Submitted:
15 January 2024
Posted:
16 January 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Strains and plasmids
2.2. Biosynthetic pathway construction
2.3. Flask fermentation conditions
2.4. Analytical methods
2.5. Model construction
2.6. Conditions for in silico simulations and optimization
3. Results
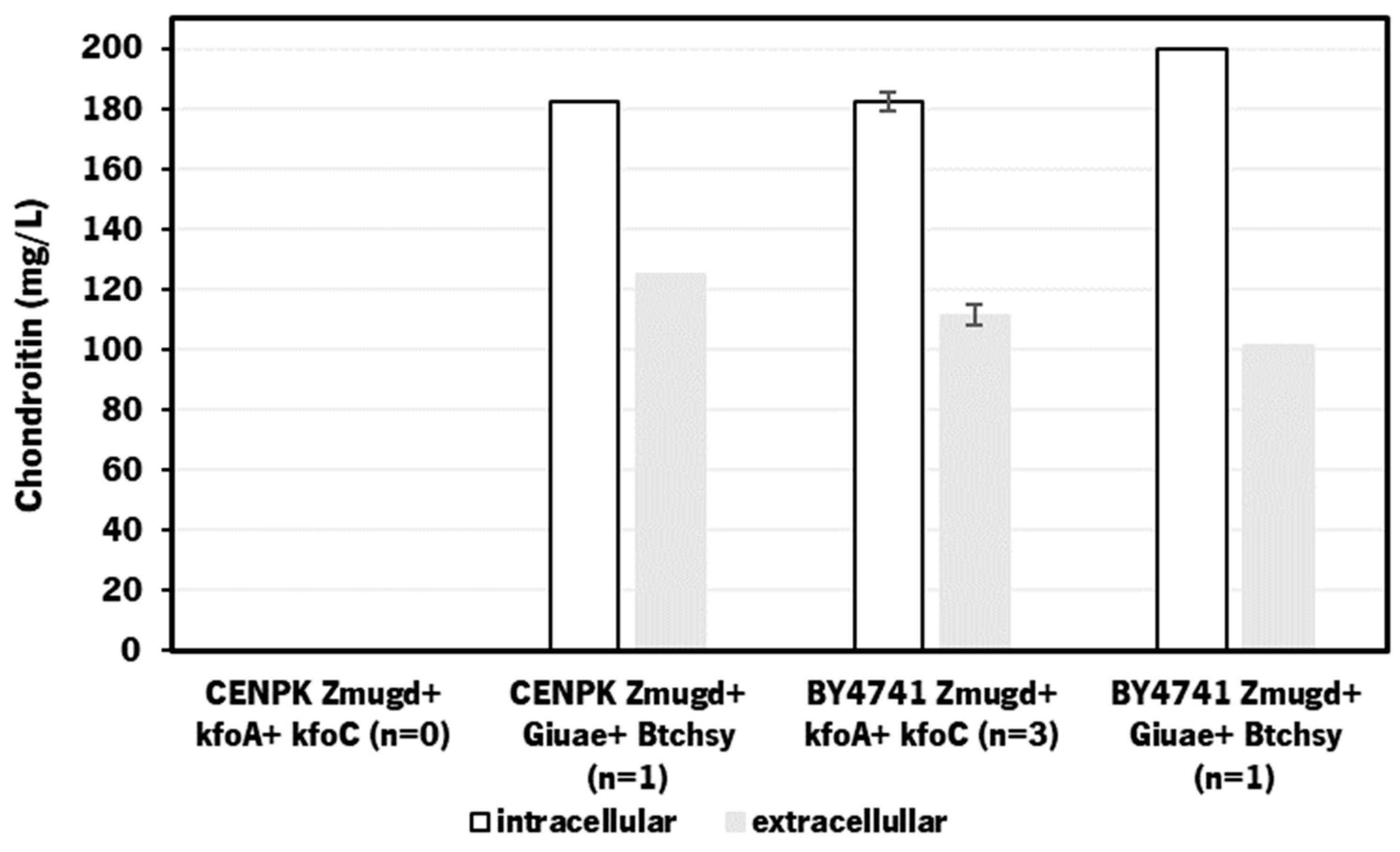
3.1. Heterologous production of chondroitin in S. cerevisae
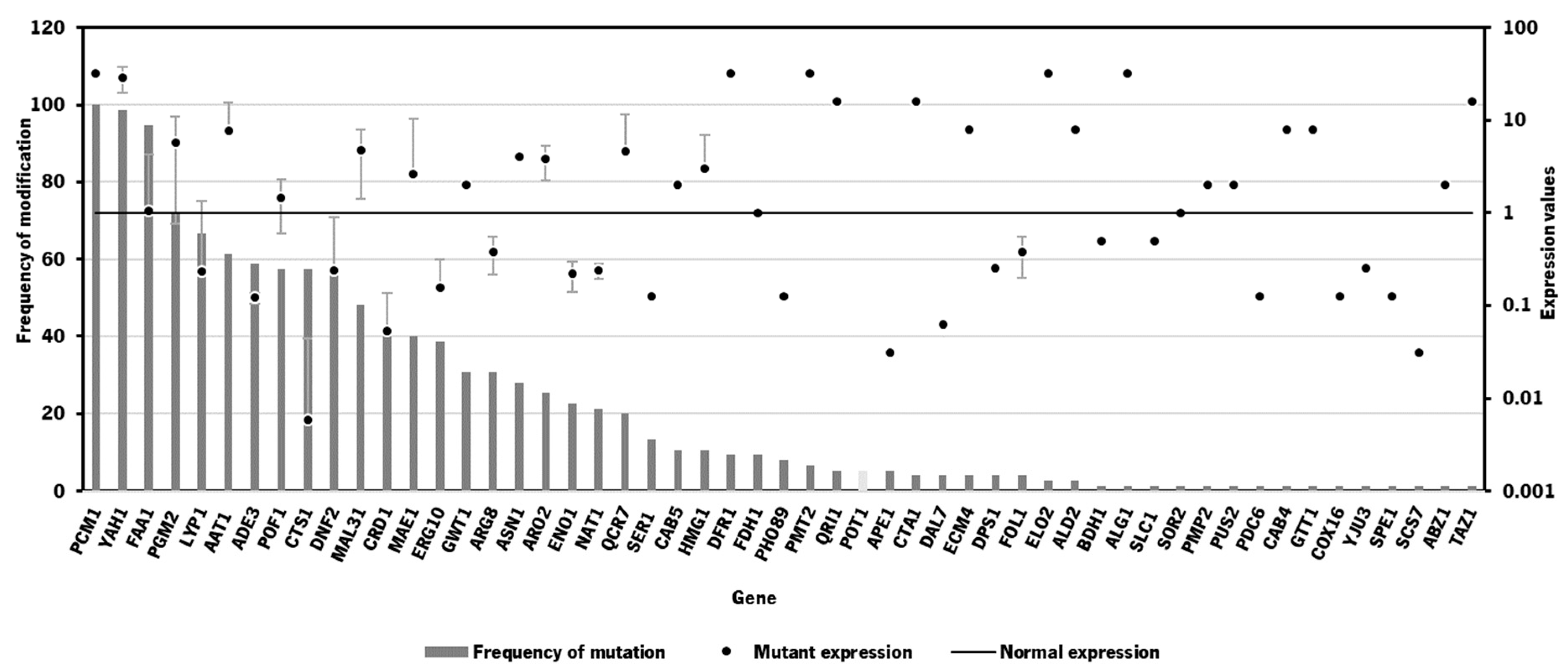
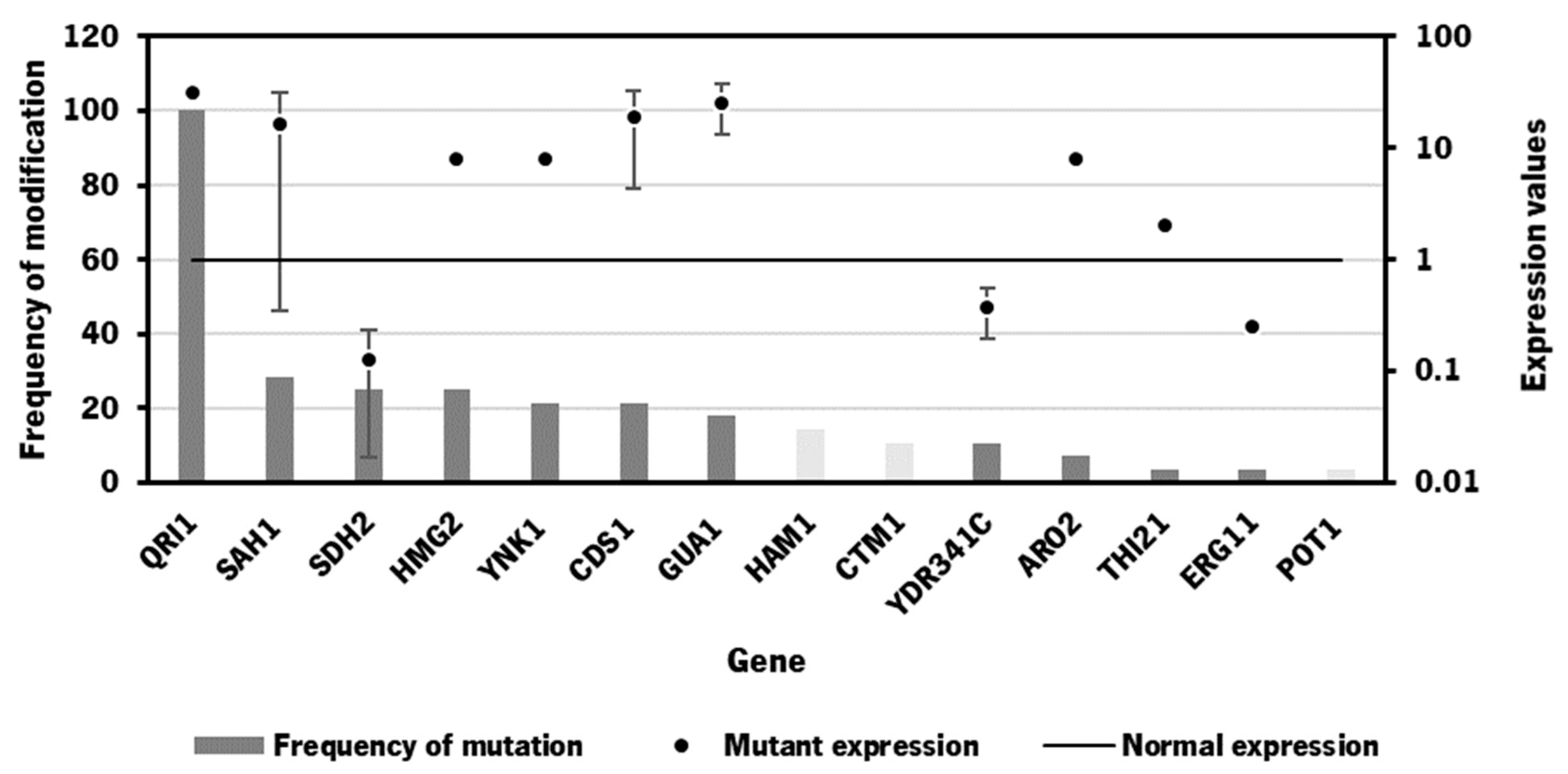
3.2. Bioinformatics tool for identification of gene targets
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Couto, M.R.; Rodrigues, J.L.; Rodrigues, L.R. Heterologous production of chondroitin. Biotechnol. Reports 2022, 33, e00710. [Google Scholar] [CrossRef]
- Cress, B.F.; Englaender, J.A.; He, W.; Kasper, D.; Linhardt, R.J.; Koffas, M.A.G. Masquerading microbial pathogens: Capsular polysaccharides mimic host-tissue molecules. FEMS Microbiol. Rev. 2014, 38, 660–697. [Google Scholar] [CrossRef]
- Cimini, D.; Carlino, E.; Giovane, A.; Argenzio, O.; Dello Iacono, I.; De Rosa, M.; Schiraldi, C. Engineering a branch of the UDP-precursor biosynthesis pathway enhances the production of capsular polysaccharide in Escherichia coli O5:K4:H4. Biotechnol. J. 2015, 10, 1307–1315. [Google Scholar] [CrossRef]
- Jin, P.; Zhang, L.; Yuan, P.; Kang, Z.; Du, G.; Chen, J. Efficient biosynthesis of polysaccharides chondroitin and heparosan by metabolically engineered Bacillus subtilis. Carbohydr. Polym. 2016, 140, 424–432. [Google Scholar] [CrossRef] [PubMed]
- He, W.; Fu, L.; Li, G.; Andrew Jones, J.; Linhardt, R.J.; Koffas, M.; Jones, J.A.; Linhardt, R.J.; Koffas, M.; Andrew Jones, J.; et al. Production of chondroitin in metabolically engineered E. coli. Metab. Eng. 2015, 27, 92–100. [Google Scholar] [CrossRef]
- Cheng, F.; Luozhong, S.; Yu, H.; Guo, Z. Biosynthesis of chondroitin in engineered Corynebacterium glutamicum. J. Microbiol. Biotechnol. 2019, 29, 392–400. [Google Scholar] [CrossRef]
- Zhou, Z.; Li, Q.; Huang, H.; Wang, H.; Wang, Y.; Du, G.; Chen, J.; Kang, Z. A microbial–enzymatic strategy for producing chondroitin sulfate glycosaminoglycans. Biotechnol. Bioeng. 2018, 115, 1561–1570. [Google Scholar] [CrossRef]
