Submitted:
10 October 2024
Posted:
11 October 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
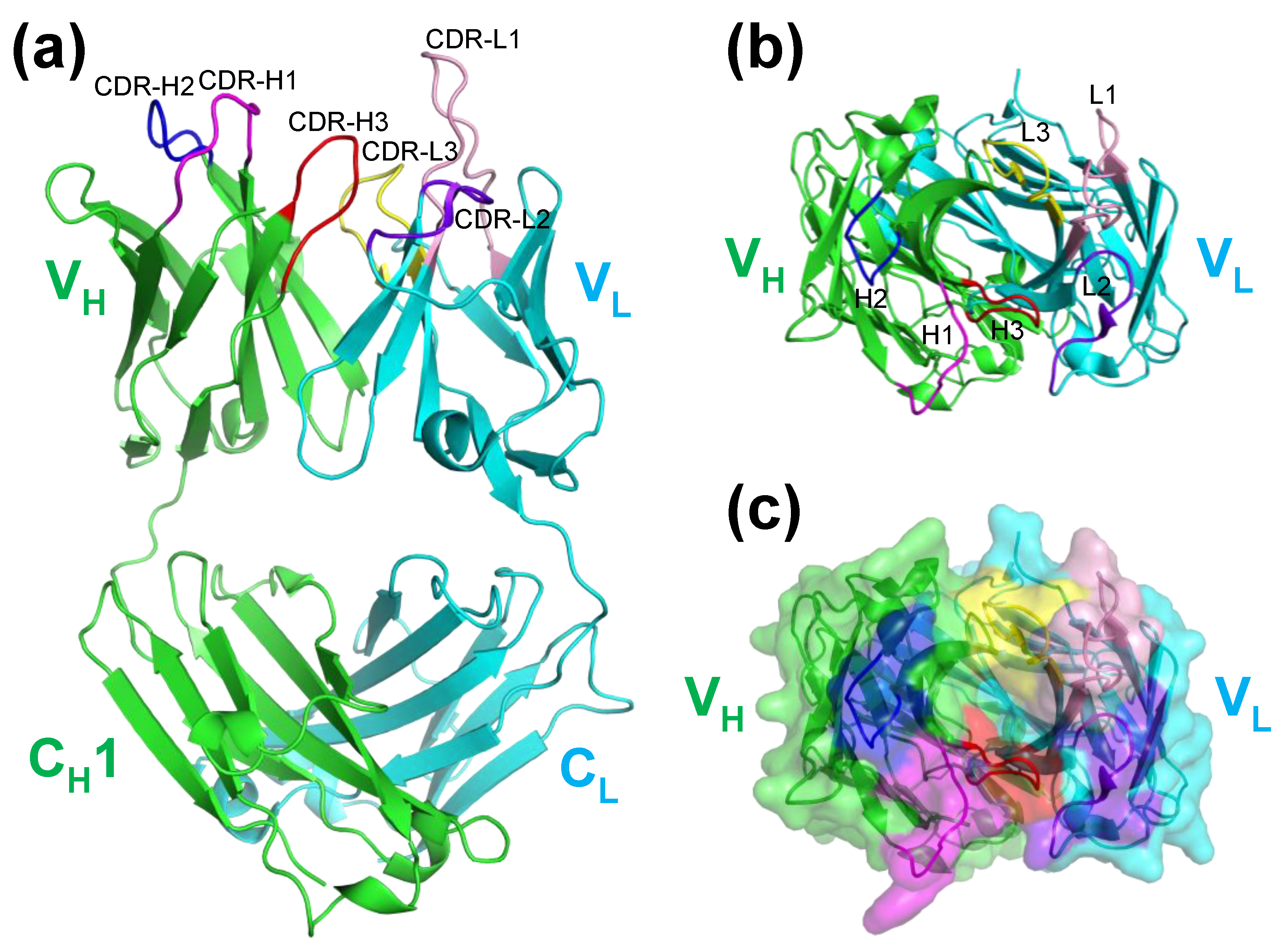
2.1. Crystal Structure of KTM219 Fab
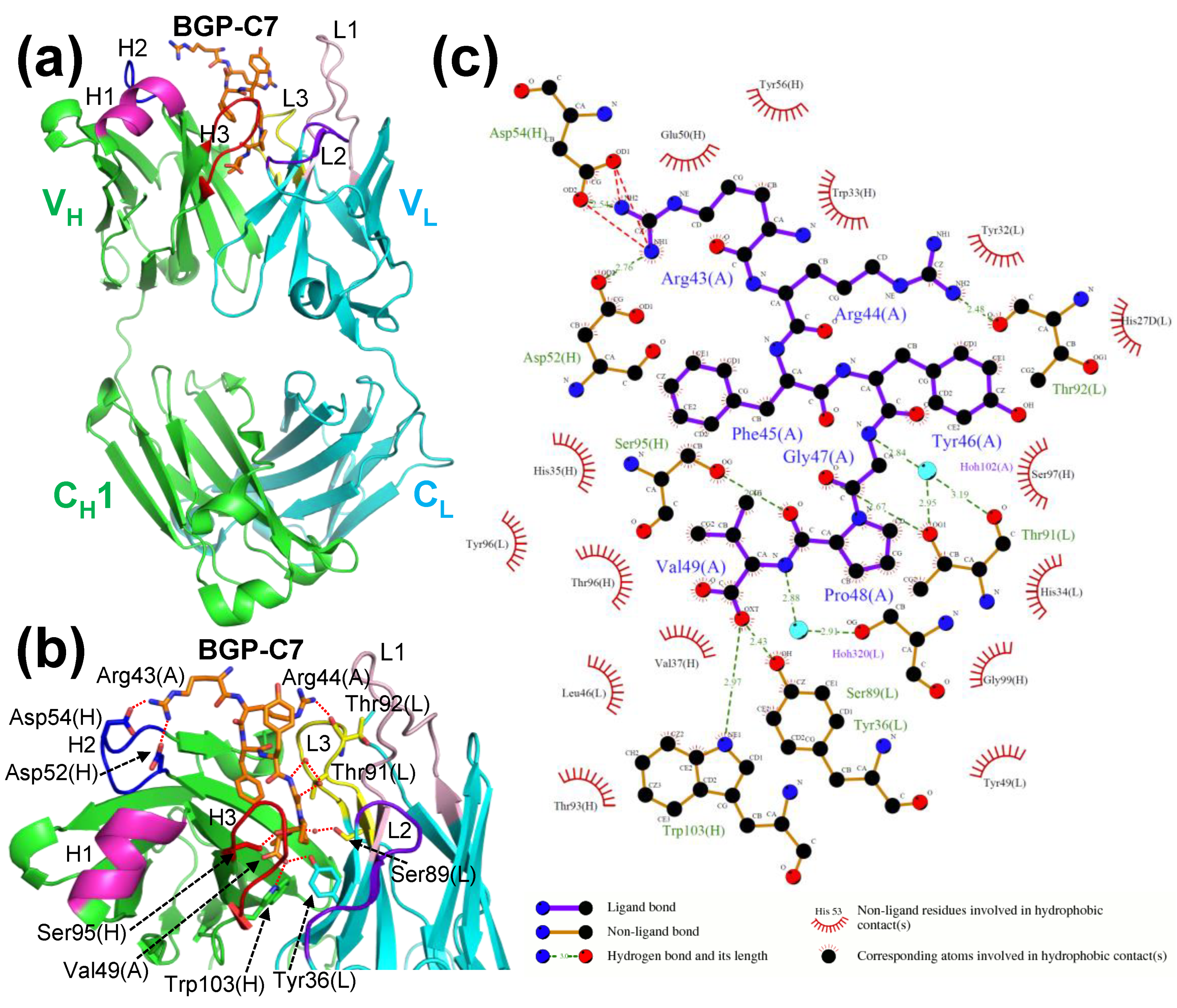
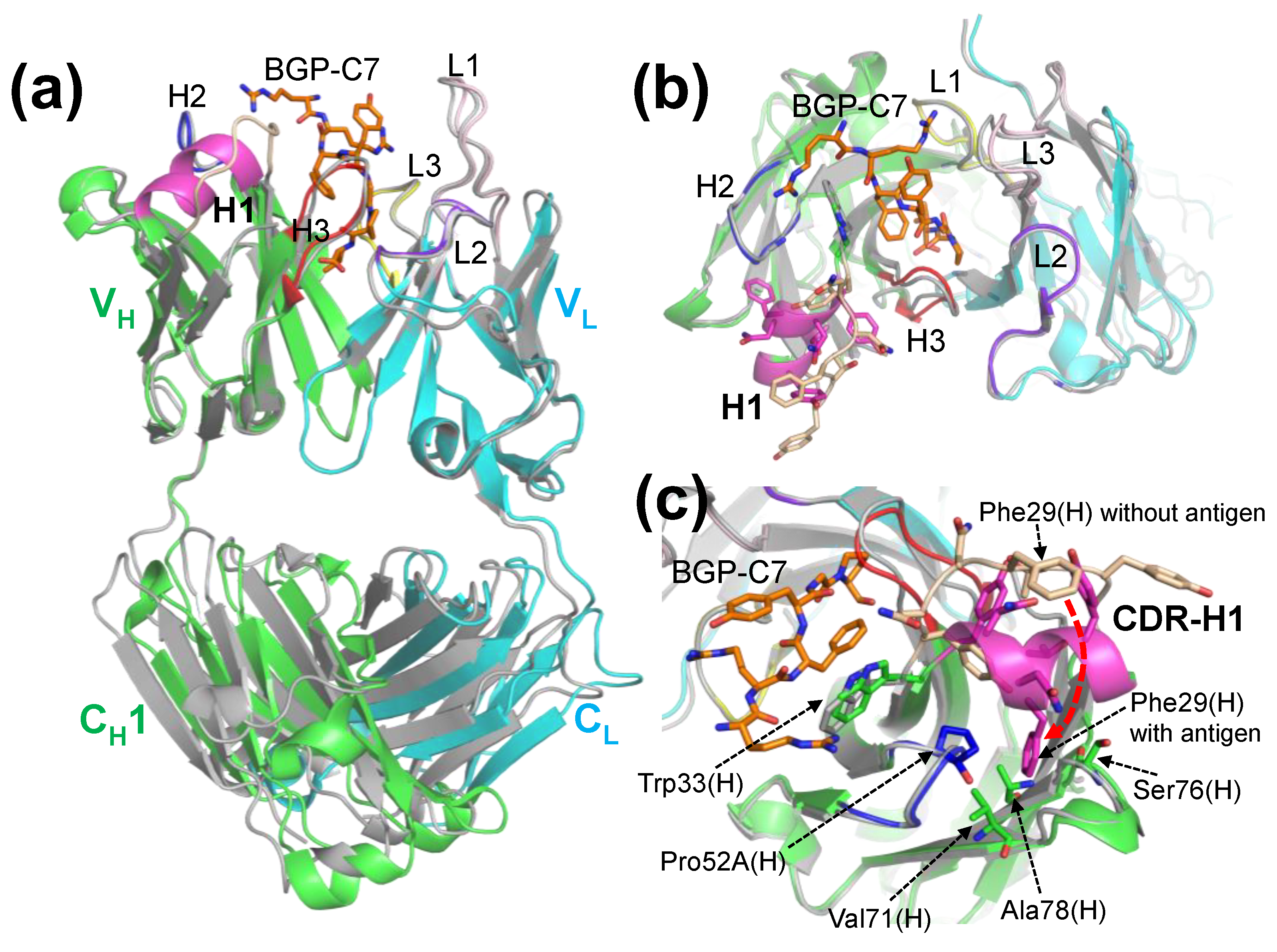
2.2. Crystal Structure of Antibody-Antigen Complex of KTM219 Fab
