M2 protein, results in resistance to adamantanes, which are no longer recommended for treatment, since in 2013, 45% of influenza A strains became resistant to them [
4].
Influenza A virus genome contains eight negative-sense, single-stranded RNA gene segments (HA, NA, M, PB1, PB2, PA, NS, and NP). Any mutation in one of these segments might lead to a remarkable change in the virus characteristics. Studying these mutations provides significant information to understand the viral genome function and the role of each mutation in viral pathogenicity [
2,
5]. From this point of view, influenza virus strain A/South Africa/3626/2013 (H1N1)pdm09 (SA-WT) is noteworthy since it is highly pathogenic for lab animals (mice, ferrets). In other words, this strain has the required characteristics to be considered a model strain to infect these lab animals without the need for the adaptation process [
6]. It was previously illustrated that this strain, compared to other H1N1pdm09 influenza A virus strains, has three strain-specific substitutions in its polymerase complex (N102T in PB2, E358E/K heterogeneity in PB2, and Q687R in PB1) [
7]. The viral polymerase complex is encoded by three gene segments (PB1, PB2, PA) [
8]. Mutations in these genes have been shown to play a critical role in strain pathogenicity as the activity of the complex affects the virus adaptation to mammals [
9]. In addition, mutations in PB2 are responsible for virus sensitivity to host temperature. And thus, its ability to reproduce in lower respiratory tracks [
10,
11]. Therefore, the mutations described by us might help to explain why this strain is highly pathogenic to mice without previous adaptation. We proposed that passaging in mice will leave only the variant most capable of replicating in mice lungs, becoming the predominant population, which will give us more information about the effect of these strain-specific substitutions in virus pathogenicity.
In this work, a mouse-adapted version of SA-WT was obtained after five passages in mice (SA-M5). Subsequently, the differences in toxicity, lethality, reproduction, and immunogenicity of SA-WT and SA-M5 were discussed.
2. Results
2.1. Sequencing analysis of SA-M5
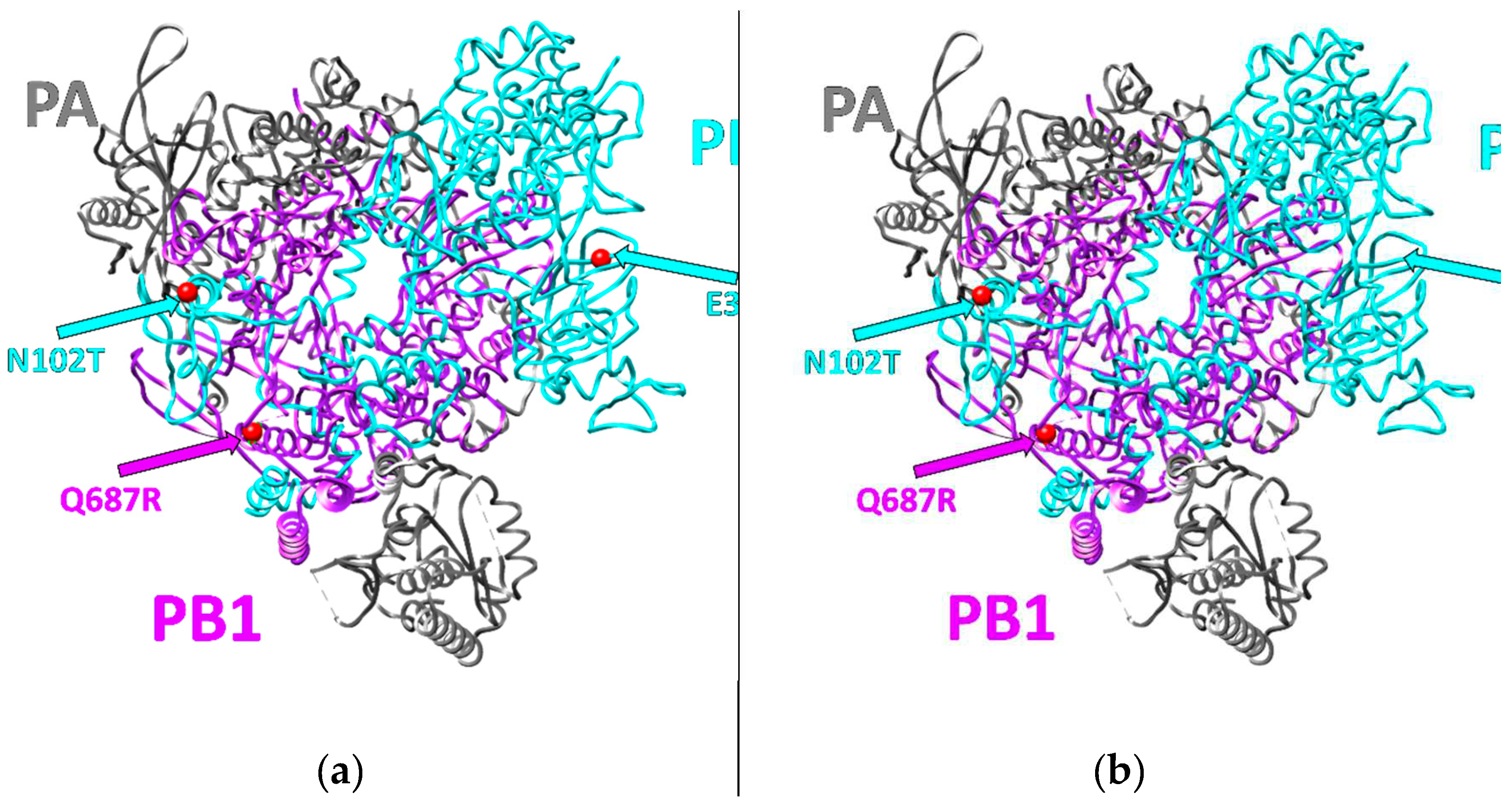
To detect the changes in the three strain-specific substitutions (N102T in PB2, E358E/K heterogeneity in PB2, and Q687R in PB1), Sequencing of PB2 and PB1 gene segments in the SA-M5 isolates was performed. Sequencing results revealed that after five sequential passages in mice, the only change that occurred was the disappearance of the A/G heterogeneity in the 1072nd position in the PB2 gene, which is responsible for E358E/K heterogeneity in PB2 protein, and the G-variant, coding for E358, was the only variant in this position in all SA-M5 virus tested isolates (
Figure 1).