- Jin, X.; Zhang, W.; Wang, Y.; Sheng, J.; Xu, R.; Li, J.; Du, G.; Kang, Z. Biosynthesis of non-animal chondroitin sulfate from methanol using genetically engineered Pichia pastoris. Green Chem. 2021, 23, 4365–4374. [Google Scholar] [CrossRef]
- Desko, M.M.; Gross, D.A.; Kohler, J.J. Effects of N-glycosylation on the activity and localization of GlcNAc-6-sulfotransferase 1. Glycobiology 2009, 19, 1068–1077. [Google Scholar] [CrossRef]
- Badri, A.; Williams, A.; Awofiranye, A.; Datta, P.; Xia, K.; He, W.; Fraser, K.; Dordick, J.S.; Linhardt, R.J.; Koffas, M.A.G.G. Complete biosynthesis of a sulfated chondroitin in Escherichia coli. Nat. Commun. 2021, 12, 1–10. [Google Scholar] [CrossRef]
- Nielsen, J. Yeast Systems Biology: Model Organism and Cell Factory. Biotechnol. J. 2019, 14, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Borodina, I. Application of synthetic biology for production of chemicals in yeast Saccharomyces cerevisiae. FEMS Yeast Res. 2015, 15, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.K.; Shukla, P. Sophisticated cloning, fermentation, and purification technologies for an enhanced therapeutic protein production: A review. Front. Pharmacol. 2017, 8, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Lopes, H.; Rocha, I. Genome-scale modeling of yeast: chronology, applications and critical perspectives. FEMS Yeast Res. 2017, 17, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Vieira, V.; Maia, P.; Rocha, M.; Rocha, I. Comparison of pathway analysis and constraint-based methods for cell factory design. BMC Bioinformatics 2019, 20, 1–15. [Google Scholar] [CrossRef]
- Maia, P.; Rocha, M.; Rocha, I. In silico constraint-based strain optimization methods: the quest for optimal cell factories. Microbiol. Mol. Biol. Rev. 2016, 80, 45–67. [Google Scholar] [CrossRef]
- Bi, X.; Liu, Y.; Li, J.; Du, G.; Lv, X.; Liu, L. Construction of multiscale genome-scale metabolic models: frameworks and challenges. Biomolecules 2022, 12, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Ng, R.H.; Lee, J.W.; Baloni, P.; Diener, C.; Heath, J.R.; Su, Y. Constraint-based reconstruction and analyses of metabolic models: open-source Python tools and applications to cancer. Front. Oncol. 2022, 12, 1–23. [Google Scholar] [CrossRef]
