Submitted:
25 June 2024
Posted:
26 June 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
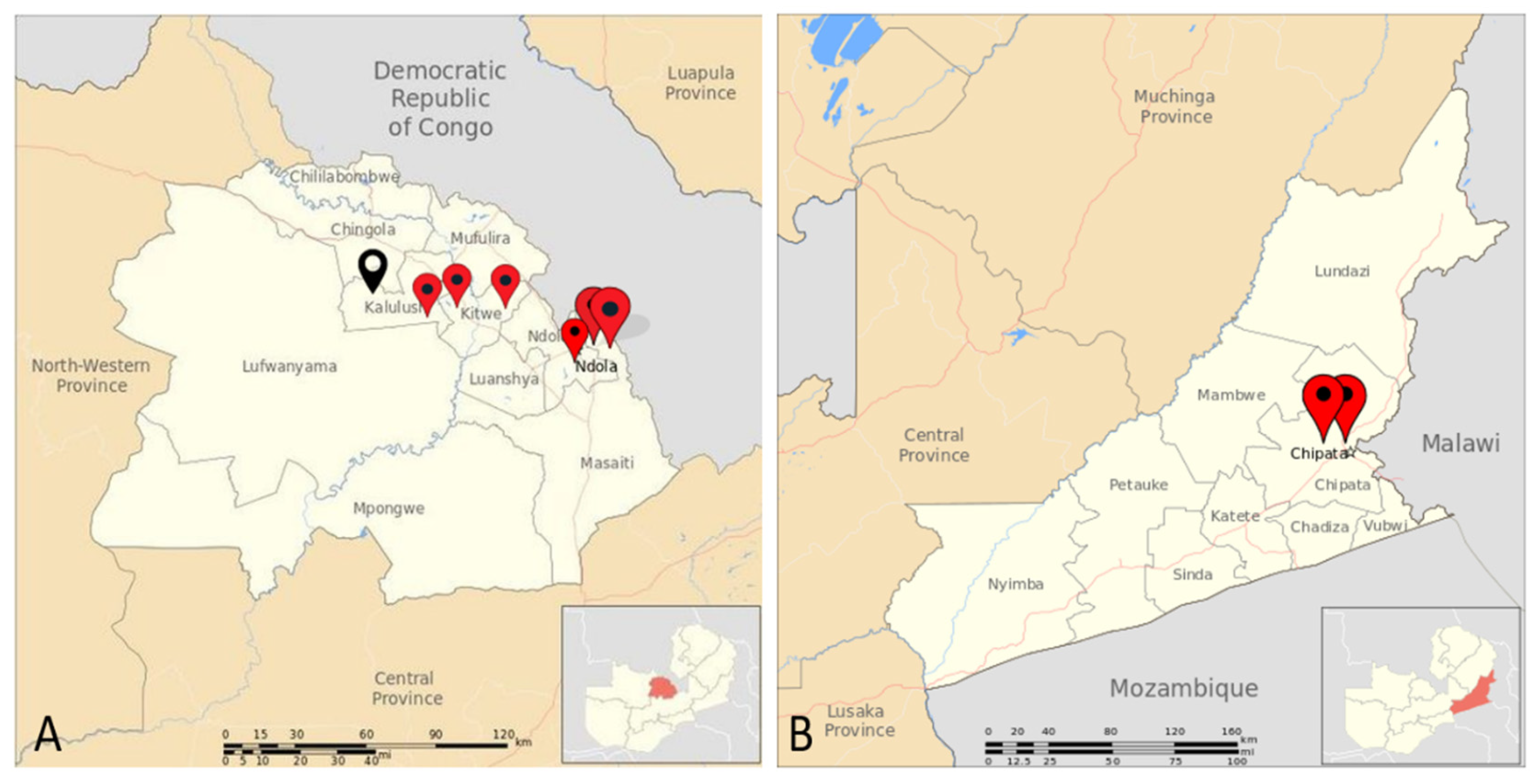
2.1. Study Design and Sites
2.2. Sample Collection
2.3. Viral Concentration
2.3.1. Skimmed Milk Flocculation
2.3.2. Bag-Mediated Filtration System (BMFS)
2.3.3. Polyethylene Glycol-Based (PEG) Concentration
2.4. Nucleic Acid Extraction
2.5. SARS-CoV-2 Detection by RT-PCR
2.6. Genomic Sequencing
2.6.2. Library Preparation and Illumina Sequencing
2.6.3. Genome Assembly and Annotation
2.7. Data Analysis
2.8. Phylogenetic Analysis
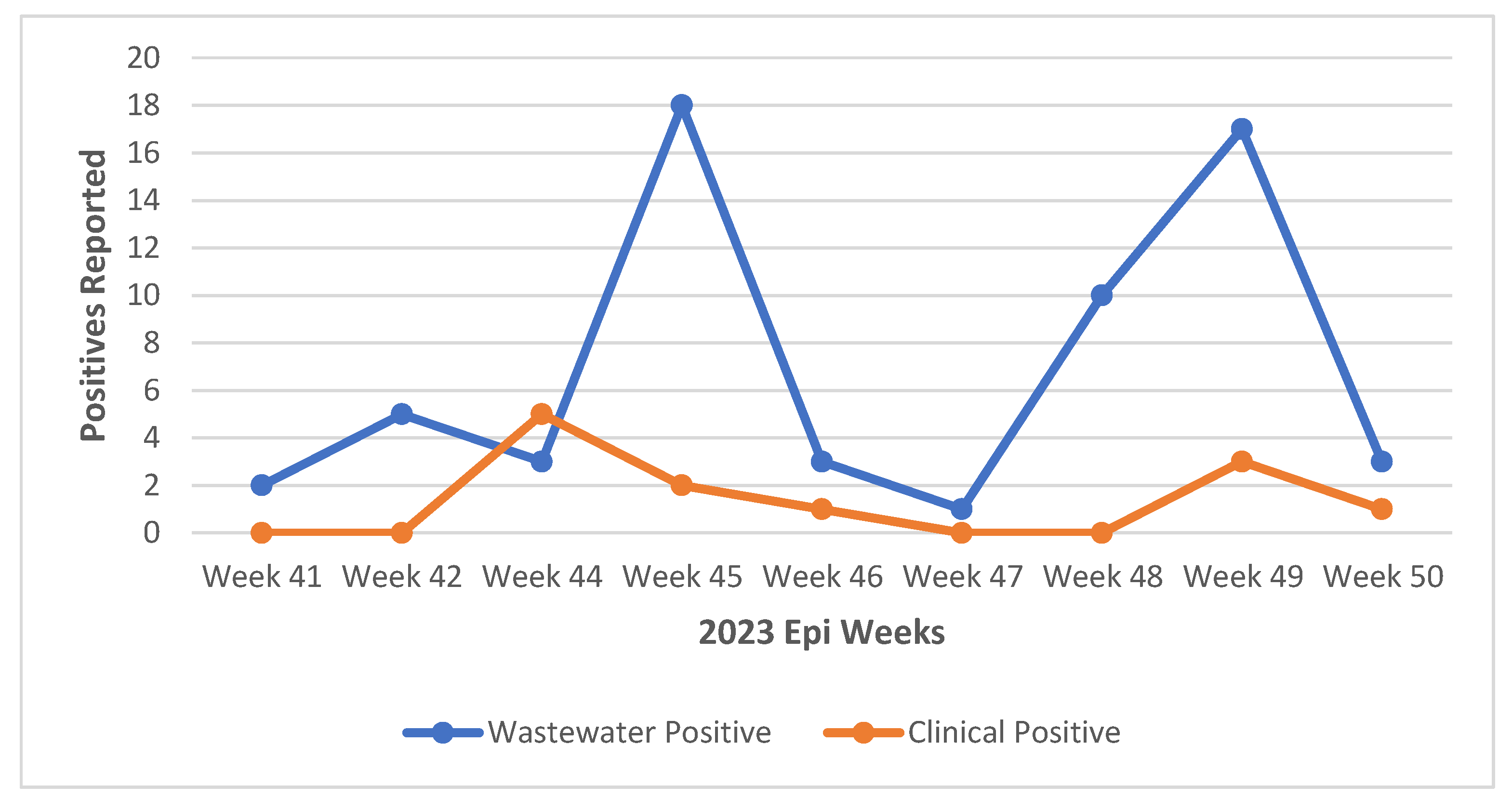
2.9. Collection of Clinical Data for COVID-19
3. Results
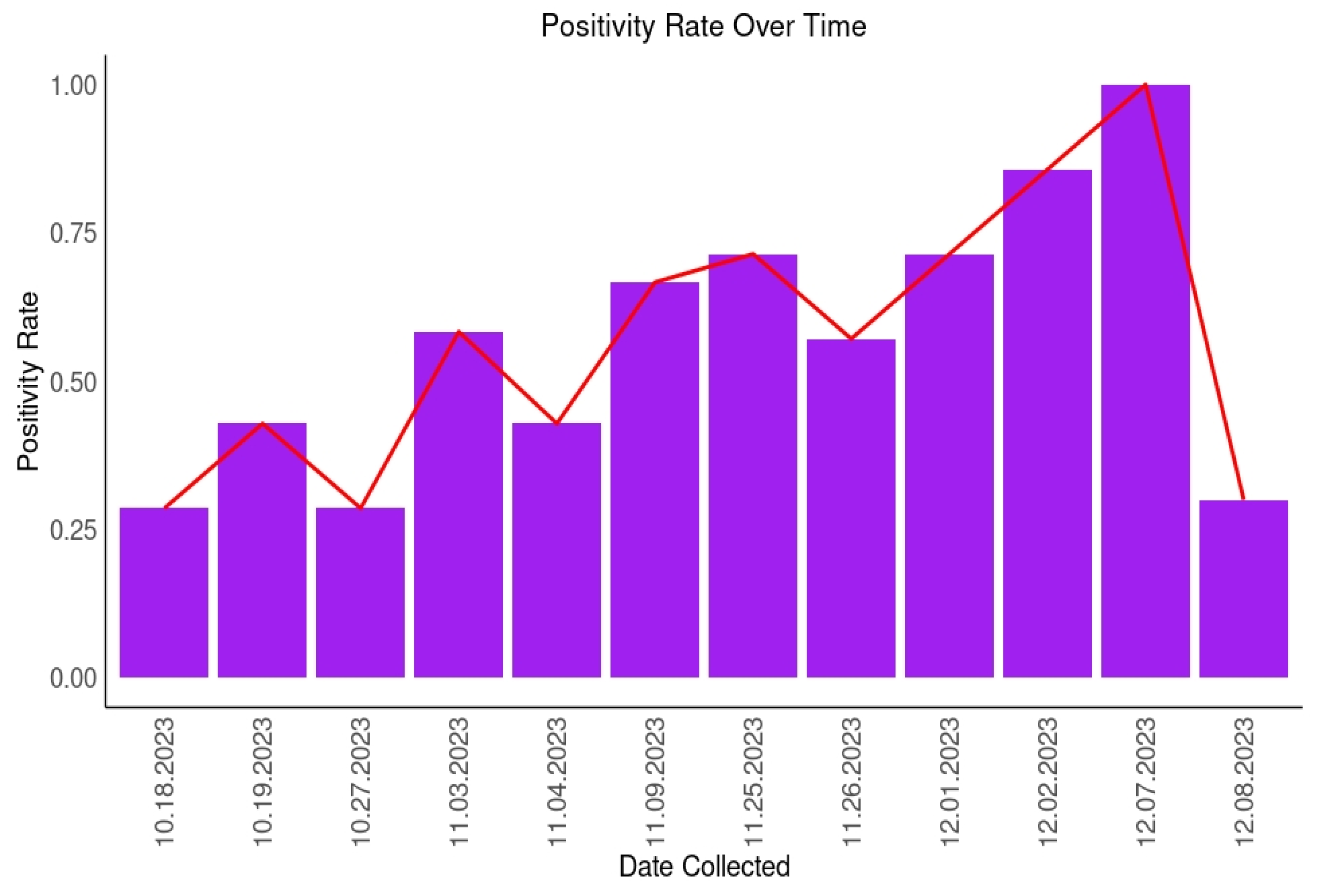
3.1. Detection of SARS-CoV-2
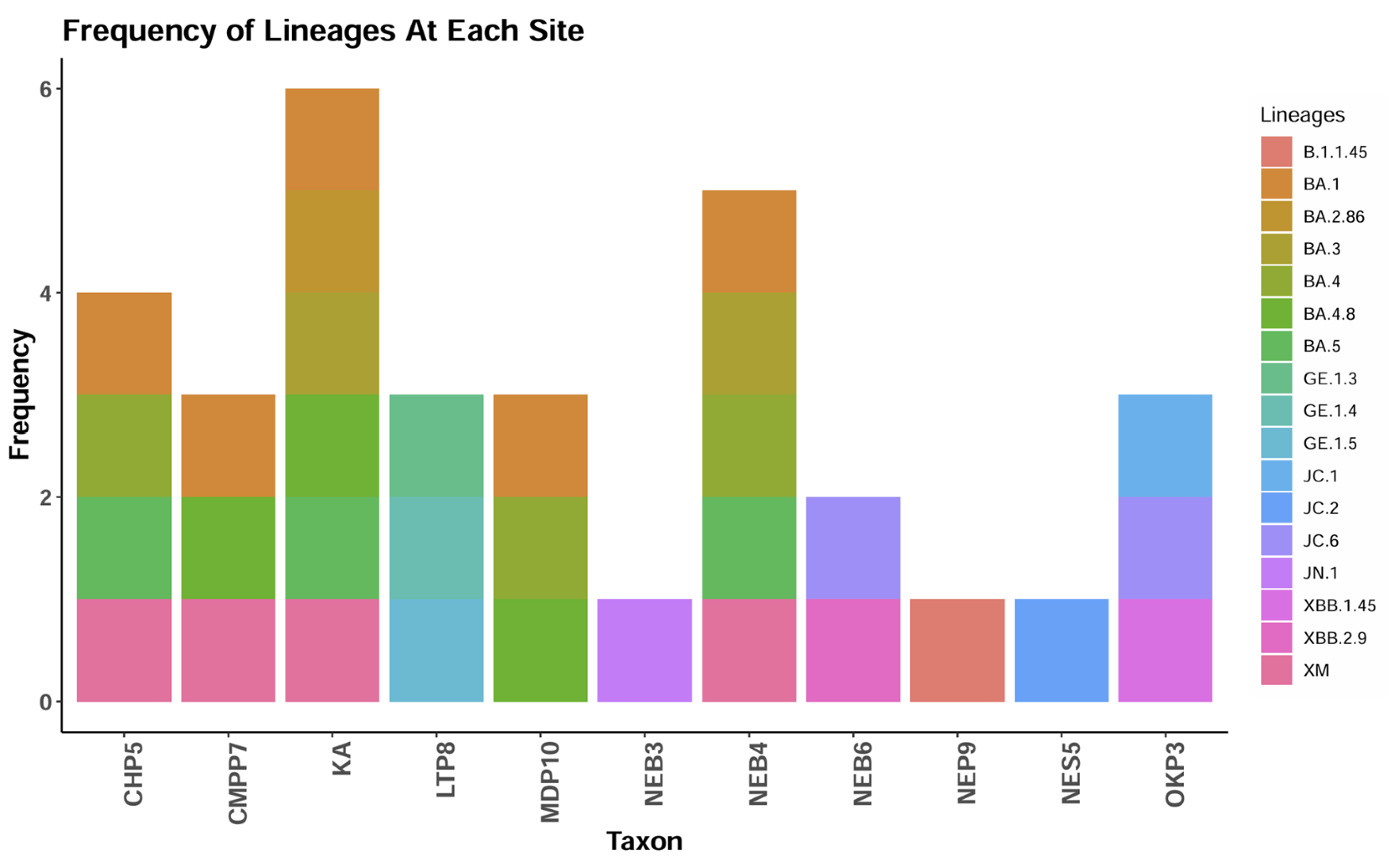
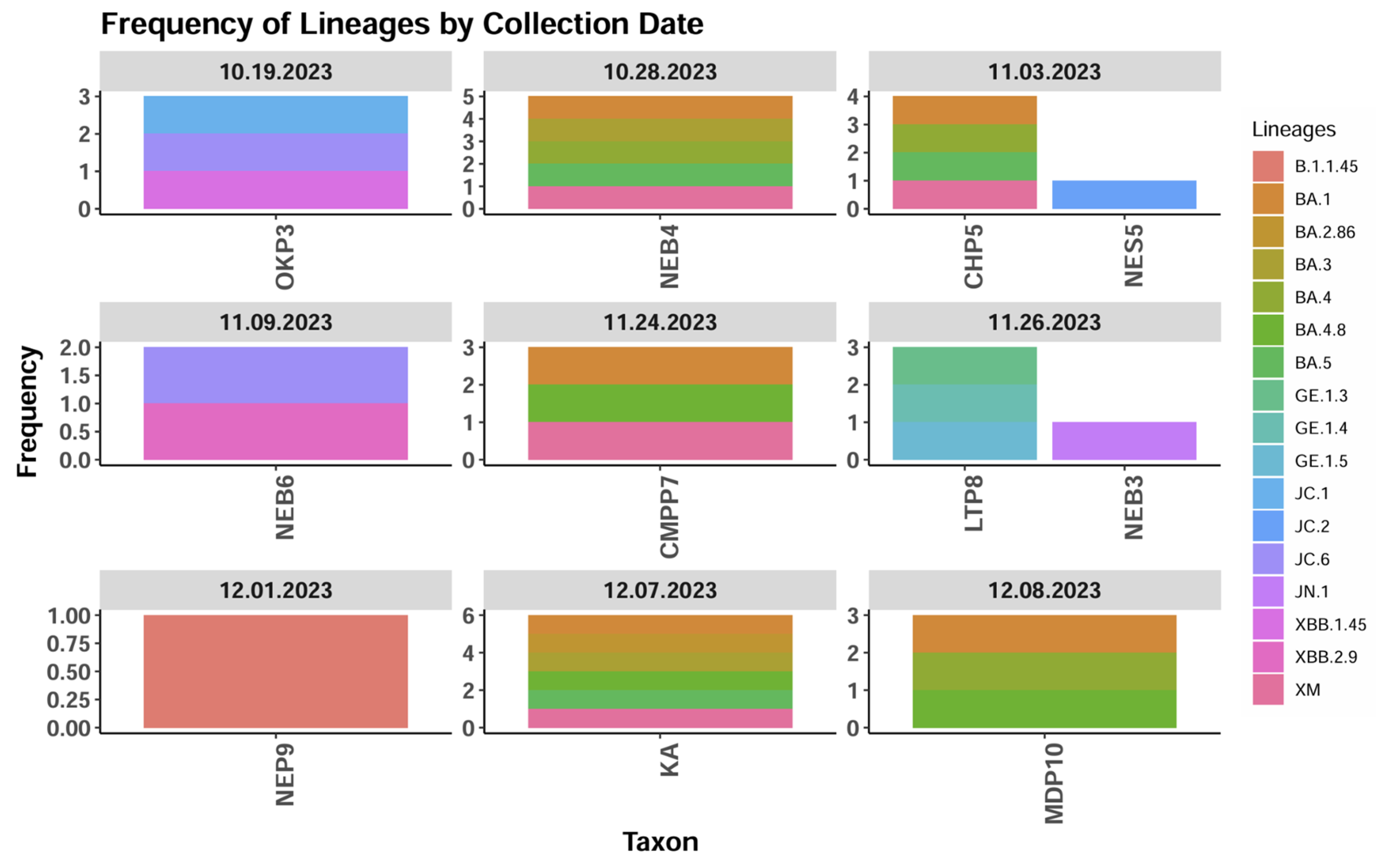
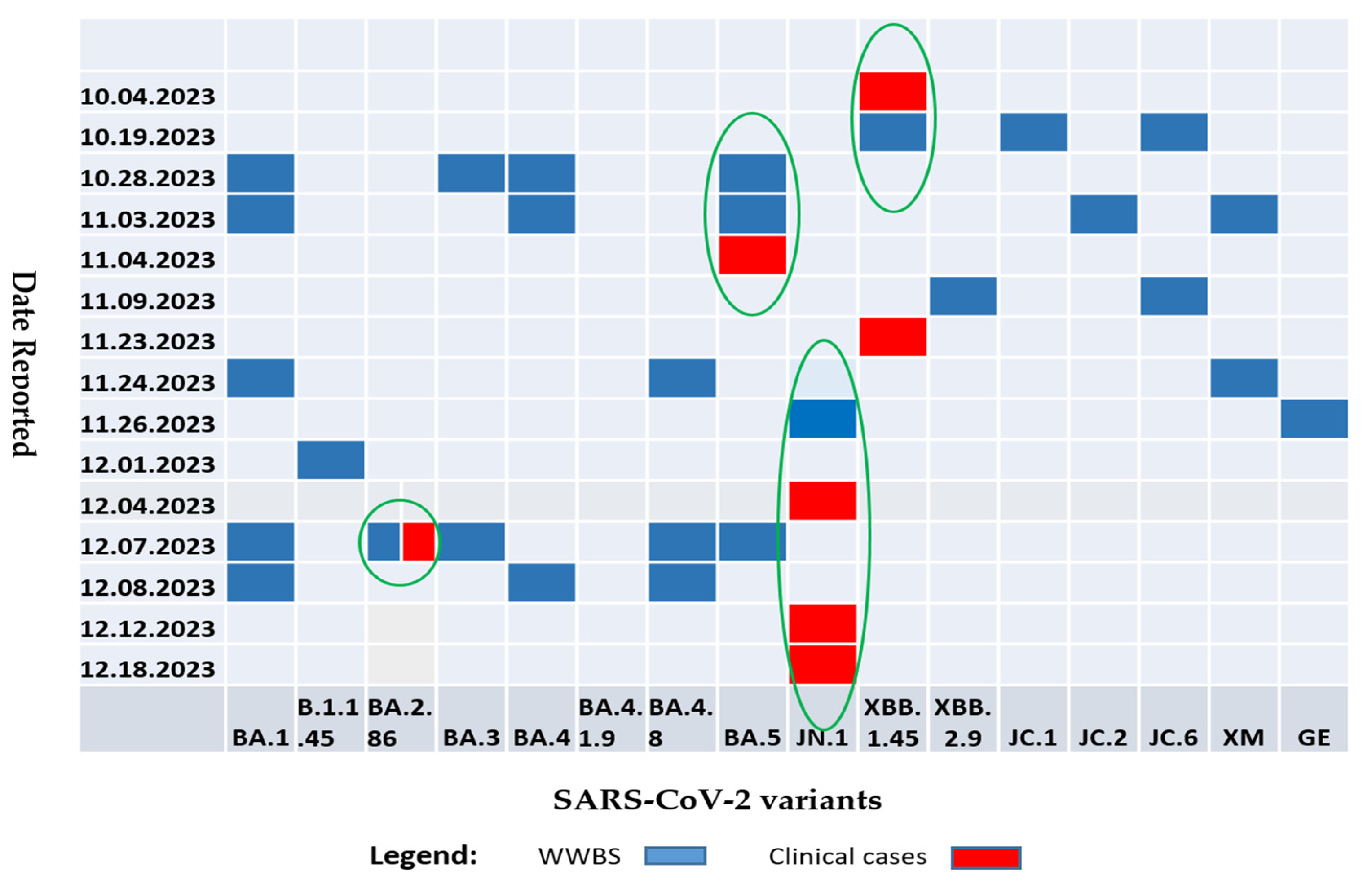
3.2. SARS-COV-2 Sequencing of Wastewater Samples
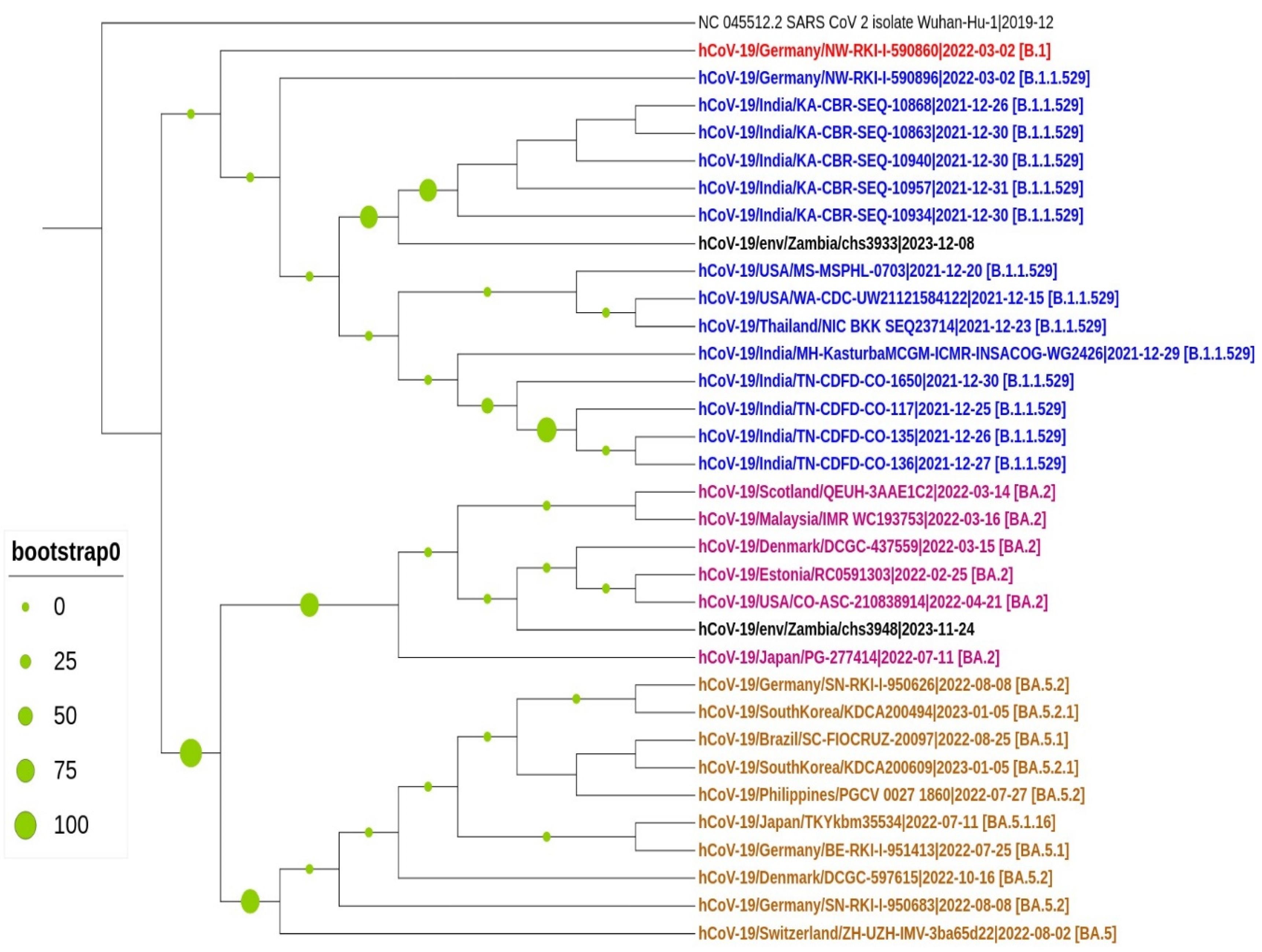
3.3. Mutation and Phylogenetic Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sharma A, Tiwari S, Deb MK, Marty JL. Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2): a global pandemic and treatment strategies. Int J Antimicrob Agents 2020;56:106054. [CrossRef]
