1. Introduction
The Eurasian grapevine (
Vitis vinifera L.) holds the title of being the most extensively cultivated and economically significant horticultural crop globally, being cultivate since the ancient times [
1]. Due to its substantial production, this crop plays a crucial part in the economies of many countries [
2]. The fruit is important because it can be used to consumption and also to the production of wine. The number grape varieties present in the world is unknown, but specialists estimate it to be around 5000 to 8000, under 14000 to 24000 different names [
3,
4,
5]. Despite this huge number, only 300 or 400 varieties account most of the grape plantings in the world [
4]. The most common varieties are Kyoho, Carbenet Sauvignon, Sultanina, Merlot, Tempranillo, Airen, Chardonnay, Syrah, Red Globe, Grenache Noir, Pinot Noir, Trebbiano Toscano [
6].
The grape variety plays an important role in the wine production chain and in the leave consumption, since in some cases the they can be more costly than the fruit [
7,
8]. Wine is one of the most popular agri-foods in the four corners of the world [
9]. In 2019, the European Union accounted for 48% of world consumption and 63% of world production [
10]. In terms of value, the wine market share totalled almost 29.6 billion euros in 2020, despite the Covid-19 pandemic crisis [
10]. The varieties used on the production of the drink directly influences its authenticity and classification, and due to its socioeconomic importance, identifying grape varieties became an important part of the production regulation. Furthermore, recent results achieved by Jones and Alves [
11] highlighted that some varieties can be prone to wamer environments, in the context of climate changes, accentuating the need of tools for grapevine variety identification.
Nowadays the identification of grapevine varieties is carried out mostly using ampelography or molecular analysis. Ampelography, defined by Chitwood et al. [
12] as "the science of phenotypic distinction of vines", is one of the most accurate ways of identifying grape varieties through visual analysis. Its authorised reference is Precis D’Ampelographie Pratique [
13]; however, it uses well-defined official descriptors provided in the identity of the plant material for grape identification [
14,
15]. Despite it wide utilisation, Ampelography depends on the person carrying it out, as with any visual analysis task, making the process subjective. It can be exposed to interference from environmental, cultural and genetic conditions, introducing uncertainty into the identification process [
14,
16]. It can be time-consuming and error-prone, just like any other human-based task, and ampelographers are becoming scarce [
17].
Molecular markers is another technique that has been used to identify grape varieties [
17]. Among the used markers, random amplified polymorphic DNA, amplified fragment length polymorphism and microsatellite markers have been used in the grape variety identification [
17]. This technique makes it possible to deal with subjectivity and environmental influence. However, it must be complemented by ampelography due to leaf characteristics that can only be assessed in the field [
3,
18,
19]. In addition, the identification of grape varieties with a focus on production control and regulation would involve several molecular analyses, increasing the costs and time required.
With the advance of computer vision techniques and easier access to data, several studies have emerged with the aim of automatically identifying grapevine varieties. Initially, they were based on classic machine learning classifiers, e.g. Support Vector Machines, Artificial Neural Networks [
20], Nearest Neighbour algorithm [
21], Partial least squares regression [
22], and using manually or statically extracted characteristics, e.g. indices, or the data directly. However, in 2012, with the advent of Deep Learning (DL), more specifically the study by Krizhevsky et al. [
23], computer vision classifiers became capable of reaching or, in some cases, surpassing human capacity. Lately, transfer learning and fine-tuning approaches have allowed these models to be applied to many general computer vision tasks, such as object detection, semantic segmentation and instance segmentation, and in other research domains, for example precision agriculture and medical image analysis. The automatic identification of grapevine varieties has followed this lead, and most studies now use DL-based classifiers in their approaches.
In this study, recent literature on the identification of grapevine varieties using ML and DL-based classification approaches was reviewed. The steps of the computer vision classification process (data preparation, choice of architecture or feature extraction and classifier selection, training and model evaluation) were described for 31 studies found in the literature, highlighting their pros and cons. Possible directions for improving this field of research are also presented. To the best of our knowledge, there are no studies in the literature with the same objective. However, this study may have some intersection with Chen et al. [
24], which aimed to review studies that used deep learning for plant image identification. Besides, Mohimont et al. [
25] reviewed studies that used computer vision and DL for yield-related precision viticulture tasks, e.g. flower counting, grape detection, berry counting and yield estimation, while Ferro and Catania [
26] surveyed the technologies employed in precision viticulture, covering topics ranging from sensors to computer vision algorithms for data processing. It is important to emphasise that the explanation of computer vision algorithms is already widespread in the literature and will not be covered in this study. One can refer to Chai et al. [
27] and Khan et al. [
28] for advances in the field of natural scenes, or Dhanya et al. [
29] for developments in the field of agriculture.
The remainder of this article is organised as follows. In
Section 2, the research questions, inclusion criteria, search strategy and extraction of the characteristics of the selected studies are described. Then, in
Section 3, the results are presented, highlighting the approach used in the stage of creating the DL-based classifier. In
Section 4, a discussion around the selected studies is presented, focussing on the pros and cons of the approaches used and also introducing techniques that can still be explored in the context of identifying grapevine varieties using DL-based methods. Finally, in
Section 5, the main conclusions are presented.
3. Results
As shown in
Table 2,
Table 3 and
Table 4, 30 studies were identified from the selected sources.
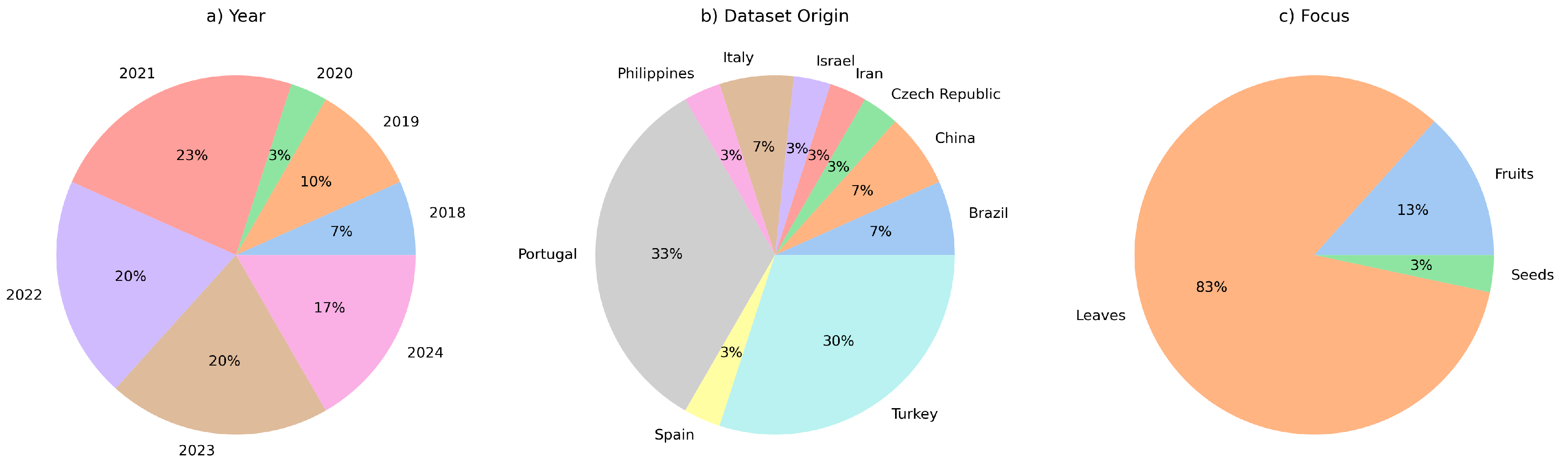
Figure 3 shows a graph comprising the countries of origin of the datasets, the years and the focus of the selected studies. The majority of the studies were published in 2021, and most of the datasets used have Portugal as their source of localisation. Therefore, this field of research has been active these days, especially in countries where the grape cultivar is economically relevant. Furthermore, most studies have focused on the leaves to identify grape varieties.