- Mardinoglu, A.; Nielsen, J. Systems medicine and metabolic modelling. J. Intern. Med. 2012, 271, 142–154. [Google Scholar] [CrossRef]
- Zhang, C.; Hua, Q. Applications of genome-scale metabolic models in biotechnology and systems medicine. Front. Physiol. 2016, 6, 1–8. [Google Scholar] [CrossRef]
- Fang, X.; Lloyd, C.J.; Palsson, B.O. Reconstructing organisms in silico: genome-scale models and their emerging applications. Nat. Rev. Microbiol. 2020, 18, 731–743. [Google Scholar] [CrossRef] [PubMed]
- Gu, C.; Kim, G.B.; Kim, W.J.; Kim, H.U.; Lee, S.Y. Current status and applications of genome-scale metabolic models. Genome Biol. 2019, 20, 1–18. [Google Scholar] [CrossRef]
- Chen, X.; Xu, G.; Xu, N.; Zou, W.; Zhu, P.; Liu, L.; Chen, J. Metabolic engineering of Torulopsis glabrata for malate production. Metab. Eng. 2013, 19, 10–16. [Google Scholar] [CrossRef]
- Xu, G.; Zou, W.; Chen, X.; Xu, N.; Liu, L.; Chen, J. Fumaric acid production in Saccharomyces cerevisiae by in silico aided metabolic engineering. PLoS One 2012, 7, 1–10. [Google Scholar] [CrossRef]
- Otero, J.M.; Cimini, D.; Patil, K.R.; Poulsen, S.G.; Olsson, L.; Nielsen, J. Industrial systems biology of Saccharomyces cerevisiae enables novel succinic acid cell factory. PLoS One 2013, 8, 1–10. [Google Scholar] [CrossRef]
- Song, C.W.; Kim, D.I.; Choi, S.; Jang, J.W.; Lee, S.Y. Metabolic engineering of Escherichia coli for the production of fumaric acid. Biotechnol. Bioeng. 2013, 110, 2025–2034. [Google Scholar] [CrossRef]
- Chen, X.; Wu, J.; Song, W.; Zhang, L.; Wang, H.; Liu, L. Fumaric acid production by Torulopsis glabrata: Engineering the urea cycle and the purine nucleotide cycle. Biotechnol. Bioeng. 2015, 112, 156–167. [Google Scholar] [CrossRef]
- Harder, B.J.; Bettenbrock, K.; Klamt, S. Model-based metabolic engineering enables high yield itaconic acid production by Escherichia coli. Metab. Eng. 2016, 38, 29–37. [Google Scholar] [CrossRef]
- Mishra, P.; Lee, N.-R.; Lakshmanan, M.; Kim, M.; Kim, B.-G.; Lee, D.-Y. Genome-scale model-driven strain design for dicarboxylic acid production in Yarrowia lipolytica. BMC Syst. Biol. 2018, 12, 12. [Google Scholar] [CrossRef]