- Liu YC, Kuo RL, Shih SR. COVID-19: The first documented coronavirus pandemic in history. Biomed J 2020;43:328–33. [CrossRef]
- Carabelli AM, Peacock TP, Thorne LG, Harvey WT, Hughes J, de Silva TI, et al. SARS-CoV-2 variant biology: immune escape, transmission and fitness. Nat Rev Microbiol 2023;21:162–77. [CrossRef]
- Bostanghadiri N, Ziaeefar P, Mofrad MG, Yousefzadeh P, Hashemi A, Darban-Sarokhalil D. COVID-19: An Overview of SARS-CoV-2 Variants—The Current Vaccines and Drug Development. Biomed Res Int 2023;2023:1879554. [CrossRef]
- Sah R, Rais MA, Mohanty A, Chopra H, Chandran D, Bin Emran T, et al. Omicron (B.1.1.529) variant and its subvariants and lineages may lead to another COVID-19 wave in the world? -An overview of current evidence and counteracting strategies. Int J Surg Open 2023;55:100625. [CrossRef]
- Cao Y, Yisimayi A, Jian F, Song W, Xiao T, Wang L, et al. BA.2.12.1, BA.4 and BA.5 escape antibodies elicited by Omicron infection. Nat 2022 6087923 2022;608:593–602. [CrossRef]
- Shah M, Woo HG. Omicron: A Heavily Mutated SARS-CoV-2 Variant Exhibits Stronger Binding to ACE2 and Potently Escapes Approved COVID-19 Therapeutic Antibodies. Front Immunol 2022;12:830527. [CrossRef]