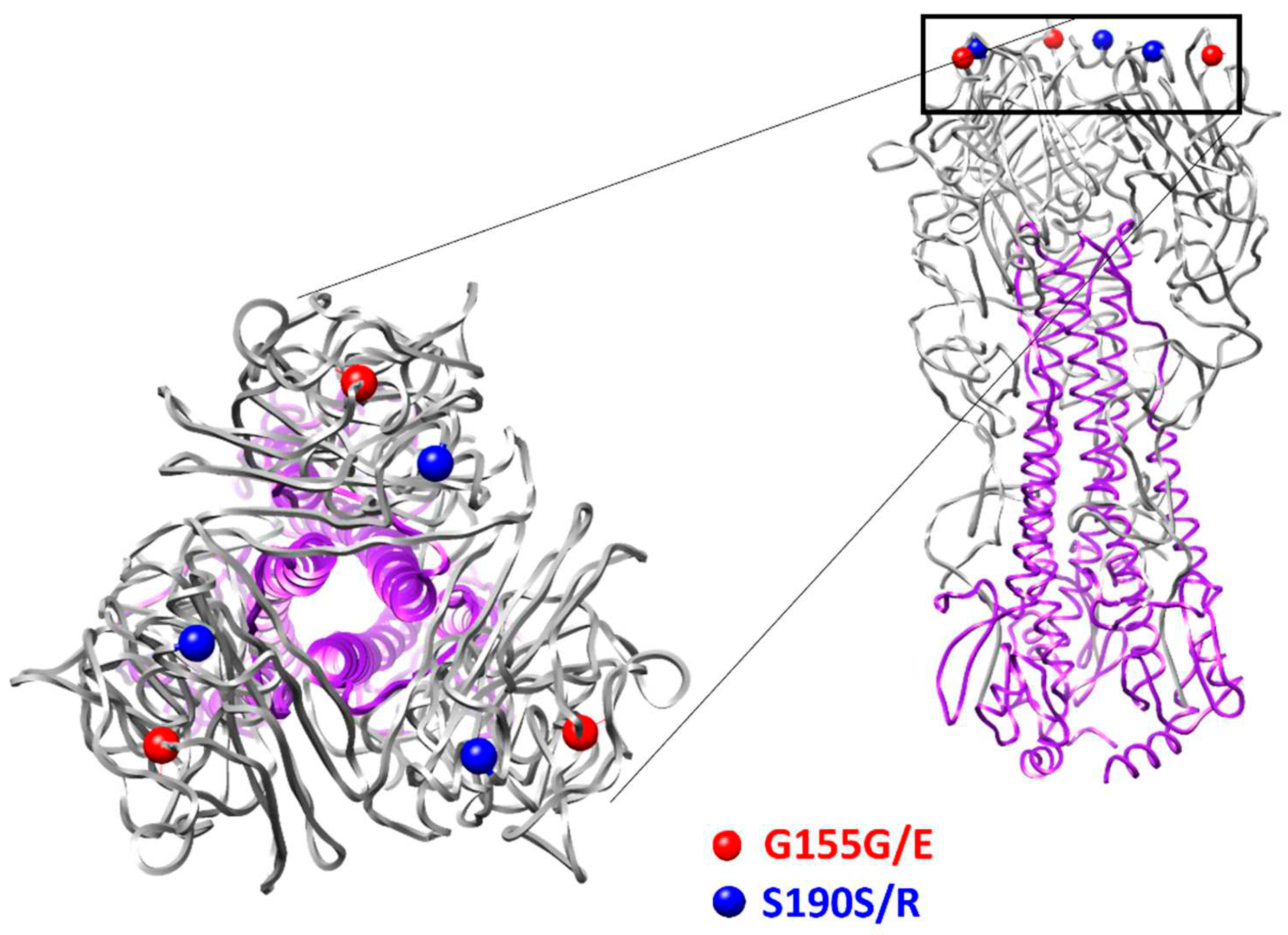
The results of full-genome sequencing of the SA-M5 revealed that SA-M5’s genome has no additional substitutions in any of its genetic segments except in the HA gene. In the HA gene, two heterogeneous positions (547R and 651M result in G155G/E variants and S190S/R, respectively) were found (
Figure 2).