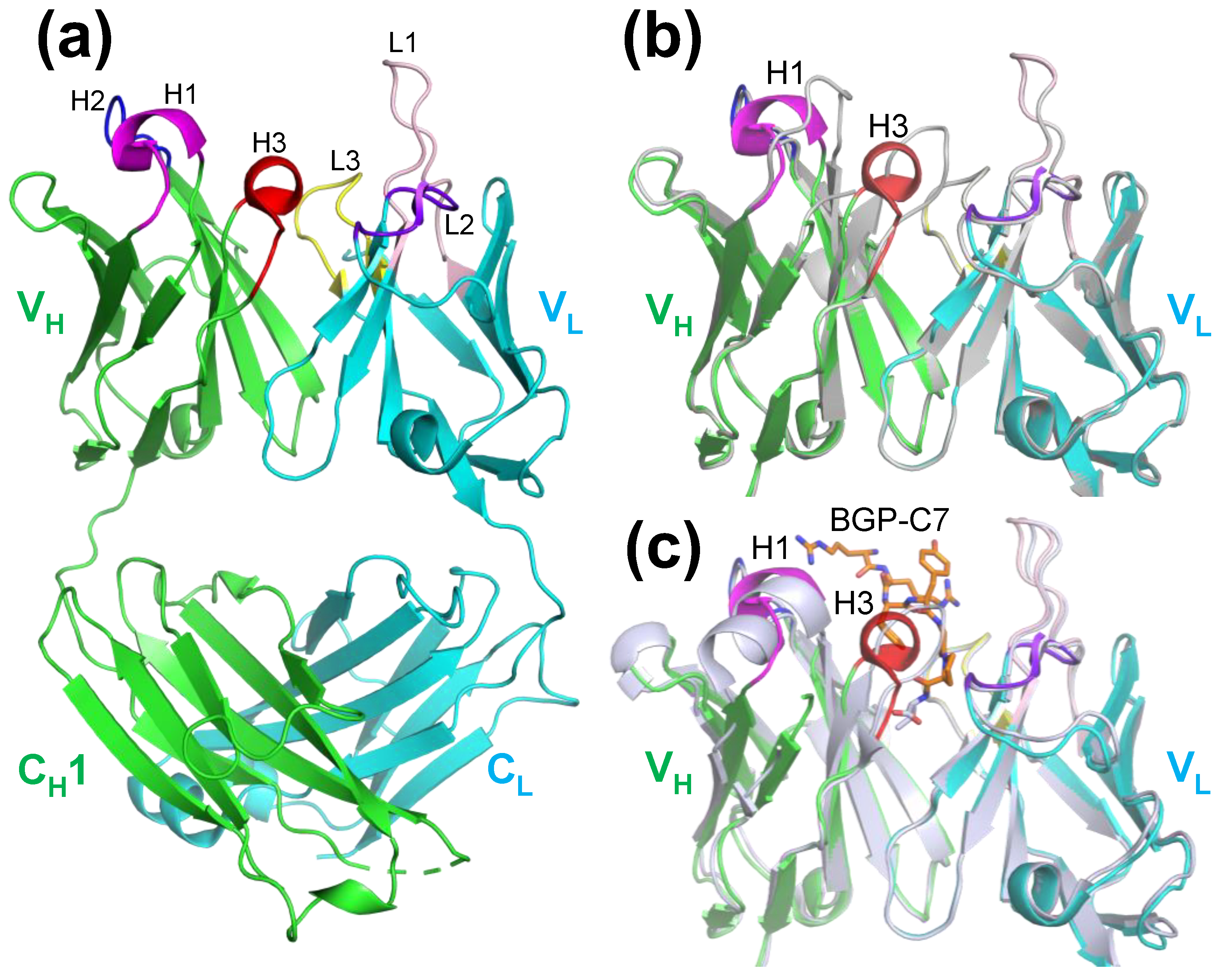
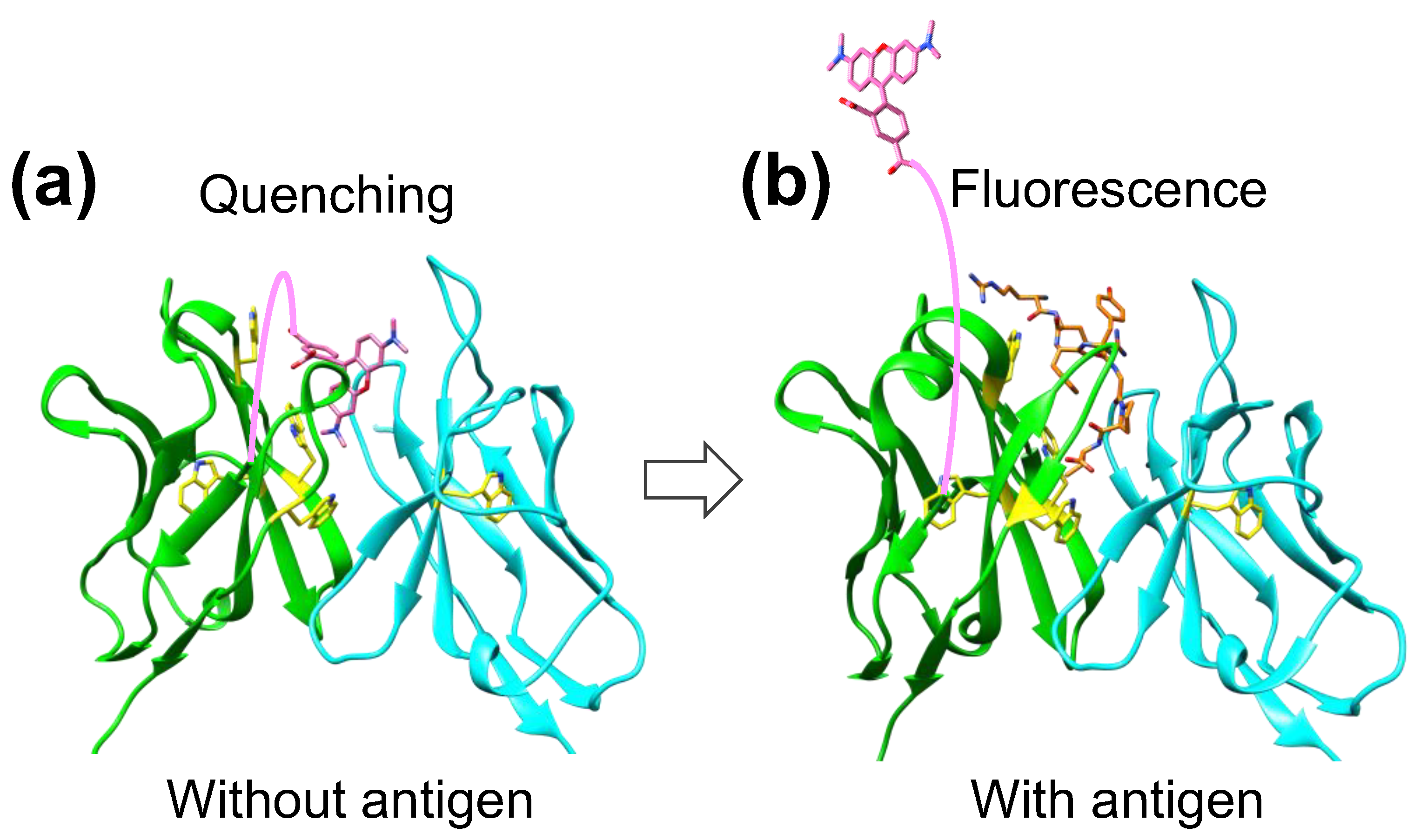
2.3. Structure of KTM219 Fab Crystal Grown in the Presence of a Fluorescent Dye TAMRA
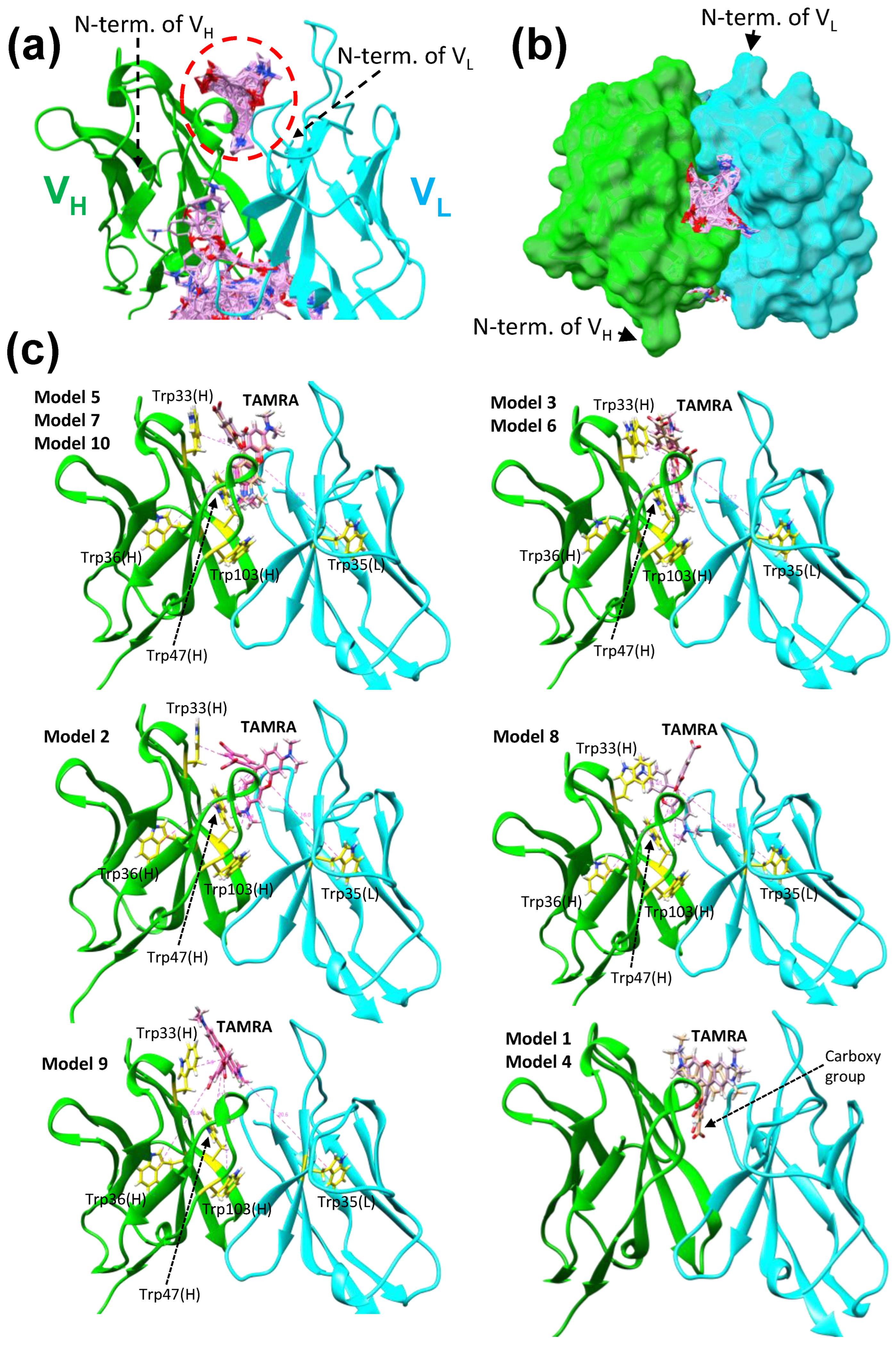
2.4. Docking Simulations of a Fluorescent Dye TAMRA to KTM219 Fab
3. Discussion