- Cavanaugh AM, Fortier S, Lewis P, Arora V, Johnson M, George K, et al. COVID-19 Outbreak Associated with a SARS-CoV-2 R.1 Lineage Variant in a Skilled Nursing Facility After Vaccination Program — Kentucky, March 2021. MMWR Recomm Reports 2021;70:639–43. [CrossRef]
- Sarkar P, Banerjee S, Saha SA, Mitra P, Sarkar S. Genome surveillance of SARS-CoV-2 variants and their role in pathogenesis focusing on second wave of COVID-19 in India. Sci Rep 2023;13:4692. [CrossRef]
- Fernandes Q, Inchakalody VP, Merhi M, Mestiri S, Taib N, Moustafa Abo El-Ella D, et al. Emerging COVID-19 variants and their impact on SARS-CoV-2 diagnosis, therapeutics and vaccines. Ann Med 2022;54:524–40. [CrossRef]
- D’Agostino Y, Rocco T, Ferravante C, Porta A, Tosco A, Cappa VM, et al. Rapid and sensitive detection of SARS-CoV-2 variants in nasopharyngeal swabs and wastewaters. Diagn Microbiol Infect Dis 2022;102:115632. [CrossRef]
- Kevadiya BD, Machhi J, Herskovitz J, Oleynikov MD, Blomberg WR, Bajwa N, et al. Diagnostics for SARS-CoV-2 infections. Nat Mater 2021;20:593–605. [CrossRef]