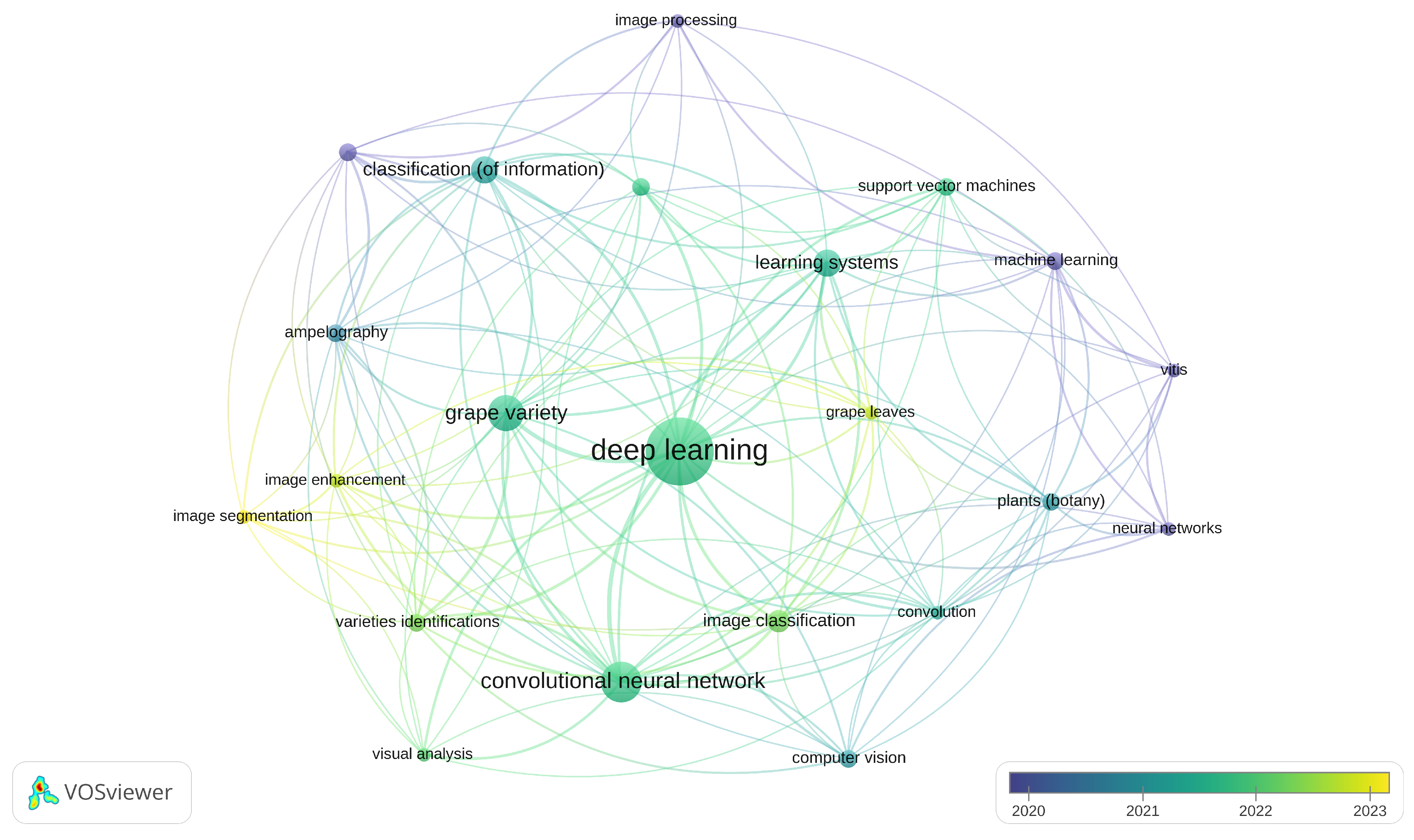
Figure 4 shows the occurrences of keywords generated for the studies included using VOSViwer [
59]. It can be seen that the number of DL-based approaches exceeds the number of ML-based approaches published in the literature over the last 6 years (ML = 7 versus DL = 24).
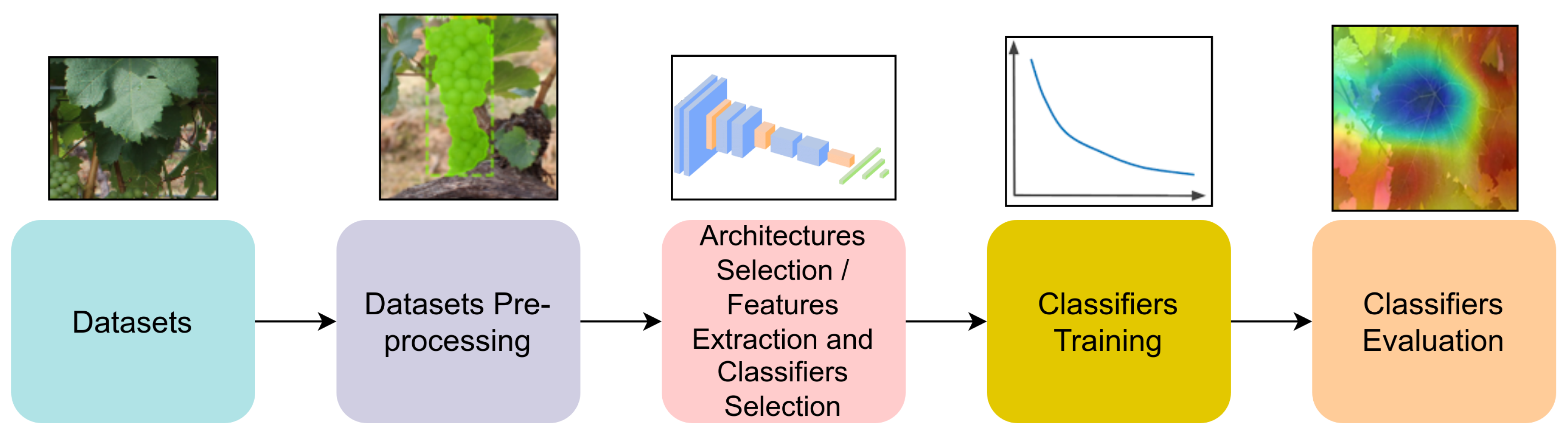
All the selected studies followed the classic process of training computer vision classifiers in their method. This process can be seen in
Figure 5. First, the data is acquired and prepared to train the classifiers. Next, pre-processing steps are applied to the data to increase the quality of the classification. Then, architectures are selected or created for DL methods, or feature extractors and classifiers are selected for ML methods, and subsequently trained on the data. In the final stage, these classifiers are evaluated. To better understand the different approaches used in the different stages, the pipeline will be followed to guide this discussion.
The datasets and benchmarks used in the studies will first be presented, followed by details of the approaches used in the pre-processing phase. Next, the architecture, or feature extractors and classifiers, and the training process adopted will be examined. Finally, the metrics and explanation techniques used to evaluate the studies will be described.
3.1. Datasets and Benchmarks
Details of the datasets used by the studies included in this review are presented in
Table 5,
Table 6 and
Table 7.