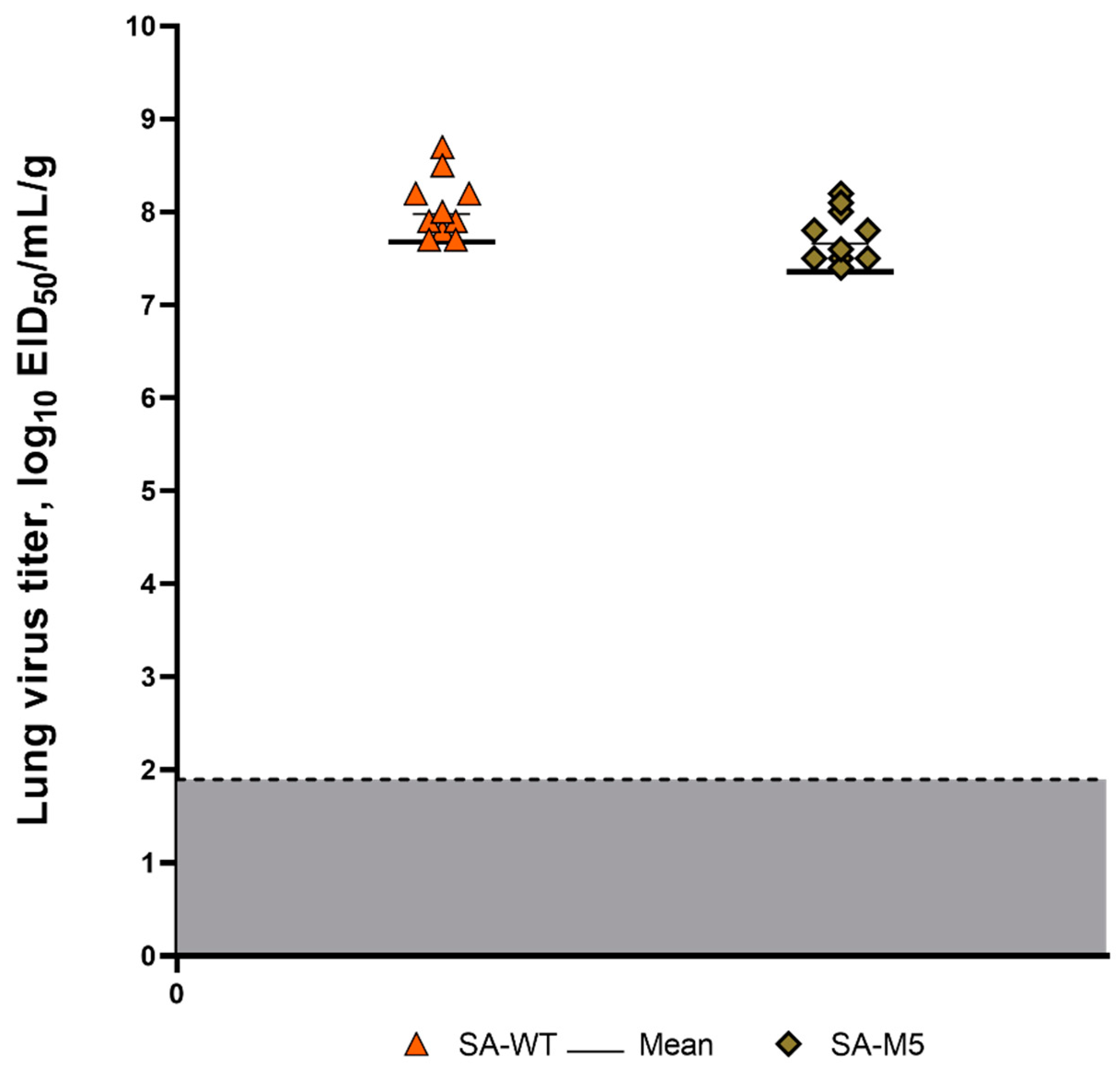
2.2. Viral replication in lung tissue
Both strains, SA-WT and SA-M5, efficiently replicated in the lower respiratory tract of the infected mice, with no significant difference between the two strains. Titration of lung homogenates showed high titers for SA-WT and SA-M5, where the average titers were 8.06 ± 0,60 log
10 EID
50/mL and 7.74 ± 0,40 log
10 EID
50/mL, respectively (
Figure 3).
2.3. SA-WT and SA-M5 toxicity in mice
To compare the toxicity of SA-WT and SA-M5, mice were inoculated intranasally with a high dose of fresh undiluted SA-WT or SA-M5 to cause lethality from acute pulmonary edema. In mice, infection with SA-WT and SA-M5 resulted in 80% and 70% lethality, respectively, within the first six days after the inoculation with the viruses. A statistical difference between the two strains was not observed (
Figure 4).