- Tembo J, Egbe NF, Maluzi K, Mulonga K, Chilufya M, Kapata N, et al. Evaluation of SARS-CoV-2 diagnostics and risk factors associated with SARS-CoV-2 infection in Zambia. Int J Infect Dis 2022;120:150–7. [CrossRef]
- Peeling RW, Heymann DL, Teo YY, Garcia PJ. Diagnostics for COVID-19: moving from pandemic response to control. Lancet 2022;399:757–68. [CrossRef]
- Gangavarapu K, Latif AA, Mullen JL, Alkuzweny M, Hufbauer E, Tsueng G, et al. Outbreak.info genomic reports: scalable and dynamic surveillance of SARS-CoV-2 variants and mutations. Nat Methods 2023;20:512–22. [CrossRef]
- Amato E, Hyllestad S, Heradstveit P, Langlete P, Moen LV, Rohringer A, et al. Evaluation of the pilot wastewater surveillance for SARS-CoV-2 in Norway, June 2022–March 2023. BMC Public Health 2023;23:1714. [CrossRef]
- Brunner FS, Brown MR, Bassano I, Denise H, Khalifa MS, Wade MJ, et al. City-wide wastewater genomic surveillance through the successive emergence of SARS-CoV-2 Alpha and Delta variants. Water Res 2022;226:119306. [CrossRef]
- Flood MT, Sharp J, Bruggink J, Cormier M, Gomes B, Oldani I, et al. Understanding the efficacy of wastewater surveillance for SARS-CoV-2 in two diverse communities. PLoS One 2023;18:e0289343. [CrossRef]