RGB images are the main way of identifying grapevine varieties in the studies included, centred on leaves, fruit and seeds. On the other hand, spectra, hyperspectral images (HSI) and the 3D point cloud have also been used as target data. Of the studies that have used RGB images, most have used datasets acquired in a controlled environment, although classifiers trained with images acquired in a controlled environment may be of limited use. Furthermore, another disadvantage is that controlled environment-based techniques are generally invasive, requiring the leaf to be removed from the plant. Similarly, studies that have used spectral signatures, through spectrometers or hyperspectral cameras, and 3D point clouds have also centred on data acquisition in a controlled environment. All the datasets that used RGB data acquired images at a maximum distance of 1.2 metres. Images acquired in the field were prone to secondary leaves and the presence of unrelated information, e.g. soil, sky, and human body parts. For the purposes of simplification, in the rest of this text RGB images will be referred to as images.
Similar to other fields of research, only a few studies provided their datasets. Peng et al. [
54] and Franczyk et al. [
55] used the grape instance segmentation dataset from Embrapa Vinho (Embrapa WGISD) [
61]. This dataset consists of 300 images belonging to 6 grape varieties. Koklu et al. [
7] provided the dataset used and, more recently, studies [
30,
39,
40,
41,
42,
46] have followed and explored the same dataset. This dataset consists of 5 classes and was acquired in a controlled environment, resulting in 500 images. De Nart et al. [
37], did not provide their dataset, although they tested their approach using the Vlah [
62] dataset. Vlah [
62] organised a dataset made up of 1009 images distributed over 11 varieties. Other datasets have also been proposed in the literature. Al-khazraji et al. [
63] proposed a dataset with 8 different grape varieties acquired in the field. Sozzi et al. [
64] proposed a dataset for cluster detection, but which can be used to identify 3 different varieties. In the same vein, Seng et al. [
65] presented a dataset with images of fruit at different stages of development that is made up of 15 different varieties.
Table 8 summarises all the publicly available datasets that, as far as we know, can be used to train and evaluate DL models with the aim of classifying different grape varieties.
Figure 6 shows examples of images for each publicly available dataset.
In the studies that provided the acquisition period, most used data obtained over a short period of time (less than a month). Carneiro et al. [
44] and Carneiro et al. [
45] used the most representative datasets, in terms of time, to identify grape varieties. It should be noted that since grapes are seasonal plants, it is very important to represent different periods of the season in the dataset in order to capture the different phenotypic characteristics of the leaves over time. Seasonal representation in the dataset directly implies the classifier’s ability to generalise.
In addition, Magalhães et al. [
43] and Fuentes et al. [
36] were the only studies that were concerned with the position of the leaves used in the classification. Magalhães et al. argue that leaves from nodes between the 7th and 8th should be used, as they are the most representative in terms of phenotypic characteristics [
66], while Fuentes et al. used mature leaves from the fifth position. De Nart et al. [
37], considered the age of the leaves in their study, excluding samples that were too young or too old, and health, discarding unhealthy leaves. In contrast, Garcia et al. [
31] considered the age of the plant, acquiring samples from plants over a year old. Other characteristics, water stress and nutrition, were not considered by the included studies.
Given that DL-based classification models are prone to overfitting, most models have used data augmentation techniques to improve the quality of the data used during training, contrasting with the ML-based classifiers. Rotations, reflections and translations are the main modifications applied to images. Furthermore, these modifications are generally only applied to the training subset. Carneiro et al. [
44] and De Nart et al. [
37] tested different techniques for data augmentation. Carneiro et al. [
44] concluded that offline geometric augmentations (zoom, rotation and flips) led the models to better classification.
Furthermore, it is clear that the datasets consisted mainly of a limited number of varieties, in contrast to the estimate made by experts, which involved at least 5000 varieties.
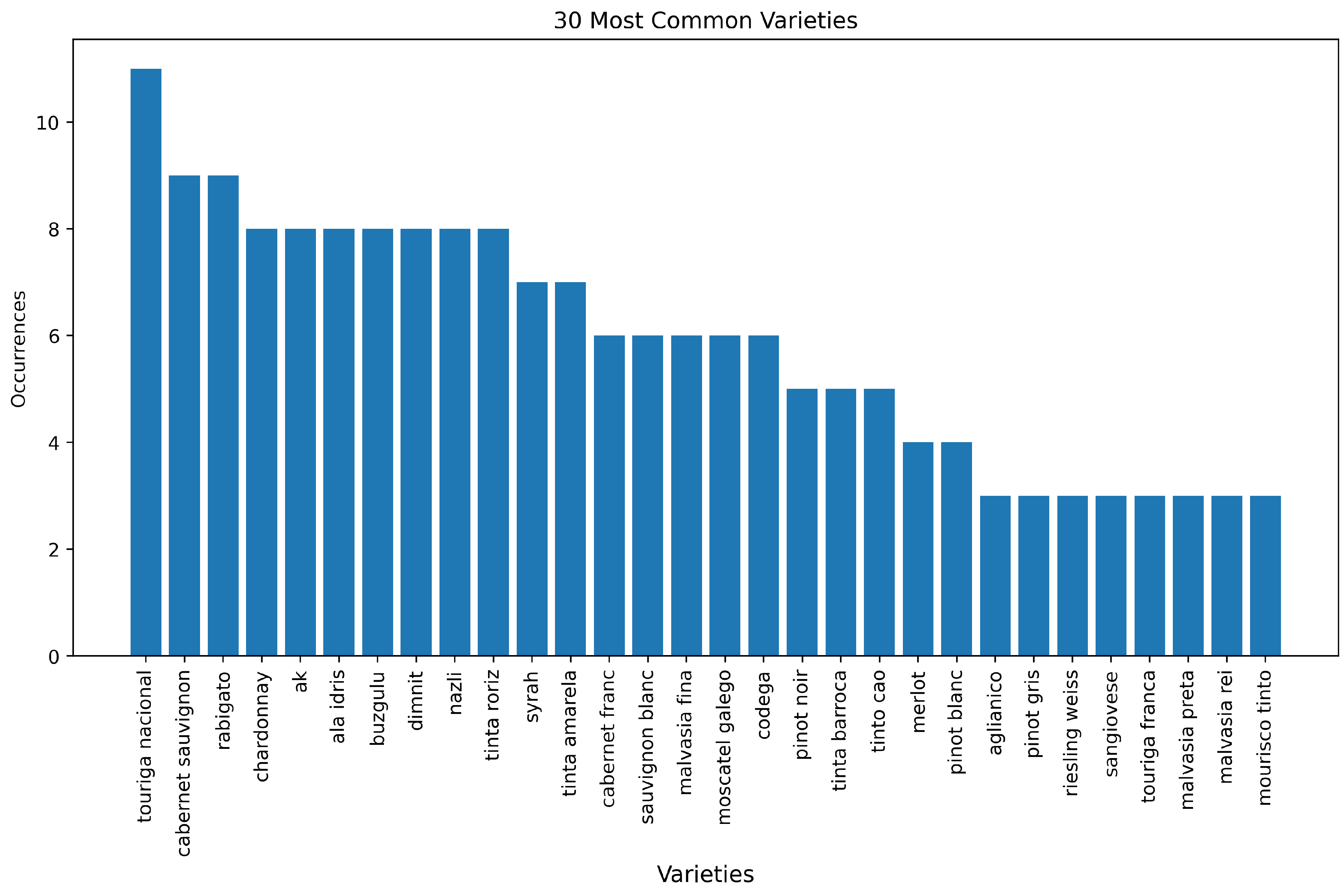
Figure 7 shows a rank of the 30 most used varieties in the included studies. It is notable that the origin of the data used in the studies and the availability of public datasets directly influence the rank. Touriga Nacional is one of the most representative varieties planted in Portugal (source of 33% of the data in the included studies), while Ak, Ala Idris, Bozgüzlü, Dimnit and Nazlı are the varieties presented in the most widely used publicly available dataset [
7]. Among the most common varieties planted around the world, Carbenet Sauvignon, Tempranillo (aka Tinta Roriz), Merlot, Chardonnay, Syrah and Pinot Noir are present in the rank. Also, one must consider that the same grape variety can be called by a synonym, for example, in Portugal, Tempranillo is known as Tinta Roriz [
67].