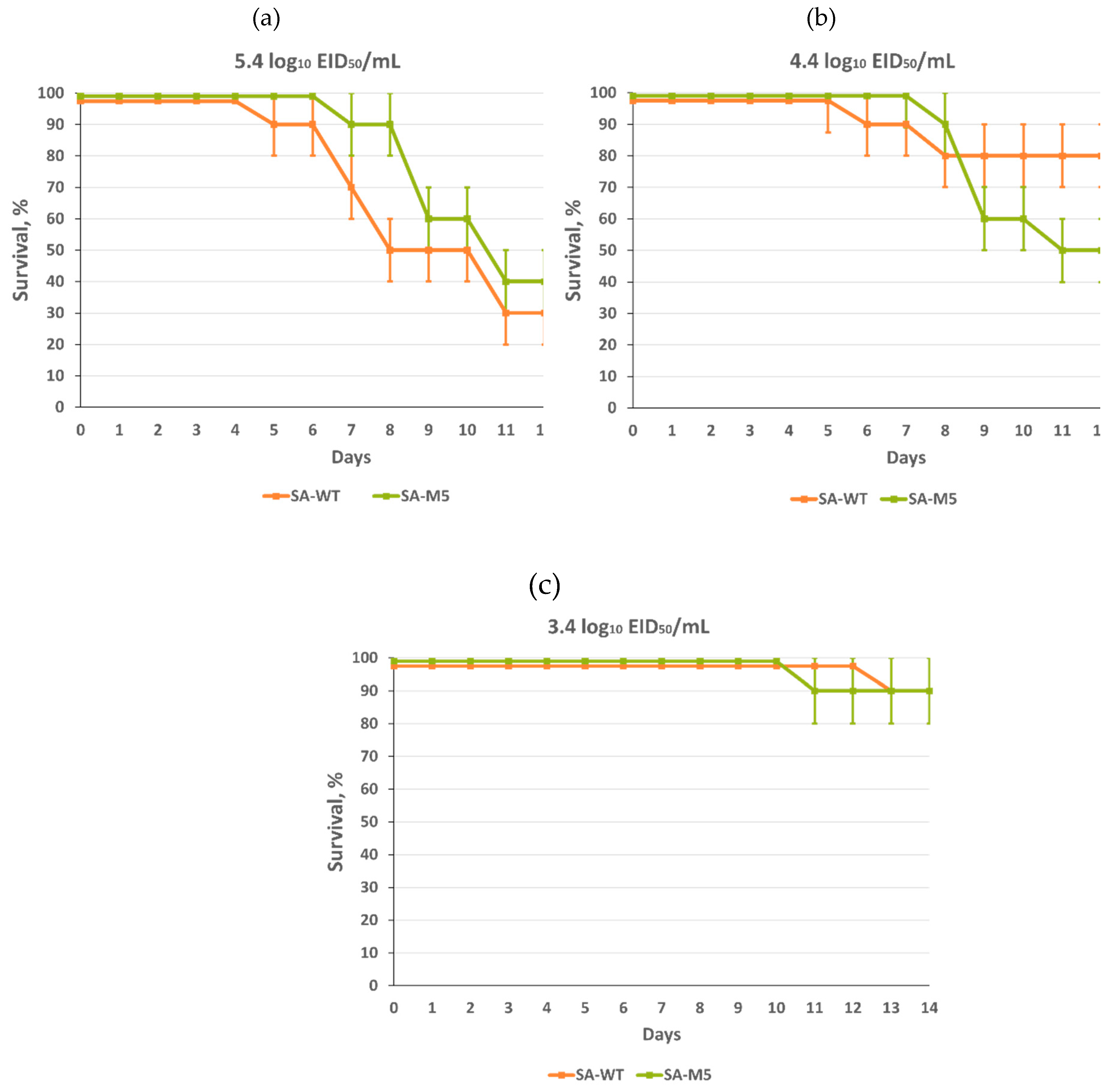
2.4. SA-WT and SA-M5 pathogenicity in mice
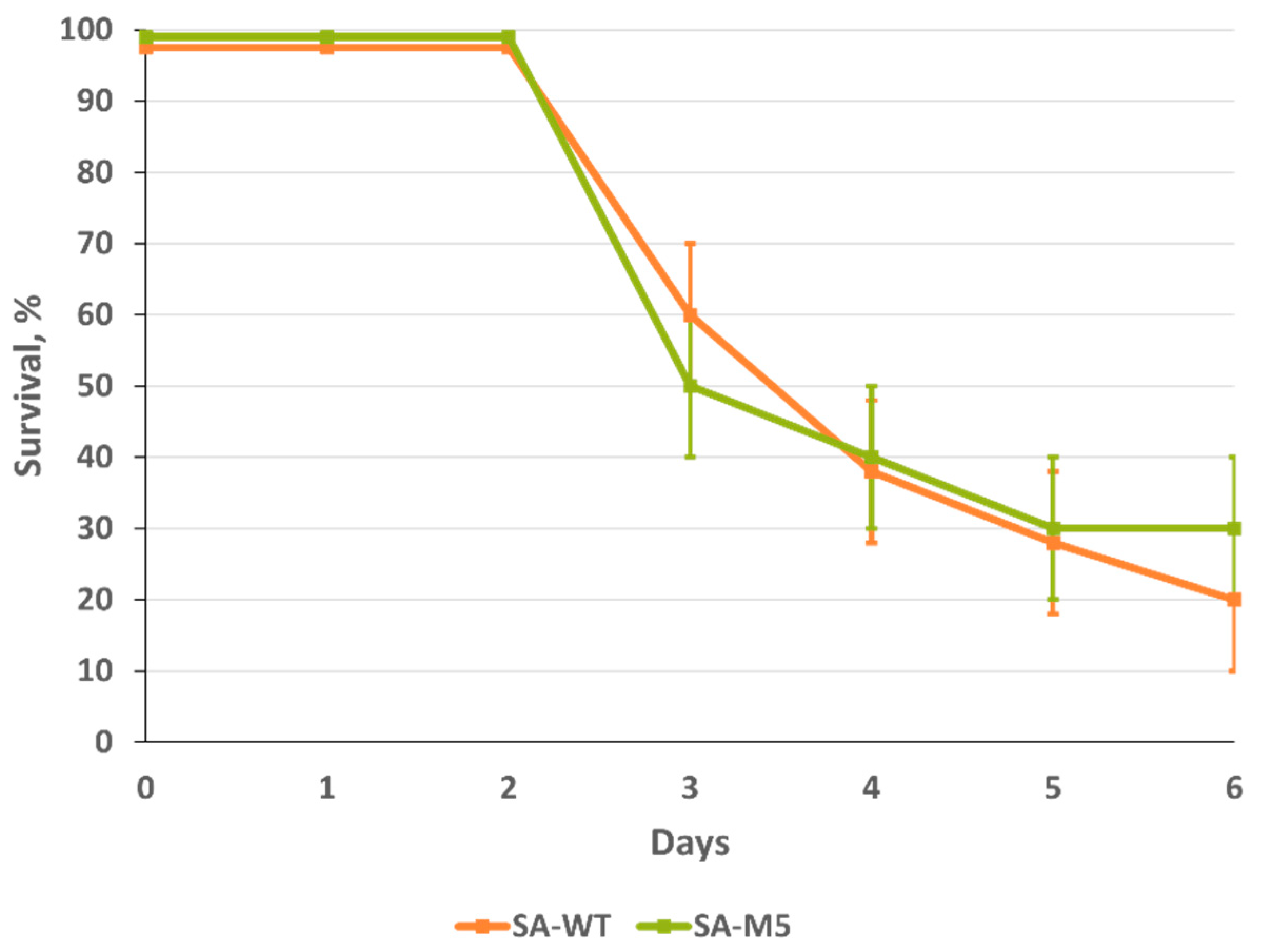
Survival study showed that both strains were highly pathogenic for mice with no significant difference between them, as mice lethality at the end of the experiment after intranasal inoculation with 5.4 log
10 EID
50/mL was 70% and 60% for SA-WT and SA-M5, respectively. Intranasal inoculation with 4.4 log
10 EID
50/mL caused 50% lethality for both strains. Even intranasal inoculation with 3.4 log
10 EID
50/mL caused 10% lethality for both strains (
Figure 5). In addition, LD
50 calculated by Reed and Muench method [
19] for both strains were almost identical as 1 LD
50 for SA-WT and 1 LD
50 for SA-M5 were 4.60 log
10 EID
50/mL and 4.65 log
10 EID
50/mL, respectively.
3.5. SA-WT and SA-M5 immunogenicity
To detect the serum antibody levels against SA-WT and SA-M5, a two-way hemagglutination inhibition test (HAI) was performed, where all sera were tested against SA-WT, and the test was also repeated against SA-M5, using the sera obtained from mice who survived until the 21st day of the experiment. The values of the two-way HAI test were identical for both strains (
Table S1). Interestingly, data showed that HAI titers of SA-M5 sera, after mice inoculation with a series of infection doses, were significantly higher than titers of SA-WT sera (
Figure 6).
3. Discussion
The presented work is a continuation of our investigations devoted to the analysis of biological properties of the influenza virus strain A/South Africa/3626/2013 (H1N1)pdm09 (SA-WT), which is a convenient model for laboratory studies of the mechanisms of influenza infection but remains unstudied. SA-WT strain has three specific substitutions in the polymerase complex (N102T in PB2, E358E/K heterogeneity also in PB2, and Q687R in PB1) [
7]. Q687R localized in the C-terminus of PB1 that forms a strong bond with PB2 through its N-terminus. Hence, Q687R in PB1 might affect the polymerase activity[
7,