- Conway MJ, Yang H, Revord LA, Novay MP, Lee RJ, Ward AS, et al. Chronic shedding of a SARS-CoV-2 Alpha variant in wastewater. BMC Genomics 2024;25:59. [CrossRef]
- Smith T, Cassell G, Bhatnagar A. Wastewater Surveillance Can Have a Second Act in COVID-19 Vaccine Distribution. JAMA Heal Forum 2021;2:E201616. [CrossRef]
- Wolfe M, Hughes B, Duong D, Chan-Herur V, Wigginton KR, White BJ, et al. Detection of SARS-CoV-2 Variants Mu, Beta, Gamma, Lambda, Delta, Alpha, and Omicron in Wastewater Settled Solids Using Mutation-Specific Assays Is Associated with Regional Detection of Variants in Clinical Samples. Appl Environ Microbiol 2022;88:e00045-22. [CrossRef]
- Ansari N, Kabir F, Khan W, Khalid F, Malik AA, Warren JL, et al. Environmental surveillance for COVID-19 using SARS-CoV-2 RNA concentration in wastewater—a study in District East, Karachi, Pakistan. Lancet Reg Heal—Southeast Asia 2024;20:100299. [CrossRef]
- Liu P, Ibaraki M, VanTassell J, Geith K, Cavallo M, Kann R, et al. A sensitive, simple, and low-cost method for COVID-19 wastewater surveillance at an institutional level. Sci Total Environ 2022;807:151047. [CrossRef]
- World Health Organization. Wastewater surveillance of SARS-CoV-2. World Heal Organ 2022:1–12.