- Bro, C.; Regenberg, B.; Förster, J.; Nielsen, J. In silico aided metabolic engineering of Saccharomyces cerevisiae for improved bioethanol production. Metab. Eng. 2006, 8, 102–111. [Google Scholar] [CrossRef]
- Yim, H.; Haselbeck, R.; Niu, W.; Pujol-Baxley, C.; Burgard, A.; Boldt, J.; Khandurina, J.; Trawick, J.D.; Osterhout, R.E.; Stephen, R.; et al. Metabolic engineering of Escherichia coli for direct production of 1,4-butanediol. Nat. Chem. Biol. 2011, 7, 445–452. [Google Scholar] [CrossRef]
- Ng, C.; Jung, M.; Lee, J.; Oh, M.-K. Production of 2,3-butanediol in Saccharomyces cerevisiae by in silico aided metabolic engineering. Microb. Cell Fact. 2012, 11, 68. [Google Scholar] [CrossRef]
- Becker, J.; Zelder, O.; Häfner, S.; Schröder, H.; Wittmann, C. From zero to hero-Design-based systems metabolic engineering of Corynebacterium glutamicum for l-lysine production. Metab. Eng. 2011, 13, 159–168. [Google Scholar] [CrossRef] [PubMed]
- Park, J.H.; Kim, T.Y.; Lee, K.H.; Lee, S.Y. Fed-batch culture of Escherichia coli for L-valine production based on in silico flux response analysis. Biotechnol. Bioeng. 2011, 108, 934–946. [Google Scholar] [CrossRef] [PubMed]
- van Ooyen, J.; Noack, S.; Bott, M.; Reth, A.; Eggeling, L. Improved L-lysine production with Corynebacterium glutamicum and systemic insight into citrate synthase flux and activity. Biotechnol. Bioeng. 2012, 109, 2070–2081. [Google Scholar] [CrossRef]
- Jung, Y.K.; Kim, T.Y.; Park, S.J.; Lee, S.Y. Metabolic engineering of Escherichia coli for the production of polylactic acid and its copolymers. Biotechnol. Bioeng. 2010, 105, 161–171. [Google Scholar] [CrossRef]
- Poblete-Castro, I.; Binger, D.; Rodrigues, A.; Becker, J.; Martins Dos Santos, V.A.P.; Wittmann, C. In-silico-driven metabolic engineering of Pseudomonas putida for enhanced production of poly-hydroxyalkanoates. Metab. Eng. 2013, 15, 113–123. [Google Scholar] [CrossRef]
- Kildegaard, K.R.; Jensen, N.B.; Schneider, K.; Czarnotta, E.; Özdemir, E.; Klein, T.; Maury, J.; Ebert, B.E.; Christensen, H.B.; Chen, Y.; et al. Engineering and systems-level analysis of Saccharomyces cerevisiae for production of 3-hydroxypropionic acid via malonyl-CoA reductase-dependent pathway. Microb. Cell Fact. 2016, 15, 1–13. [Google Scholar] [CrossRef]