12]. N102T in PB2 also might affect polymerase stability as the amino acid in this position is part of the PB2 subdomain that supports the PB1 thumb domain [
13]. As for E358E/K heterogeneity in PB2, our results correspond with literature data that this position is highly conservative in influenza subtypes that can infect humans and animals [
14]. The 358th amino acid position in PB2 is part of the cap-binding domain; therefore, a mutation in this position might alter polymerase activity, affecting the strain’s pathogenicity [
13]. Thus, each of the three substitutions of the polymerase complex may be important for the formation of the high pathogenicity of this strain.
The current study aimed to understand whether these substitutions play a role in the virus pathogenicity of SA-WT. To investigate whether these substitutions will disappear during the adaptation process and, if so, how this will affect the viral pathogenicity, strain SA-WT was passaged in mice five times. After that, the predominant population in SA-M5 kept N102T in PB2 and Q687R in and lost the heterogeneity E358E/K in PB2 as SA-M5 contains only glutamic acid in the 358th position (E358) (
Figure 1). However, further investigations showed no changes in pathogenicity as SA-M5 has the same level of pathogenicity as SA-WT, where toxicity study showed 70% and 80% lethality for SA-M5 and SA-WT respectively, with no statistical difference (
Figure 4 and
Figure 5). The same observation was in their ability to replicate in the lower respiratory tract with average titers 8.06 ± 0,60 log
10 EID
50/mL versus 7.74 ± 0,40 log
10 EID
50/mL for SA-WT versus SA-M5 (
Figure 3). In addition, both strains have almost the same 50% lethal dose (LD
50) - 4.60 log
10 EID
50/mL for SA-WT and 4.65 log
10 EID
50/mL for SA-M5. These data support our previous finding that N102T and Q687R substitutions might be the molecular basis that explains the viral pathogenicity in mice [
7]. In contrast, E358E/K heterogeneity in PB2 does not play a crucial role in strain pathogenicity as its disappearance did not affect the strain pathogenic characteristics.