- Kallem P, Hegab H, Alsafar H, Hasan SW, Banat F. SARS-CoV-2 detection and inactivation in water and wastewater: review on analytical methods, limitations and future research recommendations. Emerg Microbes Infect 2023;12:2222850. [CrossRef]
- Shafer MM, Bobholz MJ, Vuyk WC, Gregory DA, Roguet A, Haddock Soto LA, et al. Tracing the origin of SARS-CoV-2 omicron-like spike sequences detected in an urban sewershed: a targeted, longitudinal surveillance study of a cryptic wastewater lineage. The Lancet Microbe 2024;5:e335–44. [CrossRef]
- Jahn K, Dreifuss D, Topolsky I, Kull A, Ganesanandamoorthy P, Fernandez-Cassi X, et al. Early detection and surveillance of SARS-CoV-2 genomic variants in wastewater using COJAC. Nat Microbiol 2022;7:1151–60. [CrossRef]
- Fontenele RS, Kraberger S, Hadfield J, Driver EM, Bowes D, Holland LRA, et al. High-throughput sequencing of SARS-CoV-2 in wastewater provides insights into circulating variants. Water Res 2021;205:117710. [CrossRef]
- McClary-Gutierrez JS, Mattioli MC, Marcenac P, Silverman AI, Boehm AB, Bibby K, et al. SARS-CoV-2 wastewater surveillance for public health action. Emerg Infect Dis 2021;27:E1–9. [CrossRef]
- Maryam S, Ul Haq I, Yahya G, Ul Haq M, Algammal AM, Saber S, et al. COVID-19 surveillance in wastewater: An epidemiological tool for the monitoring of SARS-CoV-2. Front Cell Infect Microbiol 2023;12:978643. [CrossRef]
- Kirby AE, Walters MS, Jennings WC, Fugitt R, LaCross N, Mattioli M, et al. Using Wastewater Surveillance Data to Support the COVID-19 Response — United States, 2020-2021. MMWR Recomm Reports 2021;70:1242–4. [CrossRef]
- Yousif M, Rachida S, Taukobong S, Ndlovu N, Iwu-Jaja C, Howard W, et al. SARS-CoV-2 genomic surveillance in wastewater as a model for monitoring evolution of endemic viruses. Nat Commun 2023;14:6325. [CrossRef]
- Medema G, Heijnen L, Elsinga G, Italiaander R, Brouwer A. Presence of SARS-Coronavirus-2 RNA in Sewage and Correlation with Reported COVID-19 Prevalence in the Early Stage of the Epidemic in The Netherlands. Environ Sci Technol Lett 2020;7:511–6. [CrossRef]
- Ahmed W, Angel N, Edson J, Bibby K, Bivins A, O’Brien JW, et al. First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: A proof of concept for the wastewater surveillance of COVID-19 in the community. Sci Total Environ 2020;728:138764. [CrossRef]
- Mlejnkova H, Sovova K, Vasickova P, Ocenaskova V, Jasikova L, Juranova E. Preliminary study of Sars-Cov-2 occurrence in wastewater in the Czech Republic. Int J Environ Res Public Health 2020;17:5508. [CrossRef]
- La Rosa G, Iaconelli M, Mancini P, Bonanno Ferraro G, Veneri C, Bonadonna L, et al. First detection of SARS-CoV-2 in untreated wastewaters in Italy. Sci Total Environ 2020;736:139652. [CrossRef]
- Karthikeyan S, Nguyen A, McDonald D, Zong Y, Ronquillo N, Ren J, et al. Rapid, Large-Scale Wastewater Surveillance and Automated Reporting System Enable Early Detection of Nearly 85% of COVID-19 Cases on a University Campus. MSystems 2021;6:e0079321. [CrossRef]
- Bar-Or I, Weil M, Indenbaum V, Bucris E, Bar-Ilan D, Elul M, et al. Detection of SARS-CoV-2 variants by genomic analysis of wastewater samples in Israel. Sci Total Environ 2021;789:148002. [CrossRef]
- Albastaki A, Naji M, Lootah R, Almeheiri R, Almulla H, Almarri I, et al. First confirmed detection of SARS-COV-2 in untreated municipal and aircraft wastewater in Dubai, UAE: The use of wastewater based epidemiology as an early warning tool to monitor the prevalence of COVID-19. Sci Total Environ 2021;760:143350. [CrossRef]
- Street R, Malema S, Mahlangeni N, Mathee A. Wastewater surveillance for Covid-19: An African perspective. Sci Total Environ 2020;743:140719. [CrossRef]
- Duker EO, Obodai E, Addo SO, Kwasah L, Mensah ES, Gberbi E, et al. First Molecular Detection of SARS-CoV-2 in Sewage and Wastewater in Ghana. Biomed Res Int 2024;2024:9975781. [CrossRef]
- Simulundu E, Mupeta F, Chanda-Kapata P, Saasa N, Changula K, Muleya W, et al. First COVID-19 case in Zambia — Comparative phylogenomic analyses of SARS-CoV-2 detected in African countries. Int J Infect Dis 2021;102:455–9. [CrossRef]
- Chipimo PJ, Barradas DT, Kayeyi N, Zulu PM, Muzala K, Mazaba ML, et al. First 100 Persons with COVID-19 — Zambia, March 18–April 28, 2020. MMWR Morb Mortal Wkly Rep 2020;69:1547–8. [CrossRef]
- Katowa B, Kalonda A, Mubemba B, Matoba J, Shempela DM, Sikalima J, et al. Genomic Surveillance of SARS-CoV-2 in the Southern Province of Zambia: Detection and Characterization of Alpha, Beta, Delta, and Omicron Variants of Concern. Viruses 2022;14:1865. [CrossRef]
- Mwenda M, Saasa N, Sinyange N, Busby G, Chipimo PJ, Hendry J, et al. Detection of B.1.351 SARS-CoV-2 Variant Strain — Zambia, December 2020. MMWR Surveill Summ 2021;70:280–2. [CrossRef]