- Yang, J.E.; Park, S.J.; Kim, W.J.; Kim, H.J.; Kim, B.J.; Lee, H.; Shin, J.; Lee, S.Y. One-step fermentative production of aromatic polyesters from glucose by metabolically engineered Escherichia coli strains. Nat. Commun. 2018, 9, 79. [Google Scholar] [CrossRef]
- Kim, M.; Sang Yi, J.; Kim, J.; Kim, J.N.; Kim, M.W.; Kim, B.G. Reconstruction of a high-quality metabolic model enables the identification of gene overexpression targets for enhanced antibiotic production in Streptomyces coelicolor A3(2). Biotechnol. J. 2014, 9, 1185–1194. [Google Scholar] [CrossRef]
- Huang, D.; Wen, J.; Wang, G.; Yu, G.; Jia, X.; Chen, Y. In silico aided metabolic engineering of Streptomyces roseosporus for daptomycin yield improvement. Appl. Microbiol. Biotechnol. 2012, 94, 637–649. [Google Scholar] [CrossRef] [PubMed]
- Chemler, J.A.; Fowler, Z.L.; McHugh, K.P.; Koffas, M.A.G. Improving NADPH availability for natural product biosynthesis in Escherichia coli by metabolic engineering. Metab. Eng. 2010, 12, 96–104. [Google Scholar] [CrossRef] [PubMed]
- Bhan, N.; Xu, P.; Khalidi, O.; Koffas, M.A.G. Redirecting carbon flux into malonyl-CoA to improve resveratrol titers: Proof of concept for genetic interventions predicted by OptForce computational framework. Chem. Eng. Sci. 2013, 103, 109–114. [Google Scholar] [CrossRef]
- Xu, P.; Ranganathan, S.; Fowler, Z.L.; Maranas, C.D.; Koffas, M.A.G.G. Genome-scale metabolic network modeling results in minimal interventions that cooperatively force carbon flux towards malonyl-CoA. Metab. Eng. 2011, 13, 578–587. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Li, F.; Sánchez, B.J.; Zhu, Z.; Li, G.; Domenzain, I.; Marcišauskas, S.; Anton, P.M.; Lappa, D.; Lieven, C.; et al. A consensus S. cerevisiae metabolic model Yeast8 and its ecosystem for comprehensively probing cellular metabolism. Nat. Commun. 2019, 10. [Google Scholar] [CrossRef]
- Rocha, I.; Maia, P.; Evangelista, P.; Vilaça, P.; Soares, S.; Pinto, J.P.; Nielsen, J.; Patil, K.R.; Ferreira, E.C.; Rocha, M. OptFlux: an open-source software platform for in silico metabolic engineering. BMC Syst. Biol. 2010, 4, 1–12. [Google Scholar] [CrossRef]