As for HA protein, it contains the binding site to sialic acid residues on the cellular membrane of the host cell and plays a remarkable role in the fusion process in the cytoplasm. In addition, it has a significant role in immunogenicity because it is the most abundant glycoprotein on influenza envelop [
15]. One of the primary viral strategies to avoid host response is mutations in the antigenic sites in HA [
16]. These mutations have always drowned the attention because they are the main reason that vaccines for influenza are changed annually. Unexpectedly, passaging in mice led to the appearance of the mutations in hemagglutinin (HA) gene. Two heterogeneities (G155G/E and S190S/R) were identified additionally (
Figure 2).
SA-WT and SA-M5 differ in their HAI titers, which indicates that the immune response to SA-M5 was significantly higher than against SA-WT. This difference can be explained by the two heterogeneities (G155G/E and S190S/R) in the HA of SA-M5. At the amino acid position 155 in the HAs of H1N1pdm09 viruses, G residue is the common residue, while E residue is the rare one in the natural population. This position is also a part of antigenic site A [
17]. The 155E variant had increased replication ability in epithelial cells (Calu-3 culture). The increased growth capacity in epithelial cells could be the cause of the increased immunogenicity of this variant in mice models. It is interesting that in the two-way HAI test, the titers with different antigens were equal, and possibly, this was caused by the heterogeneity of the virus population. The 190 position is a part of the receptor binding pocket of HA. In addition, the R variant is usually a result of egg adaptation. The mechanism of egg adaptation was illustrated early [
18,
19]. It was shown that the S190R variant of the A/Hunan/26/2016 (H1N1)pdm09 virus had increased pathogenicity for mice [
18]. However, in our study, the presence of this component in the population of the virus didn’t increase its pathogenicity (
Figure 5).
The SA-M5 variant of the SA-WT is perhaps even more attractive as a model virus than SA-WT itself for studying various aspects of the pathogenesis of influenza infection when screening anti-influenza chemotherapy drugs, etc. due to its high immunogenicity.
4. Materials and Methods
4.1. Viruses
Two virus strains were used in this study. 1 – egg-derived A/South Africa/3626/2013 (H1N1)pdm09 (SA-WT): influenza virus strain A/South Africa/3626/2013 (H1N1)pdm09 was obtained from The Francis Crick Institute (London, UK) ID# 2014701384. 2 - A/South Africa/3626/2013 (H1N1)pdm09 E1E2/M5 (SA-M5): was obtained from SA-WT strain after the 5 passages in mice (see below section “Virus adaptation to mice”)
4.2. Mice
Female CBA mice, aged 8-12 weeks, the animals were kept in polycarbonate cages with free access to food and water.
4.3. Virus adaptation to mice
Five mice were lightly anesthetized with ether and inoculated intranasally with 50 μL of PBS containing 4.8 log10 EID50 of influenza SA-WT, divided equally between the nostrils. On the third day after the infection, mice were sacrificed then, their lungs were harvested in sterile conditions, after which they were homogenized using a small bead mill Tissue Lyser LT (QIAGEN, Germany) in 1.0 mL of PBS containing antibiotic-antimycotic (Invitrogen, UK), a pool from the homogenates was prepared to perform the second passage of mice. After that, the pool was diluted 10-fold, and each mouse was inoculated with 50 μL of the diluted mixture. This process was repeated until the homogenized lungs from the fifth passage were obtained.
4.4. Viral replication in lung tissue
For each of the two strains, SA-WT and SA-M5, respectively, a group of 10 mice was inoculated with the virus, and on the third day after the infection, mice were sacrificed, and their lungs were harvested and homogenized in PBS. Homogenates were titrated in 10-11-day-old chicken embryos supplied by the «Sinyavino» poultry farm (Kirovsk Area, Leningrad region, Russia). Eggs were incubated for 72 hours at 32°C. The log
10 EID
50/mL calculation was based on the Reed and Muench method [
20].
4.5. Viral genome sequencing
Samples were homogenized using a T10 basic ULTRA-TURRAX disintegrator (IKA, Germany) and the total RNA was extracted and purified from homogenates of mice organs by Rizol reagent (Diaem, Russia). Viral RNA from chorioallantoic fluid was extracted by precipitation method using a Ribo-prep kit (AmpliSens, Moscow, Russia). RT-PCR was performed with a Biomaser RT-PCR kit (Biolabmix, Novosibirsk, Russia) with electrophoretic detection of products and purification from the gel with a Clean-Up column kit (Evrogen, Moscow, Russia). The viral genome sequencing was performed with Applied Biosystems 3130xl Genetic Analyzer according to the manufacturer’s recommendations using BigDye™ Terminator v3.1 Cycle Sequencing Kit (Thermo Fischer Scientific, USA). The primers used for RT-PCR reaction and subsequent sequencing are listed in
Table S1.