3.2. Pre-Processing
In the image context, Liu et al. [
51] used the complemented images in training, so that each colour channel in the resulting image was the complement of the corresponding channel in the original image. Pereira et al. [
58] tested many types of pre-processing: fixed-point FastICA algorithm [
68], canny edge detector [
69], greyscale morphology processing [
70], background removal with the segmentation method proposed by Pereira et al. [
71], and the proposed four-corners-in-one method. The FastICA algorithm is an independent component analysis method based on kurtosis maximisation that was applied to blind source separation. The idea behind applying independent component analysis (ICA) to images is that each image can be understood as a linear superposition of weighted features and then ICA decomposes them into a statistically independent source base with minimal loss of information content to achieve detection and classification [
72,
73]. Unlike ICA, grey-scale morphological processing is a method of extracting vine leaves based on classical image processing. Firstly, the image is transformed into greyscale, based on its tonality and intensity information. Next, morphological greyscale processing is applied to remove colour overlap in the leaf vein and background. Linear intensity adjustment is then used to increase the difference in grey values between the leaf vein and its background. Finally, the Otsu threshold [
74] is calculated to separate the veins from the background, and detailed processing is carried out to connect lines and remove isolated points [
70].
The next method used by Pereira et al. [
58] was proposed in Pereira et al. [
71] and is also based on classical image processing. This study presents a method for segmenting grape leaves from fruit and background in images acquired in field. This approach is based on growing regions using a colour model and thresholding techniques and can be separated into three stages: pre-processing; segmentation; and post-processing. In pre-processing, the image is resized, the histogram is adjusted to increase contrast and then the resulting image and the raw image are converted to the hue, saturation and intensity colour model, and the original image is also converted to the CIELAB (L*a*b*) colour model. In the segmentation phase, the luminance component of the raw image (L*) is used to detect the shadow regions, then the shadow and non-shadow regions are processed, removing the marks and the background with different approaches for each. Finally, in the post-processing step, the method fills in small holes using morphological operations. These holes are usually due to the presence of diseases, pests, insects, sunburn and dust on the leaves. The method achieved 94.80% average accuracy. Finally, the same authors also propose a new pre-processing method called four-corners-in-one. The idea is to concentrate all the non-removed pixels in the north-west corner of the image, after segmenting the leaves of the vines in an image; then a left-shift sequence is performed followed by a sequence of up-shift operations on the coloured pixels in the image. This algorithm is replicated for the other three corners. According to the authors, this method obtained the best classification accuracy in the set of experiments carried out.
Carneiro et al. [
47] and Carneiro et al. [
45] evaluated the use of segmentation models to remove the background from images acquired in the field before classification using DL. Both studies applied a U-Net [
75], and Carneiro et al. [
47] also tested SegNet [
76] to segment the data before classification. The results show that performance can be reduced if secondary leaves are removed and the model trained with the segmented leaves pays more attention to the centre leaves. Abbasi and Jalal [
30], Garcia et al. [
31] and Marques et al. [
34] did the same for ML-based algorithms, although with the aim of separating better leaf regions to extract better features. They used K-Means, an unspecified thresholding technique, and Otsu thresholding, respectively.
Doğan et al. [
40] used Enhanced Super Resolution Generative Adversarial Networks (ESRGAN) [
77] to increase the resolution of the images, after decreasing it, so that the method could work as a generator of new samples. The idea is to apply a Generative Adversarial Network [
78] to recover a high-resolution image from a low-resolution one. The authors applied this approach as a data augmentation technique, decreasing the resolution of the images in the dataset and increasing it again using ESRGAN, so that these new images were considered new samples.
In addition, ML applications have also employed filtering, e.g. median filter [
30], and indices, e.g. Red Green Blue Vegetation Index [
34], to improve the features extracted before classification.
In the spectra scenario, the approach to obtaining the spectra and their processing before classification has a crucial impact. Xu et al. [
32] applied threshold segmentation to the 810 nm band to obtain the leaf region and then averaged this region to generate a spectral signature. To filter the spectra, they applied the standard normal variable and the first derivative, while to de-noise the data they applied an algorithm based on empirical mode decomposition. In addition, the authors discarded the first 78 wavelengths. Gutierrez et al. [
35] selected the spectra of a pixel to represent the licence and then calculated Pearson’s
r between that pixel and a group of pixels, if it was greater than 0.9, it would be used. The authors applied the standard normal variate (SNV) and Savitzky-Golay (SG) filtering with two orders of derivatives to pre-process the spectra. The first 25 wavelengths were discarded to avoid noise. Fuentes et al.[
36] averaged the values from 5 different measurement points to calculate the spectra that would represent each leaf. Fernandes et al. [
56] after calculating the reflectance using the acquired spectra, applied the SG filter, logarithm, multiplicative scatter correction (MSC), SNV, first derivative and second derivative to the data, comparing the results for each approach.
3.3. Architecture and Training
Considering that most of the studies included in the study applied DL-based techniques to classify grape varieties, the main approach used in the architecture was an ensemble of transfer-learning and fine-tuning. However, a few different techniques were also employed: hand-crafted architectures, Fused Deep Features and extraction using DL-based models plus Support Vector Machines (SVM) classifiers. Otherwise, in ML-based studies, SVM with raw spectra emerged as the most widely used approaches.
3.3.1. Deep Learning
AlexNet [
23], VGG-16 [
79], ResNet [
80], DenseNet [
81], Xception [
82], MobileNetV2 [
83], EfficientNetV2 [
84], Inception V3 [
85], Inception ResNet [
86] were the Convolutional Neural Networks (CNNs) architectures employed in the image-based studies included in this review. These networks were first trained on ImageNet and then transfer learning and fine-tuning were employed in two stages. In the first step, their classifiers are replaced by a new one and the convolutional weights are frozen so that only the new classifier is trained and has its weights updated (transfer learning). In the second step, all the weights are unfrozen and the entire architecture is retrained (tuning). Detailed information on each architecture used can be found in Alzubaidi et al. [
87]. In a different way, Carneiro et al. [
47] and Kunduracioglu and Pacal [
38] used Vision Transformers throught ViT [
88], Swin-Transformers [
89], MobileViT [
90], Deit [
91], and MaxViT [
92] but followed the same learning strategy.
Unlike other studies based on image classification, Peng et al. [
54], Lv [
42], and Doğan et al. [
40] used Fused Deep Features to identify grapevine varieties. This approach consists of extracting features from images from more than one source, concatenating all the extracted features and then classifying them. Peng et al. [
54] extracted features from AlexNet, ResNet and GoogLeNet, then fused the features using the Canonical Correlation Analysis algorithm [
93] and then classified the vine varieties using an SVM classifier. In addition, the authors trained the aforementioned architectures with fully connected classifiers, which resulted in worse performance than the proposed method. They argued that the small size of the dataset is the main reason why it is difficult to obtain better results using CNN directly. Lv [
42] merged the results of VGG-19, ViT, Inception Resnet, DenseNet and ResNext, but instead of merging the extracted features, the final classification of each model was used so that voting strategies were applied to obtain the final classification. Doğan et al. [
40] merged attributes from VGG-19 and MobileNetV2. The difference is that the authors used a Genetic Based Support Vector Machine to select the best features for classification, improving the results by 3 percentage points. The final classification of the selected feature was done using an SVM. Koklu et al. [
7] also used the features extracted from a pre-trained CNN architecture plus an SVM classifier. The idea was to extract features from the logits of the first fully connected layer of MobileNetV2 and use them to test the performance of four different SVM kernels: Linear, Quadratic, Cubic and Gaussian. In addition, the authors carried out experiments using the Chi-Square test to select the 250 most representative features in the logits.