- Lewis, N.E.; Hixson, K.K.; Conrad, T.M.; Lerman, J.A.; Charusanti, P.; Polpitiya, A.D.; Adkins, J.N.; Schramm, G.; Purvine, S.O.; Lopez-Ferrer, D.; et al. Omic data from evolved E. coli are consistent with computed optimal growth from genome-scale models. Mol. Syst. Biol. 2010, 6, 1–13. [Google Scholar] [CrossRef]
- Zitzler, E.; Laumanns, M.; Thiele, L. SPEA2: Improving the Strength Pareto Evolutionary Algorithm. Evol. Methods Des. Optim. Control with Appl. to Ind. Probl. 2001, 95–100. [Google Scholar] [CrossRef]
- Pereira, V.; Cruz, F.; Rocha, M. MEWpy: a computational strain optimization workbench in Python. Bioinformatics 2021, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Deb, K.; Pratap, A.; Agarwal, S.; Meyarivan, T. A fast and elitist multiobjective genetic algorithm: NSGA-II. IEEE Trans. Evol. Comput. 2002, 6, 182–197. [Google Scholar] [CrossRef]
- Mahadevan, R.; Schilling, C.H. The effects of alternate optimal solutions in constraint-based genome-scale metabolic models. Metab. Eng. 2003, 5, 264–276. [Google Scholar] [CrossRef] [PubMed]
- Entian, K.-D.; Kötter, P. 25 Yeast genetic strain and plasmid collections. In Methods in Microbiology; Stansfield, I., Stark, M.J.R., Eds.; Elsevier; Vol. 36, pp. 629–666. ISBN 0123694787.
- Brachmann, C.B.; Davies, A.; Cost, G.J.; Caputo, E.; Li, J.; Hieter, P.; Boeke, J.D. Designer deletion strains derived from Saccharomyces cerevisiae S288C: A useful set of strains and plasmids for PCR-mediated gene disruption and other applications. Yeast 1998, 14, 115–132. [Google Scholar] [CrossRef]
- Chen, Y.; Partow, S.; Scalcinati, G.; Siewers, V.; Nielsen, J. Enhancing the copy number of episomal plasmids in Saccharomyces cerevisiae for improved protein production. FEMS Yeast Res. 2012, 12, 598–607. [Google Scholar] [CrossRef] [PubMed]
- Couto, M.R.; Rodrigues, J.L.; Rodrigues, L.R. Cloning, expression and characterization of UDP-glucose dehydrogenases. Life 2021, 11, 1201. [Google Scholar] [CrossRef] [PubMed]
- Gietz, R.D.; Woods, R.A. Transformation of yeast by lithium acetate/single-stranded carrier DNA/polyethylene glycol method. In Methods in Enzymology; 2002; Vol. 350, pp. 87–96.