4.6. Toxicity in mice
Mice were inoculated with a high dose of fresh, undiluted virus under anesthesia with ether, to study the virus's ability to cause acute pulmonary edema which is also known as the viral toxic effect, after which they were observed daily during the first six days post-infection for detecting lethality from acute pulmonary edema as previously described in [
6].
4.7. Pathogenicity in mice
A Series of 10-fold dilutions was prepared in PBS for the model strain SA-WT and the strain SA-M5 obtained after five passages in mice. Using 10 mice per group, mice were inoculated intranasally under ether anesthesia by 50 μL of the previously prepared series. For the first 14 days post-infection mice lethality was observed, and the LD
50 for each virus was calculated by the Reed and Muench method [
20] and expressed as the log
10 EID
50/mL required to give 1 LD
50.
4.8. Hemagglutination inhibition test
A Standard hemagglutination inhibition test was performed against four hemagglutination units 4 HAU for both strains using the sera obtained from mice who survived until the 21st day of the experiment. On the 21st day after the infection, mice were sacrificed, and the blood serum was collected from each mouse separately. Subsequently, the sera were treated with RDE (Denka, Japan) and incubated for 1 hour at 56°C to inactivate the nonspecific hemagglutination inhibitors in the serum, after which the sera were incubated with 4% erythrocytes-PBS solution overnight at 4°C to avoid nonspecific hemagglutination. Before incubation with viruses, sera were incubated for 1 hour at 56°C again and left to cool at room temperature. For each serum, a series of 2-fold dilutions were prepared using PBS containing either SA-WT 4 HAU or SA-M5 4 HAU, then 1% erythrocytes-PBS solution was added, subsequently, the agglutination of erythrocytes was observed, and hemagglutination inhibition titers were calculated accordingly [
21,
22].
4.9. 3D structure
The UCSF Chimera 1.15 program was used to build the 3D structure. The 3D samples were obtained from Protein Data Bank PDB.
4.10. Ethics statement
All of the experiments on mice and chicken embryos were performed according to European Union legislation [
23], and the study was approved by the Institutional Local Ethical Committee (IEM, St Petersburg, Russia). After each experiment, animals were humanely euthanized.
4.11. Statistics
GraphPad Prizm 8 program was used to perform statistical analysis. A p-value < 0.05 was considered statistically significant.
5. Conclusions
The adapted version (SA-M5) that was obtained after five passages in mice of influenza A virus A/South Africa/3626/2013 (H1N1)pdm09 (SA-WT) lost one of the three strain-specific substitutions, namely E358E/K heterogeneity in PB2, without any change in its pathogenicity. These findings support the conclusion that E358E/K in PB2 does not play an essential role in the virus's pathogenicity. Additionally, the increase in the virus's immunogenicity, which was noticed in the adapted version, could be explained by the two heterogeneities (G155G/E and S190S/R) that appeared in the HA. In light of these findings, SA-M5 is a more favorable model strain than SA-WT as its higher pathogenicity could be a preferred factor when screening anti-influenza chemotherapy drugs due to its high immunogenicity.
Supplementary Materials
The following supporting information can be downloaded at:
www.mdpi.com/xxx/s1, Table S1: Oligonucleotides used for full-genome sequencing of A/South Africa/3626/2013 virus genome.; Table S2: Two-way hemagglutination inhibition test.
Author Contributions
Conceptualization, I.K., L.P., and L.R.; methodology, A.R., E.S., and M.F.; validation, I.K., E.S., and M.F.; formal analysis, E.S. and M.F.; investigation, A.C., E.B., E.K., A.T., and M.F.; resources, L.P. and L.R.; data curation, E.S. and M.F.; writing—original draft preparation, E.S. and M.F.; writing—review and editing, L.P., I.K., and M.F.; visualization, M.F.; supervision, I.K. and L.P.; project administration, L.R. All authors have read and agreed to the published version of the manuscript.
Funding
The manuscript was funded by the budget of the Institute of Experimental Medicine (State registration number of research work AAAA-A19-119021490149-7) and received no external funding.
Institutional Review Board Statement
The animal study protocol was approved by the Ethics Committee of the Institutional Local Ethical Committee (IEM, St Petersburg, Russia) (1/20 was approved on 27 February 2020).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Estimating Disease Burden of Influenza Available online: https://www.who.int/europe/activities/estimating-disease-burden-of-influenza (accessed on 22 October 2023).
- Shao, W.; Li, X.; Goraya, M.U.; Wang, S.; Chen, J.-L. Evolution of Influenza A Virus by Mutation and Re-Assortment. Int. J. Mol. Sci. 2017, 18, 1650. [CrossRef]
- Tosh, P.K.; Jacobson, R.M.; Poland, G.A. Influenza Vaccines: From Surveillance Through Production to Protection. Mayo Clin. Proc. 2010, 85, 257–273. [CrossRef]
- Lampejo, T. Influenza and Antiviral Resistance: An Overview. Eur. J. Clin. Microbiol. Infect. Dis. Off. Publ. Eur. Soc. Clin. Microbiol. 2020, 39, 1201–1208. [CrossRef]
- Dadonaite, B.; Gilbertson, B.; Knight, M.L.; Trifkovic, S.; Rockman, S.; Laederach, A.; Brown, L.E.; Fodor, E.; Bauer, D.L.V. The Structure of the Influenza A Virus Genome. Nat. Microbiol. 2019, 4, 1781–1789. [CrossRef]
- Kiseleva, I.; Rekstin, A.; Al Farroukh, M.; Bazhenova, E.; Katelnikova, A.; Puchkova, L.; Rudenko, L. Non-Mouse-Adapted H1N1pdm09 Virus as a Model for Influenza Research. Viruses 2020, 12, 590. [CrossRef]
- Al Farroukh, M.; Kiseleva, I.; Bazhenova, E.; Stepanova, E.; Puchkova, L.; Rudenko, L. Understanding the Variability of Certain Biological Properties of H1N1pdm09 Influenza Viruses. Vaccines 2022, 10, 395. [CrossRef]
- Bouvier, N.M.; Palese, P. THE BIOLOGY OF INFLUENZA VIRUSES. Vaccine 2008, 26, D49–D53.