Regarding transfer-learning and fine-tuning, Carneiro et al. [
48] and Ahmed et al. [
60] analysed the impact of different frozen layers. Carneiro et al. [
48] obtained the same metrics for different frozen layers, while Ahmed et al. [
60] showed that freezing the half-convolution part of the model leads to better classification. On the other hand, Carneiro et al. [
48] also stated, using Explainable Artificial Intelligence (XAI), that freezing the entire convolution part or training the entire model leads to more similar results, in terms of explainability, than training only half of the convolutional part of the model, whereby in the latter case the background part of the images contributed more to classification than in the first case.
Some optimisers, global pooling techniques and losses were used for training. Among the optimisers available in the literature for training machine learning models, Stochastic Descent Gradient (SGD) and Adam [
94] were used in the selected studies. In addition to SGD, it was possible to use an adaptive learning rate scaler or a momentum technique to improve the training process.
All the image-based studies that have used a global pooling method have opted for Global Average Pooling, with the aim of reducing the CNN activation maps before classification. The losses used were Cross Entropy loss (CE) and Focal Loss (FL) [
95]. Focal Loss is a modification of the CE loss that reduces the weight of easy examples and thus concentrates training on difficult cases. It was first used in object detection studies, due to its huge imbalance between detected bins of "objects" and "non-objects", however Mukhoti et al. [
96] concluded that it can also be used to deal with calibration errors of multi-class classification models, in the sense that the probability values they associate with the labels of the classes they predict overestimate the probabilities of those labels being correct in the real world. Carneiro et al. [
50] used Focal Loss to mitigate the imbalance in the dataset used.
3.3.2. Machine Learning
The training process for ML-based approaches can be seen in three stages: 1) feature extraction, 2) dimension reduction and/or feature selection and 3) classification.
The included studies targeting HSI and point clouds did not apply any feature extraction process, using the raw spectra/points as a representation of the samples. On the other hand, a number of descriptors were adopted when classifying the images. Most of the descriptors extracted were based on colour (e.g. RGB statistical values, histogram operations, transformation to CEILAB L*a*b* colour space), shape (e.g. roundness, aspect ratio, convex area, perimeter) and texture (e.g. entropy, contrast, energy, homogeneity). In a different way, Fuentes et al. [
36] applied fractal dimension analysis and Abassi and Jalal used KAZE [
97] as feature descriptors. Fractal dimension analysis measures the complexity of shapes. Abassi and Jalal applied it to leaf shapes using the box-counting method. On the other hand, KAZE is a feature detector-descriptor algorithm that exploits non-linear diffusion filtering in the detection and description of multi-scale features, so that these features use a non-linear scale space instead of a Gaussian scale space. The extracted features are robust to changes in size (scaling), orientation (rotation), small distortions (limited affine transformations) and, furthermore, these features are distinct at various image scales [
98]. In the 3D point cloud scenario, the Pair-Wise Iterative Closest point was applied to obtain the similarity between pairs of point clouds.
For feature reduction or selection studies targeting images, recursive removal of highly correlated features [
34] and Principal Component Analysis (PCA) [
36] were applied. Also, Garcia et al. [
31] evaluated the use of three selection methods: Kendall’s Rank Coefficient, Wrapper and embedding. In the Kendall’s Rank Coefficient, features were selected by their level of correlation, removing redundant features. The Wrapper method used a logistic regression model as an estimator to select the best combination of characteristics. The embedding method used a Random Forest to determine the weights of each characteristic. According to the authors, the best performance was achieved by the embedding method.
Finally, SVM, k-NN, decision tree, Linear Discriminant Analysis, Logistic Regression, Softmax Regression, Gaussian Naive Bayes and Artificial Neural Networks (ANN) were used for classification. It is worth noting that the studies that used HSI opted only for SVMs or ANNs. On the other hand, for those targeting images, all classifiers were explored.
3.4. Evaluation
Aiming to quantitatively evaluate trained models, accuracy is the most used metric, followed by the F1 Score. Some studies also use precision, recall, Area-Under-The-Curve (AUC), specificity, or the Matthews correlation coefficient (MCC).
On the other hand, as in other areas of research, some studies based on DL use XAI to qualitatively evaluate their models. XAI is a set of processes and methods aimed at enabling humans to understand, adequately trust and effectively manage the emerging generation of artificially intelligent models [
99]. The techniques employed by the selected studies are model-agnostic for post-hoc explainability,which means that no modification to the architecture was necessary in order to apply them.
Nasiri et al. [
53] and Pereira et al.[
58] extracted the filters learnt by their models. In addition, Nasiri et al. [
53] also produced Saliency Maps. Carneiro et al. [
49] and Liu et al. [
51] used Grad-CAM [
100] to obtain heatmaps focused on the pixel’s contribution to a specific class. Carneiro et al. [
44] also used Grad-CAM to evaluate models, but instead of analysing the heatmaps generated, they used them to calculate the classification similarity between pairs of trained models. The authors calculated the heatmaps for the test subset for the models and calculated the cosine similarity between these heatmaps for the pairs of models. The authors concluded that, among the data augmentation approaches used, static geometric transformations generate representations more similar to RandAugment than to CutMix.
Carneiro et al. [
50] used Local Interpretable Model-Agnostic Explanations (LIME) [
101] for the same purpose. Furthermore, Carneiro et al. [
47] extracted attention maps from ViT and checked the impact of sample rotation using them.
To generate saliency maps, Nasiri et al. [
53] began by calculating the derivative of a class score function, which can be approximated by a first-order Taylor expansion. The elements of the calculated derivative were then rearranged. Grad-CAM is a technique proposed by Selvaraju et al. [
100] that aims to explain how a model concludes that an image belongs to a certain class. The idea is to use the gradient of the score of the predicted class in relation to the activation maps of a selected convolutional layer. The selection of the convolutional layer is arbitrary. As a result, heat maps are obtained containing the regions that contribute positively to image classification. According to the authors, obtaining explanations of the predictions using Grad-CAM makes it possible to increase human confidence in the model and, at the same time, to understand classification errors. Like Grad-CAM, LIME [
101] is an explainability approach used to explain individual machine learning model predictions for a specific class. Unlike Grad-CAM, it is not restricted to CNNs, so it is applicable to any machine learning classifier. The idea behind LIME is to train an explainable surrogate model with a new dataset composed of perturbed samples (e.g. hiding parts of the image) derived from the target data, so that it becomes a good approximation of the original model locally (in the neighbourhood of the target data). Then, from the surrogate interpretative model, it is possible to obtain the regions that have contributed to the classification, both positively and negatively.
Author Contributions
Conceptualization, G.A.C., A.C., and J.S.; methodology, G.A.C.; validation, J.S. and A.C.; formal analysis, G.A.C., A.C., and J.S.; investigation, G.A.C.; resources, G.A.C., A.C., and J.S.; data curation, G.A.C.; writing—original draft preparation, G.A.C and J.S; writing—review and editing, G.A.C, A.C., and J.S.; visualization, G.A.C.; supervision, J.S and A.C..; project administration, J.S.; funding acquisition, J.S. All authors have read and agreed to the published version of the manuscript.