- Saasa N, M’kandawire E, Ndebe J, Mwenda M, Chimpukutu F, Mukubesa AN, et al. Detection of Human Adenovirus and Rotavirus in Wastewater in Lusaka, Zambia: Demonstrating the Utility of Environmental Surveillance for the Community. Pathogens 2024;13:486. [CrossRef]
- Prata C, Ribeiro A, Cunha Â, Gomes NCM, Almeida A. Ultracentrifugation as a direct method to concentrate viruses in environmental waters: virus-like particle enumeration as a new approach to determine the efficiency of recovery. J Environ Monit 2012;14:64–70. [CrossRef]
- Fagnant-Sperati CS, Ren Y, Zhou NA, Komen E, Mwangi B, Hassan J, et al. Validation of the bag-mediated filtration system for environmental surveillance of poliovirus in Nairobi, Kenya. J Appl Microbiol 2021;130:971–81. [CrossRef]
- Philo SE, Keim EK, Swanstrom R, Ong AQW, Burnor EA, Kossik AL, et al. A comparison of SARS-CoV-2 wastewater concentration methods for environmental surveillance. Sci Total Environ 2021;760:144215. [CrossRef]
- Sievers F, Higgins DG. The Clustal Omega Multiple Alignment Package. Methods Mol. Biol., vol. 2231, Humana, New York, NY; 2021, p. 3–16. [CrossRef]
- Khandia R, Singhal S, Alqahtani T, Kamal MA, El-Shall NA, Nainu F, et al. Emergence of SARS-CoV-2 Omicron (B.1.1.529) variant, salient features, high global health concerns and strategies to counter it amid ongoing COVID-19 pandemic. Environ Res 2022;209:112816. [CrossRef]
- Shrestha S, Malla B, Angga MS, Sthapit N, Raya S, Hirai S, et al. Long-term SARS-CoV-2 surveillance in wastewater and estimation of COVID-19 cases: An application of wastewater-based epidemiology. Sci Total Environ 2023;896:165270. [CrossRef]
- Fernández-de-Mera IG, Rodríguez del-Río FJ, de la Fuente J, Pérez-Sancho M, Hervás D, Moreno I, et al. Detection of environmental SARS-CoV-2 RNA in a high prevalence setting in Spain. Transbound Emerg Dis 2021;68:1487–92. [CrossRef]
- World Health Organization. Environmental surveillance for SARS-COV-2 to complement public health surveillance—Interim Guidance. 2022.
- Krogsgaard LW, Benedetti G, Gudde A, Richter SR, Rasmussen LD, Midgley SE, et al. Results from the SARS-CoV-2 wastewater-based surveillance system in Denmark, July 2021 to June 2022. Water Res 2024;252:121223. [CrossRef]
- Wu F, Xiao A, Zhang J, Moniz K, Endo N, Armas F, et al. SARS-CoV-2 RNA concentrations in wastewater foreshadow dynamics and clinical presentation of new COVID-19 cases. Sci Total Environ 2022;805:150121. [CrossRef]
- Gupta P, Liao S, Ezekiel M, Novak N, Rossi A, LaCross N, et al. Wastewater Genomic Surveillance Captures Early Detection of Omicron in Utah. Microbiol Spectr 2023;11:e0039123. [CrossRef]
- Zhang D, Duran SSF, Lim WYS, Tan CKI, Cheong WCD, Suwardi A, et al. SARS-CoV-2 in wastewater: From detection to evaluation. Mater Today Adv 2022;13:100211. [CrossRef]
- Wolter N, Jassat W, Walaza S, Welch R, Moultrie H, Groome M, et al. Early assessment of the clinical severity of the SARS-CoV-2 omicron variant in South Africa: a data linkage study. Lancet 2022;399:437–46. [CrossRef]
- Viana R, Moyo S, Amoako DG, Tegally H, Scheepers C, Althaus CL, et al. Rapid epidemic expansion of the SARS-CoV-2 Omicron variant in Southern Africa. Nature 2022;603:679–86. [CrossRef]
- Tegally H, Wilkinson E, Giovanetti M, Iranzadeh A, Fonseca V, Giandhari J, et al. Detection of a SARS-CoV-2 variant of concern in South Africa. Nature 2021;592:438–43. [CrossRef]
- Shempela DM, Chambaro HM, Sikalima J, Cham F, Njuguna M, Morrison L, et al. Detection and Characterisation of SARS-CoV-2 in Eastern Province of Zambia; A Retrospective Genomic Surveillance Study. Int J Mol Sci 2024;25:6338. [CrossRef]
- Liddor Naim M, Fu Y, Shagan M, Bar-Or I, Marks R, Sun Q, et al. The Rise and Fall of Omicron BA.1 Variant as Seen in Wastewater Supports Epidemiological Model Predictions. Viruses 2023;15:1862. [CrossRef]
- Santiago GA, Volkman HR, Flores B, González GL, Charriez KN, Huertas LC, et al. SARS-CoV-2 Omicron Replacement of Delta as Predominant Variant, Puerto Rico. Emerg Infect Dis 2023;29:855–7. [CrossRef]
- Robles-Escajeda E, Mohl JE, Contreras L, Betancourt AP, Mancera BM, Kirken RA, et al. Rapid Shift from SARS-CoV-2 Delta to Omicron Sub-Variants within a Dynamic Southern U.S. Borderplex. Viruses 2023;15:658. [CrossRef]
- Aguayo-Acosta A, Oyervides-Muñoz MA, Rodriguez-Aguillón KO, Ovalle-Carcaño A, Romero-Castillo KD, Robles-Zamora A, et al. Omicron and Delta variant prevalence detection and identification during the fourth COVID-19 wave in Mexico using wastewater-based epidemiology. IJID Reg 2024;10:44–51. [CrossRef]