- Cox, N.J.; Kitame, F.; Kendal, A.P.; Maassab, H.F.; Naeve, C. Identification of Sequence Changes in the Cold-Adapted, Live Attenuated Influenza Vaccine Strain, A/Ann Arbor/6/60 (H2N2). Virology 1988, 167, 554–567. [CrossRef]
- Massin, P.; van der Werf, S.; Naffakh, N. Residue 627 of PB2 Is a Determinant of Cold Sensitivity in RNA Replication of Avian Influenza Viruses. J. Virol. 2001, 75, 5398–5404. [CrossRef]
- McCauley, J.W.; Penn, C.R. The Critical Cut-off Temperature of Avian Influenza Viruses. Virus Res. 1990, 17, 191–198. [CrossRef]
- Sugiyama, K.; Obayashi, E.; Kawaguchi, A.; Suzuki, Y.; Tame, J.R.H.; Nagata, K.; Park, S.-Y. Structural Insight into the Essential PB1–PB2 Subunit Contact of the Influenza Virus RNA Polymerase. EMBO J. 2009, 28, 1803–1811. [CrossRef]
- Pflug, A.; Guilligay, D.; Reich, S.; Cusack, S. Structure of Influenza A Polymerase Bound to the Viral RNA Promoter. Nature 2014, 516, 355–360. [CrossRef]
- Isakova-Sivak, I.; Stepanova, E.; Mezhenskaya, D.; Matyushenko, V.; Prokopenko, P.; Sychev, I.; Wong, P.-F.; Rudenko, L. Influenza Vaccine: Progress in a Vaccine That Elicits a Broad Immune Response. Expert Rev. Vaccines 2021, 20, 1097–1112. [CrossRef]
- Sriwilaijaroen, N.; Suzuki, Y. Molecular Basis of the Structure and Function of H1 Hemagglutinin of Influenza Virus. Proc. Jpn. Acad. Ser. B 2012, 88, 226–249. [CrossRef]
- Zhu, R.; Xu, S.; Sun, W.; Li, Q.; Wang, S.; Shi, H.; Liu, X. HA Gene Amino Acid Mutations Contribute to Antigenic Variation and Immune Escape of H9N2 Influenza Virus. Vet. Res. 2022, 53, 43. [CrossRef]
- Ilyushina, N.A.; Komatsu, T.E.; Ince, W.L.; Donaldson, E.F.; Lee, N.; O’Rear, J.J.; Donnelly, R.P. Influenza A Virus Hemagglutinin Mutations Associated with Use of Neuraminidase Inhibitors Correlate with Decreased Inhibition by Anti-Influenza Antibodies. Virol. J. 2019, 16, 149. [CrossRef]
- Chen, Y.; Bai, T.; Zhu, W.; Gao, R.; Deng, Z.; Shi, Y.; Zou, S.; Huang, Y.; Li, X.; Li, F.; et al. The S190R Mutation in the Hemagglutinin Protein of Pandemic H1N1 2009 Influenza Virus Increased Its Pathogenicity in Mice. Sci. China Life Sci. 2018, 61, 836–843. [CrossRef]
- Nicolson, C.; Harvey, R.; Engelhardt, O.G.; Robertson, J.S. The Ability of a Non-Egg Adapted (Cell-Like) A(H1N1)Pdm09 Virus to Egg-Adapt at HA Loci Other than 222 and 223 and Its Effect on the Yield of Viral Protein. PLOS ONE 2016, 11, e0166761. [CrossRef]
- REED, L.J.; MUENCH, H. A SIMPLE METHOD OF ESTIMATING FIFTY PER CENT ENDPOINTS12. Am. J. Epidemiol. 1938, 27, 493–497. [CrossRef]
- Manual for the Laboratory Diagnosis and Virological Surveillance of Influenza Available online: https://www.who.int/publications-detail-redirect/manual-for-the-laboratory-diagnosis-and-virological-surveillance-of-influenza (accessed on 22 October 2023).
- Klimov, A.; Balish, A.; Veguilla, V.; Sun, H.; Schiffer, J.; Lu, X.; Katz, J.M.; Hancock, K. Influenza Virus Titration, Antigenic Characterization, and Serological Methods for Antibody Detection. In Influenza Virus: Methods and Protocols; Kawaoka, Y., Neumann, G., Eds.; Methods in Molecular Biology; Humana Press: Totowa, NJ, 2012; pp. 25–51 ISBN 978-1-61779-621-0.
-
Directive 2010/63/EU of the European Parliament and of the Council of 22 September 2010 on the Protection of Animals Used for Scientific Purposes Text with EEA Relevance; 2010; Vol. 276;.
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).