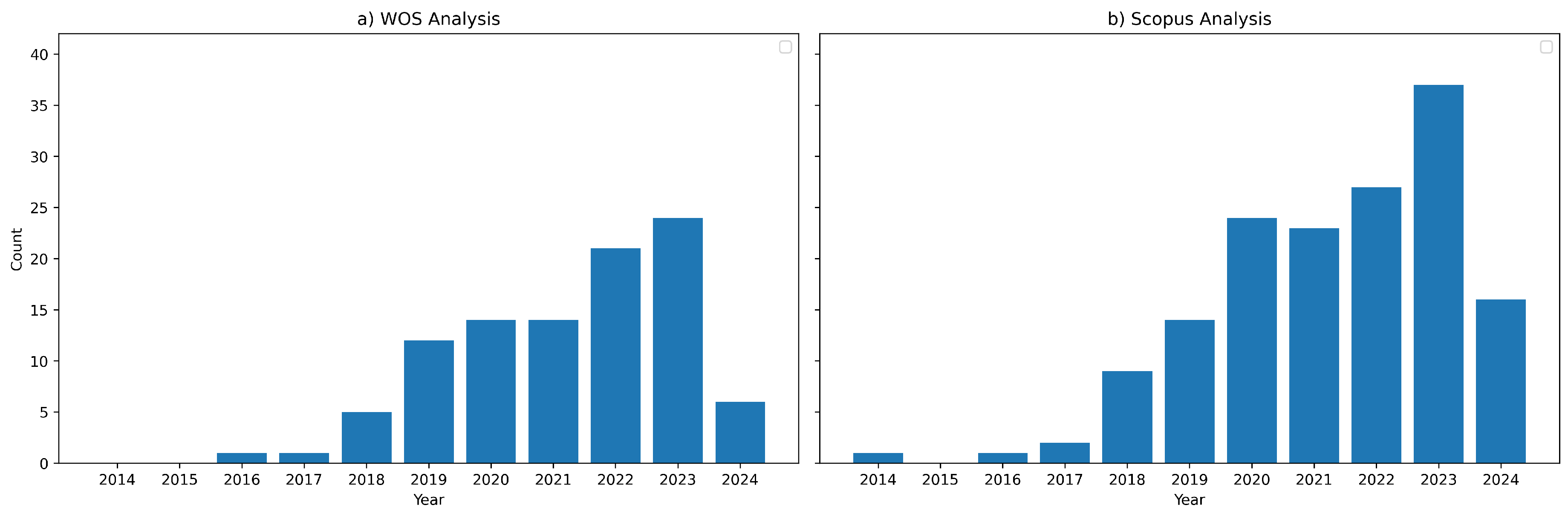
Figure 1.
Results from queries about studies in the field of machine and deep learning applied to grapevine viticulture identification without date filtering for Scopus and Web of Science. Details of the queries are shown on
Section 2.3.
Figure 1.
Results from queries about studies in the field of machine and deep learning applied to grapevine viticulture identification without date filtering for Scopus and Web of Science. Details of the queries are shown on
Section 2.3.
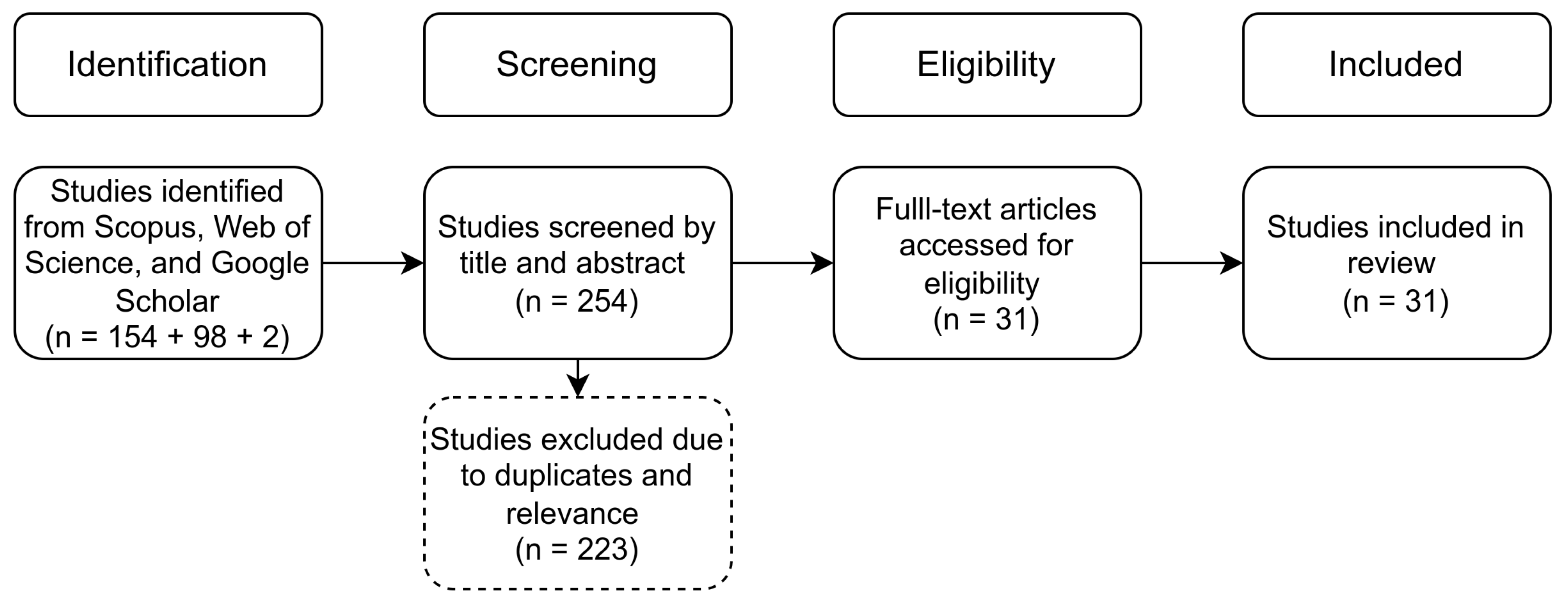
Figure 2.
PRISMA workflow applied in this study.
Figure 2.
PRISMA workflow applied in this study.
Figure 3.
Years, countries of origin of the datasets and focus of the selected studies. Most of the studies were published in 2021. Portugal was the biggest source of datasets. The studies focus on leaves rather than fruits and seeds to identify grape varieties.
Figure 3.
Years, countries of origin of the datasets and focus of the selected studies. Most of the studies were published in 2021. Portugal was the biggest source of datasets. The studies focus on leaves rather than fruits and seeds to identify grape varieties.
Figure 4.
Co-occurrences of keywords for the included studies. Figure generated with VOSViewer. [
59]
Figure 4.
Co-occurrences of keywords for the included studies. Figure generated with VOSViewer. [
59]
Figure 5.
Classic pipeline for training DL-based models. Starting with data acquisition and ending with model evaluation.
Figure 5.
Classic pipeline for training DL-based models. Starting with data acquisition and ending with model evaluation.
Figure 6.
Example images for each publicly available dataset. The images have been cropped into squares, so they cannot represent the actual size of the images in the dataset.
Figure 6.
Example images for each publicly available dataset. The images have been cropped into squares, so they cannot represent the actual size of the images in the dataset.
Figure 7.
Example images for each publicly available dataset. The images have been cropped into squares, so they cannot represent the actual size of the images in the dataset.
Figure 7.
Example images for each publicly available dataset. The images have been cropped into squares, so they cannot represent the actual size of the images in the dataset.
Table 1.
Details of the search carried out. Access to both search engines was made on April 22, 2024.
Table 1.