- Bitter, T.; Muir, H.M. A modified uronic acid carbazole reaction. Anal. Biochem. 1962, 4, 330–334. [Google Scholar] [CrossRef]
- Shaw, J.A.; Mol, P.C.; Bowers, B.; Silverman, S.J.; Valdivieso, M.H.; Duran, A.; Cabib, E. The function of chitin synthases 2 and 3 in the Saccharomyces cerevisiae cell cycle. J. Cell Biol. 1991, 114, 111–123. [Google Scholar] [CrossRef] [PubMed]
- Lesage, G.; Bussey, H. Cell Wall Assembly in Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 2006, 70, 317–343. [Google Scholar] [CrossRef] [PubMed]
- Orlean, P. Architecture and biosynthesis of the Saccharomyces cerevisiae cell wall. Genetics 2012, 192, 775–818. [Google Scholar] [CrossRef]
- Klis, F.M.; Mol, P.; Hellingwerf, K.; Brul, S. Dynamics of cell wall structure in Saccharomyces cerevisiae. FEMS Microbiol. Rev. 2002, 26, 239–256. [Google Scholar] [CrossRef]
- Zhang, Q.; Yao, R.; Chen, X.; Liu, L.; Xu, S.; Chen, J.; Wu, J. Enhancing fructosylated chondroitin production in Escherichia coli K4 by balancing the UDP-precursors. Metab. Eng. 2018, 47, 314–322. [Google Scholar] [CrossRef]
- Wu, J.; Zhang, Q.; Liu, L.; Chen, X.; Liu, J.; Luo, Q. Recombinant Escherichia coli for high efficiency production of fructosylated chondroitin and method for making thereof 2019.
- D’ambrosio, S.; Alfano, A.; Cassese, E.; Restaino, O.F.; Barbuto Ferraiuolo, S.; Finamore, R.; Cammarota, M.; Schiraldi, C.; Cimini, D. Production and purification of higher molecular weight chondroitin by metabolically engineered Escherichia coli K4 strains. Sci. Rep. 2020, 10, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Barros, M.H.; Tzagoloff, A. Regulation of the heme A biosynthetic pathway in Saccharomyces cerevisiae. FEBS Lett. 2002, 516, 119–123. [Google Scholar] [CrossRef] [PubMed]
- Sánchez, B.J.; Zhang, C.; Nilsson, A.; Lahtvee, P.; Kerkhoven, E.J.; Nielsen, J. Improving the phenotype predictions of a yeast genome-scale metabolic model by incorporating enzymatic constraints. Mol. Syst. Biol. 2017, 13, 935. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, Y.; Zhou, Z.; Wang, P.; Xi, X.; Hu, S.; Xu, R.R.; Du, G.; Li, J.; Chen, J.; et al. Synthesis of bioengineered heparin by recombinant yeast Pichia pastoris. Green Chem. 2022, 3180–3192. [Google Scholar] [CrossRef]

| Strains | Relevant genotype | Source |
|---|---|---|
| Escherichia coli NZY5α | fhuA2 Δ(argF-lacZ)U169 phoA glnV44Φ80 Δ(lacZ)M15 gyrA96 recA1 relA1 endA1 thi-1 hsdR17 | NZYTech (MB00401) |
| Saccharomyces cerevisiae CEN.PK2-1C | MATa ura3-52 his3Δ1 leu2-3,112 trp1-289 MAL2-8c SUC2 | Euroscarf 30000A [52] |
| S. cerevisiae BY4741 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 | Euroscarf Y00000 [53] |
| Plasmids | Description | Source |
| pSP-GM1 | pUC ori, AmpR, 2 µ ori, URA3 PTEF1 PPGK1 | Addgene #64739 [54] |
| pBEVY-L | pUC ori, AmpR, 2 µ ori, LEU2 PGPD PADH1 | ATCC 51226 |
| pUC57_Giuae | pMB1 ori, AmpR; pUC57 carrying Giardia intestinalis uridine diphosphate-glucosamine-4-epimerase gene (GiUAE) codon-optimized for S. cerevisiae | NZYTech |
| pUC57_Btchsy1 | pMB1 ori, AmpR; pUC57 carrying Bos taurus chondroitin synthase 1 gene (BtCHSY) codon-optimized for S. cerevisiae | NZYTech |