- Zabidi NZ, Liew HL, Farouk IA, Puniyamurti A, Yip AJW, Wijesinghe VN, et al. Evolution of SARS-CoV-2 Variants: Implications on Immune Escape, Vaccination, Therapeutic and Diagnostic Strategies. Viruses 2023;15:944. [CrossRef]
- Tao K, Tzou PL, Nouhin J, Gupta RK, de Oliveira T, Kosakovsky Pond SL, et al. The biological and clinical significance of emerging SARS-CoV-2 variants. Nat Rev Genet 2021;22:757–73. [CrossRef]
- Wilhelm A, Agrawal S, Schoth J, Meinert-Berning C, Bastian D, Orschler L, et al. Early Detection of SARS-CoV-2 Omicron BA.4 and BA.5 in German Wastewater. Viruses 2022;14. [CrossRef]
- Baldovin T, Amoruso I, Fonzo M, Bertoncello C, Groppi V, Pitter G, et al. Trends in SARS-CoV-2 clinically confirmed cases and viral load in wastewater: A critical alignment for Padua city (NE Italy). Heliyon 2023;9:e20571. [CrossRef]
- Triggiano F, De Giglio O, Apollonio F, Brigida S, Fasano F, Mancini P, et al. Wastewater-based Epidemiology and SARS-CoV-2: Variant Trends in the Apulia Region (Southern Italy) and Effect of Some Environmental Parameters. Food Environ Virol 2023;15:331–41. [CrossRef]
- La Rosa G, Brandtner D, Bonanno Ferraro G, Veneri C, Mancini P, Iaconelli M, et al. Wastewater surveillance of SARS-CoV-2 variants in October–November 2022 in Italy: detection of XBB.1, BA.2.75 and rapid spread of the BQ.1 lineage. Sci Total Environ 2023;873:162339. [CrossRef]
- Qvesel AG, Bennedbæk M, Larsen NB, Gunalan V, Krogsgaard LW, Rasmussen M, et al. SARS-CoV-2 Variants BQ.1 and XBB.1.5 in Wastewater of Aircraft Flying from China to Denmark, 2023. Emerg Infect Dis 2023;29:2559–61. [CrossRef]
- Lambrou AS, South E, Ballou ES, Paden CR, Fuller JA, Bart SM, et al. Early Detection and Surveillance of the SARS-CoV-2 Variant BA.2.86 — Worldwide, July–October 2023. MMWR Recomm Reports 2023;72:1162–7. [CrossRef]
- Rasmussen M, Moller FT, Gunalan V, Baig S, Bennedbak M, Christiansen LE, et al. First cases of SARS-CoV-2 BA.2.86 in Denmark, 2023. Eurosurveillance 2023;28:2300460. [CrossRef]
- Wannigama DL, Amarasiri M, Phattharapornjaroen P, Hurst C, Modchang C, Chadsuthi S, et al. Increased faecal shedding in SARS-CoV-2 variants BA.2.86 and JN.1. Lancet Infect Dis 2024;24:e348–50. [CrossRef]
- Espinosa-Gongora C, Berg C, Rehn M, Varg JE, Dillner L, Latorre-Margalef N, et al. Early detection of the emerging SARS-CoV-2 BA.2.86 lineage through integrated genomic surveillance of wastewater and COVID-19 cases in Sweden, weeks 31 to 38 2023. Eurosurveillance 2023;28. [CrossRef]
- Wannigama DL, Amarasiri M, Phattharapornjaroen P, Hurst C, Modchang C, Chadsuthi S, et al. Tracing the new SARS-CoV-2 variant BA.2.86 in the community through wastewater surveillance in Bangkok, Thailand. Lancet Infect Dis 2023;23:e464–6. [CrossRef]
- Du C, Peng Y, Lyu Z, Yue Z, Fu Y, Yao X, et al. Early Detection of the Emerging SARS-CoV-2 BA.2.86 Lineage Through Wastewater Surveillance Using a Mediator Probe PCR Assay — Shenzhen City, Guangdong Province, China, 2023. China CDC Wkly 2024;6:332–8. [CrossRef]
- Erster O, Bar-Or I, Azar R, Assraf H, Kabat A, Mannasse B, et al. Incursion of SARS-CoV-2 BA.2.86.1 variant into Israel: National-scale wastewater surveillance using a novel quantitative real-time PCR assay. Sci Total Environ 2024;933:173164. [CrossRef]
- Planas D, Bruel T, Staropoli I, Guivel-Benhassine F, Porrot F, Maes P, et al. Resistance of Omicron subvariants BA.2.75.2, BA.4.6, and BQ.1.1 to neutralizing antibodies. Nat Commun 2023;14:1–11. [CrossRef]
- World Health Organization. Initial rapid evaluation of JN.1. vol. 1. 2023.
- Li P, Liu Y, Faraone JN, Hsu CC, Chamblee M, Zheng Y-M, et al. Distinct patterns of SARS-CoV-2 BA.2.87.1 and JN.1 variants in immune evasion, antigenicity, and cell-cell fusion. MBio 2024;15:e00751-24. [CrossRef]
- Karyakarte RP, Das R, Rajmane M V, Dudhate S, Agarasen J, Pillai P, et al. Appearance and Prevalence of JN.1 SARS-CoV-2 Variant in India and Its Clinical Profile in the State of Maharashtra. Cureus 2024;16:e56718. [CrossRef]
- Bartel A, Grau JH, Bitzegeio J, Werber D, Linzner N, Schumacher V, et al. Timely Monitoring of SARS-CoV-2 RNA Fragments in Wastewater Shows the Emergence of JN.1 (BA.2.86.1.1, Clade 23I) in Berlin, Germany. Viruses 2024;16:102. [CrossRef]
| Site (n) | Positive n (%) | 95% CI |
|---|---|---|
| Chambishi (n=18) | 7(39%) | 17.3-64.3 |
| Chipata Motel Ponds (13) | 2(15%) | 1.9-45.4 |
| Chipata Motel Pumps (20) | 3(15%) | 3.2-37.9 |
| Lubuto (18) | 6(33%) | 13.3-59.0 |
| Mindolo (18) | 11(61%) | 35.7-82.7 |
| New Kanini (26) | 12(46%) | 26.6-66.6 |
| Nkana East (26) | 16(62%) | 40.6-79.8 |
| Old Kanini (16) | 5(31%) | 11.0-58.7 |
| Total (155) | 62(40%) | 32.2-48.2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