Details of the search carried out. Access to both search engines was made on April 22, 2024.
| Database |
Website |
Query |
| Scopus |
https://www.scopus.com/home.uri |
TITLE-ABS-KEY (("grape variety" OR "grapevine") AND ("classification" OR "identification" OR "detection") AND "deep learning" OR "machine learning") |
| Web of Science |
https://www.webofscience.com/ |
TS=(("grape variety" OR "grapevine") AND ("classification" OR "identification" OR "detection") AND ("deep learning" OR "machine learning")) |
Table 2.
A summary of the selected studies that used ML to classify grape varieties. The year, location of the data set, description, the part of the plant where the images were focused, the feature extractors and classifiers used, as well as the results are presented.
Table 2.
A summary of the selected studies that used ML to classify grape varieties. The year, location of the data set, description, the part of the plant where the images were focused, the feature extractors and classifiers used, as well as the results are presented.
| Study |
Year |
Data Location |
Dataset Description |
Focus |
Features Extractor |
Classifiers |
Results |
| Abassi and Jalal [30] |
2024 |
Turkey |
500 RGB images distributed between 5 classes acquired in controlled environment |
Leaves |
Kaze and Blob |
Softmax Regression |
83.20 (Acc) |
| Garcia et al. [31] |
2022 |
Philippines |
1149 RGB images dsitributed between 7 classes acquired in controlled environment |
Leaves |
Color, Texture, and shape mensurement analysis |
SVM, k-NN, and Decision Tree |
89.00 (F1) |
| Xu et al. [32] |
2021 |
China |
480 spectras distributed between 4 classes acquired in controlled environment |
Fruits |
Raw Signature |
SVM |
99.31 (Acc) |
| Landa et al. [33] |
2021 |
Israel |
400 3D point clouds distributed between 8 classes acquired in a controlled environment |
Seeds |
Pair-wise using Iterative Closest Point |
Linear Discriminant Analisys |
93.00 (Acc) |
| Marques et al. [34] |
2019 |
Portugal |
240 RGB images distributed between 3 classes acquired in controlled environment |
Leaves |
Color and Shape Features |
Linear Discriminant Analisys, Logistic Regression, k-NN, Decision Tree, Gaussian Naive Bayes, SVM |
86.90 (F1) |
| Gutiérrez et al. [35] |
2018 |
Italy |
2400 spectras distributed between 30 varieties acquired in field using vehicle |
Leaves |
Raw Signature |
SVM and ANN |
0.99 (F1) |
| Fuentes et al. [36] |
2018 |
Spain |
138 RGB and 144 spectra distributed between 16 varities in a controlled environment |
Leaves |
Fractal dimensions, color and shape mensurement; Raw Spectra |
ANN |
71.44 (Acc) |
Table 3.
A summary of the selected studies that used DL to classify grape varieties published between 2023 and 2024. The year, location of the data set, description, the part of the plant where the images were focused, the architectures used, as well as the results are presented.
Table 3.
A summary of the selected studies that used DL to classify grape varieties published between 2023 and 2024. The year, location of the data set, description, the part of the plant where the images were focused, the architectures used, as well as the results are presented.
| Study |
Year |
Data Location |
Dataset Description |
Focus |
Architecture |
Results |
| De Nart et al. [37] |
2024 |
Italy |
26382 RGB images distributed between 27 classes acquired in field and in a controlled environment |
Leaves |
MobileNetV2, EfficientNet, ResNet, Inception ResNet V2, and Inception V3 |
1.00 (Acc) |
| Kunduracioglu and Pacal [38] |
2024 |
Turkey |
500 RGB images distributed between 5 classes acquired in controlled environment |
Leaves |
VGG-16, ResNet, Xception, Inception, EfficientNetV2, DenseNet, SwinTransformers, MobileViT, ViT, Deit, MaxVit |
100.00 (F1) |
| Rajab et al. [39] |
2024 |
Turkey |
500 RGB images distributed between 5 classes acquired in controlled environment |
Leaves |
VGG-16 and VGG-19 |
100.00 (Acc) |
| Doğan et al. [40] |
2024 |
Turkey |
7000 RGB images distributed between 5 classes acquired in controlled environment |
Leaves |
Fused Deep Features + SVM |
1.00 (F1) |
| Sun et al. [41] |
2023 |
Turkey |
500 RGB images distributed between 5 classes acquired in controlled environment |
Leaves |
Handcraft |
91.58 (F1) |
| Lv [42] |
2023 |
Turkey |
2800 RGB images distributed between 5 classes acquired in controlled environment |
Leaves |
VGG-19, ViT, Inception ResNet, DenseNet, ResNext |
0.98 (F1) |
| Magalhães et al. [43] |
2023 |
Portugal |
40428 RGB images distributed between 26 classes acquired in a controlled environment |
Leaves |
MobileNetV2, ResNet-34 and VGG-11 |
94.75 (F1) |
| Carneiro et al. [44] |
2023 |
Portugal |
6216 RGB images distributed between 14 classes acquired in field |
Leaves |
EfficientNetV2S |
0.89 (F1) |
| Carneiro et al. [45] |
2023 |
Portugal |
675 RGB images distributed between 12 classes; 4354 RGB images distributed between 14 classes; both acquired in field |
Leaves |
EfficientNetV2S |
0.88 (F1) |
| Gupta and Gill [46] |
2023 |
Turkey |
2500 RGB images distributed between 5 classes acquired in controlled environment |
Leaves |
EfficientNetB5 |
0.86 (Acc) |
Table 4.
A summary of the selected studies that used DL to classify grape varieties published between 2018 and 2022. The year, location of the data set, description, the part of the plant where the images were focused, the architectures used, as well as the results are presented.
Table 4.
A summary of the selected studies that used DL to classify grape varieties published between 2018 and 2022. The year, location of the data set, description, the part of the plant where the images were focused, the architectures used, as well as the results are presented.
| Study |
Year |
Data Location |
Dataset Description |
Focus |
Architecture |
Results |
| Ahmed et al. [47] |
2022 |
Turkey |
500 RGB images distributed between 5 classes acquired in controlled environment |
Leaves |
DenseNet201 |
98.02 (F1) |
| Carneiro et al. [47] |
2022 |
Portugal |
28427 RGB images distributed between 6 classes acquired in field |
Leaves |
Xception |
0.92 (F1) |
| Carneiro et al. [48] |
2022 |
Portugal |
6922 RGB images distributed between 12 classes acquired in-field |
Leaves |
Xception |
0.92 (F1) |
| Carneiro et al. [49] |
2022 |
Portugal |
6922 RGB images distributed between 12 classes acquired in-field |
Leaves |
Vision Transformer (ViT_B) |
0.96 (F1) |
| Koklu et al. [7] |
2022 |
Turkey |
2500 RGB images distributed between 5 classes acquired in controlled environment |
Leaves |
MobileNetV2 + SVM |
97.60 (Acc) |
| Carneiro et al. [50] |
2021 |
Portugal |
6922 RGB images distributed between 12 classes acquired in-field |
Leaves |
Xception |
0.93 (F1) |
| Liu et al. [51] |
2021 |
China |
5091 RGB images distributed between 21 classes acquired in-field |
Leaves |
GoogLeNet |
99.91 (Acc) |
| Škrabánek et al. [52] |
2021 |
Czech Republic |
7200 RGB images distributed between 7 classes acquired in-field |
Fruits |
DenseNet |
98.00 (Acc) |
| Nasiri et al. [53] |
2021 |
Iran |
300 RGB images distributed between 6 classes acquired in a controlled environment |
Leaves |
VGG-16 |
99.00 (Acc) |
| Peng et al.[54] |
2021 |
Brazil |
300 RGB images distributed between 6 classes acquired in-field |
Fruits |
Fused Deep Features |
96.80 (F1) |
| Franczyk et al. [55] |
2020 |
Brazil |
3957 RGB images distributed between 5 classes acquired in-field |
Fruits |
ResNet |
99.00 (Acc) |
| Fernandes et al. [56] |
2019 |
Portugal |
35933 Spectra distributed between 64 classes in-field |
Leaves |
Handcraft and SVM |
0.98 (AUC) |
| Adão et al. [57] |
2019 |
Portugal |
3120 RGB images distributed between 6 classes acquired in controlled environment |
Leaves |
Xception |
100.00 (Acc) |
| Pereira et al. [58] |
2019 |
Portugal |
224 RGB images distributed between 6 classes acquired in controlled environment |
Leaves |
AlexNet |
77.30 (Acc) |
Table 5.