| pETM6_kfoCA | pETM6 carrying chondroitin synthase, kfoC, and, uridine diphosphate-glucosamine-4-epimerase, kfoA, genes from E. coli O5:K4:H4 | [5] |
| pSP-GM1_Zmugd | pSP-GM1 carrying Zymomonas mobilis uridine diphosphate glucose 6-dehydrogenase gene (Zmugd) | [55] |
| pSP-GM1_Giuae_Zmugd | pSP-GM1 carrying GiUAE and Zmugd | This study |
| pBEVY_Btchsy | pBEVY-L carrying BtCHSY | This study |
| pSP-GM1_kfoA_Zmugd | pSP-GM1 carrying kfoA and Zmugd | This study |
| pBEVY_kfoC | pBEVY-L carrying kfoC | This study |
| Compound ID code | Compound name | Template model | Corrected model 1% chitin | ||
|---|---|---|---|---|---|
| Stoichiometry | Percentage (%) | Stoichiometry | Percentage (%) | ||
| s_0001_ce | (1→3)-β-D-glucan [cell envelope] | 0.748514964 | 33.88 | 0.748514964 | 33.54 |
| s_0004_ce | (1→6)-β-D-glucan [cell envelope] | 0.250091654 | 11.32 | 0.250091654 | 11.21 |
| s_0773_c | glycogen [cytoplasm] | 0.361414528 | 16.36 | 0.361414528 | 16.20 |
| s_1107_c | mannan [cytoplasm] | 0.710939625 | 32.18 | 0.710939625 | 31.86 |
| s_1520_c | trehalose [cytoplasm] | 0.138275712 | 6.26 | 0.138275712 | 6.20 |
| s_0509_c | chitin [cytoplasm] | 0 | 0.00 | 0.022313288 | 1.00 |
| Solution | BPCY | Genes modified | Predicted phenotype (pFBA) | FVA | |||
|---|---|---|---|---|---|---|---|
| Under expression |
Over expression |
Biomass (h-1) | Chondroitin flux (mmol/gDW/h) | Minimum chondroitin flux (mmol/gDW/h) | Maximum chondroitin flux (mmol/gDW/h) | ||
| - | - | - | - | 0.8612 | 0.0000 | - | - |
| 1 | 0.04375 | - | QRI1 | 0.7317 | 0.5980 | 0.5980 | 0.9358 |
| 2 | 0.04375 | - | GNA1 | 0.7317 | 0.5980 | 0.5980 | 0.9358 |
| 3 | 0.04375 | - | PCM1 | 0.7317 | 0.5980 | 0.5980 | 0.9358 |
| Solution | BPCY | WYIELD | Genes modified | Predicted phenotype (pFBA) | FVA | ||||
|---|---|---|---|---|---|---|---|---|---|
| Knock-out | Under expression |
Over expression |
Biomass (h-1) | Chondroitin flux (mmol/gDW/h) | Minimum chondroitin flux (mmol/gDW/h) | Maximum chondroitin flux (mmol/gDW/h) | |||
| 1 | 0.04375 | 0.60872 | DNF2, CTS1 | CRD1, LYP1, FAA1 | YAH1, QRI1, PCM1, CAB5 | 0.7317 | 0.5980 | 0.5980 | 0.9358 |
| 2 | 0.04374 | 0.60872 | FAA1, POT1, CTS1 | MAL31, ARG8 | YAH1, PCM1, ALD2 | 0.7316 | 0.5980 | 0.5980 | 0.9358 |
| 3 | 0.04335 | 0.60876 | ENO1, CTS1 | ERG10 | FAA1, YAH1, AAT1, POF1, CTA1, PCM1, MAL31 | 0.7248 | 0.5981 | 0.5981 | 0.9365 |
| Solution | BPCY | WYIELD | Genes modified | Predicted phenotype (pFBA) | FVA | ||||
|---|---|---|---|---|---|---|---|---|---|
| Knock-out | Under expression |
Over expression |
Biomass (h-1) | Chondroitin flux (mmol/gDW/h) | Minimum chondroitin flux (mmol/gDW/h) | Maximum chondroitin flux (mmol/gDW/h) | |||
| 1 | 0.04375 | 0.83445 | - | - | QRI1 | 0.7317 | 0.5980 | 0.5980 | 0.9358 |
| 2 | 0.01488 | 2.80306 | CTM1 | CDS1 | QRI1 | 0.0541 | 0.6131 | 0.6131 | 3.7416 |
| 3 | 0.04359 | 0.83489 | HAM1 | SDH2 | QRI1 | 0.7288 | 0.5980 | 0.5980 | 0.9364 |
| Overexpressions | Biomass (h-1) | Chondroitin flux (mmol/gDW/h) | BPCY |
|---|---|---|---|
| PCM1, QRI1 | 0.7317 | 0.5980 | 0.04375 |
| PCM1, GNA1 | 0.7317 | 0.5980 | 0.04375 |
| QRI1, GNA1 | 0.7317 | 0.5980 | 0.04375 |
| PCM1, QRI1, GNA1 | 0.7317 | 0.5980 | 0.04375 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