Datasets characteristics for each selected study that used ML.
Table 5.
Datasets characteristics for each selected study that used ML.
| Study |
Acquisition Device |
Publicly Available |
Acquisition Period |
D. Augmentation |
Acquisition Environment |
| Abassi and Jalal [30] |
Camera – Prosilica GT2000C |
Yes |
- |
No |
Special Illumination box |
| Garcia et al. [31] |
Smartphone |
No |
- |
No |
Controlled Environment |
| Xu et al. [32] |
Spectrograph – ImSpectorV10 |
No |
1 Day |
No |
Box with defined distance |
| Landa et al. [33] |
Microscope – Nikon SMZ25 |
No |
1 Day |
No |
Special Illumination box covered in aluminium foil |
| Marques et al. [34] |
Camera – Canon 600D |
No |
Season 2017 |
No |
Controlled Environment |
| Gutierrez et al. [35] |
Hyperspectral Imaging Camera – Resonon Pika L |
No |
Two days |
No |
In field using a vehicle at 5 km/h |
| Fuentes et al. [36] |
Scanner – Hewlett Packard Scanjet G3010; Spectrometer – Ocean Optics HR2000+ |
No |
- |
No |
Controlled Environment |
Table 6.
Datasets characteristics for each selected study that used DL published between 2023 and 2024.
Table 6.
Datasets characteristics for each selected study that used DL published between 2023 and 2024.
| Study |
Acquisition Device |
Publicly Available |
Acquisition Period |
D. Augmentation |
Acquisition Environment |
| De Nart et al. [37] |
Mixed Camera and Smartphone |
Yes |
Seasons 2020 and 2021 |
Flips, rotation, scale, CutMix and Zoom |
In field and Controlled Environment |
| Kunduracioglu and Pacal [38] |
Camera - Prosilica GT2000C |
Yes |
- |
Flips, rotation, scale, CutMix and Zoom |
Special illumination box |
| Rajab et al. [39] |
Camera - Prosilica GT2000C |
Yes |
- |
- |
Special illumination box |
| Doğan et al. [40] |
Camera - Prosilica GT2000C |
Yes |
- |
Static augmentations and artificially generated images |
Special illumination box |
| Sun et al. [41] |
Camera - Prosilica GT2000C |
Yes |
- |
Rotations, flips and scale |
Special illumination box |
| Lv [42] |
Camera - Prosilica GT2000C |
Yes |
- |
Random erasing, zoom, scale and Gaussian noise |
Special illumination box |
| Magalhães et al. [43] |
Kyocera TASKalfa 2552ci |
No |
1 day in June 2021 |
Blur, rotations, variations in brightness, horizontal flips, and gaussian noise |
Special illumination box |
| Carneiro et al. [44] |
Smartphones |
No |
Seasons 2021 and 2020 |
Static augmentations, CutMix and RandAugment |
In-field |
| Carneiro et al. [45] |
Mixed Camera and Smartphone |
No |
1 Season and 2 Seasons |
Rotations, Flips and Zoom |
In-field |
| Gupta and Gill [46] |
Camera - Prosilica GT2000C |
Yes |
- |
Angle, scaling factor, translation |
Special illumination box |
Table 7.
Datasets characteristics for each selected study that used DL published between 2018 and 2022.
Table 7.
Datasets characteristics for each selected study that used DL published between 2018 and 2022.
| Study |
Acquisition Device |
Publicly Available |
Acquisition Period |
D. Augmentation |
Acquisition Environment |
| Ahmed et al. [60] |
Camera - Prosilica GT2000C |
Yes |
- |
Flip, rotation, sharpen, variation in brightness |
Special illumination box |
| Carneiro et al. [47] |
Mixed Camera and Smartphone |
No |
Seasons 2017 and 2020 |
Rotation, shift, flip, and brightness changes |
In-field |
| Carneiro et al. [48] |
Camera - Canon EOS 600D |
No |
- |
Rotations, shifts, variations in brightness and flips |
In-field |
| Carneiro et al. [49] |
Camera - Canon EOS 600D |
No |
1 Season |
Rotations, shifts, variations in brightness and flips |
In-field |
| Koklu et al. [7] |
Camera - Prosilica GT2000C |
Yes |
- |
Angle, scaling factor, translation |
Special illumination box |
| Carneiro et al. [50] |
Camera - Canon EOS 600D |
No |
1 Season |
Rotations, shifts, variations in brightness and flips |
In-field |
| Liu et al. [51] |
Camera - Canon EOS 70D |
No |
- |
Scaling, transposing, rotation and flips |
In-field |
| Škrabánek et al. [52] |
Camera – Canon EOS 100D and Canon EOS 1100D |
No |
2 days in August 2015 |
- |
In-field |
| Nasiri et al. [53] |
Camera – Canon SX260 HS |
On request |
1 day in July 2018 |
Rotation, height and width shift |
Capture station with artificial light |
| Peng et al.[54] |
Mixed Camera and Smartphone |
Yes |
1 day in April 2017 and 1 day in April 2018 |
- |
In-field |
| Franczyk et al. [55] |
Mixed Camera and Smartphone |
Yes |
1 day in April 2017 and 1 day in April 2018 |
- |
In-field |
| Fernandes et al. [56] |
Spectrometer – OceanOptics Flame-S |
No |
4 days in July 2017 |
- |
In-field |
| Adão et al. [57] |
Camera - Canon EOS 600D |
No |
Season 2017 |
Rotations, contrasts/brightness, vertical/horizontal mirroring, and scale variations |
Controlled environment with white background |
| Pereira et al. [58] |
Mixed Camera and Smartphone |
No |
Seasons 2016 and 2019 |
Translation, reflection, rotation |
In-field |
Table 8.
List of publicly available datasets that can be used to classify grapevine varieties.
Table 8.
List of publicly available datasets that can be used to classify grapevine varieties.
| Dataset |
Number of Classes |
Number of images |
Classes |
Balanced |
| Koklu et al. [7] |
5 |
500 |
Ak, Ala Idris, Bozgüzlü, Dimnit, Nazlı |
Yes, 100 images per class |
| Al-khazraji et al. [63] |
8 |
8000 |
deas al-annz, kamali, halawani, thompson seedless, aswud balad, riasi, frinsi, shdah |
Yes, 1000 images per class |
| Santos et al. [61] |
6 |
300 |
Chardonnay, Cabernet Franc, Cabernet Sauvignon, Sauvignon Blanc, Syrah |
No |
| Sozzi et al. [64] |
3 |
312 |
Glera, Chardonnay, Trebbiano |
No |
| Seng et al. [65] |
15 |
2078 |
Merlot, Cabernet Sauvignon, Saint Macaire, Flame Seedless, Viognier, Ruby Seedless, Riesling, Muscat Hamburg, Purple Cornichon, Sultana, Sauvignon Blanc, Chardonnay |
No |
| Vlah [62] |
11 |
1009 |
Auxerrois, Cabernet Franc, Cabernet Sauvignon, Chardonnay, Merlot, Müller Thurgau, Pinot Noir, Riesling, Sauvignon Blanc, Syrah, Tempranillo |
No |