4. Materials and Methods
4.1. Protein Expression and Purification
4.2. Crystallization
4.3. Data Collection, Structure Determination and Refinement
4.4. Docking Simulations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Nan, X.; Yang, L.; Cui, Y. Lateral flow immunoassay for proteins. Clin Chim Acta 2023, 544, 117337. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Li, Q.; Wang, X.; Liu, X.; Zhang, Y.; Li, R.; Guo, J.; Zhang, G. Lateral flow immunoassays for antigens, antibodies and haptens detection. Int J Biol Macromol 2023, 242, 125186. [Google Scholar] [CrossRef] [PubMed]
- Spicuzza, L.; Campagna, D.; Di Maria, C.; Sciacca, E.; Mancuso, S.; Vancheri, C.; Sambataro, G. An update on lateral flow immunoassay for the rapid detection of SARS-CoV-2 antibodies. AIMS Microbiol 2023, 9, 375–401. [Google Scholar] [CrossRef] [PubMed]
- Rani, A. Q.; Zhu, B.; Ueda, H.; Kitaguchi, T. Recent progress in homogeneous immunosensors based on fluorescence or bioluminescence using antibody engineering. Analyst 2023, 148, 1422–1429. [Google Scholar] [CrossRef]
- Abe, R.; Ohashi, H.; Iijima, I.; Ihara, M.; Takagi, H.; Hohsaka, T.; Ueda, H. "Quenchbodies": quench-based antibody probes that show antigen-dependent fluorescence. J. Am. Chem. Soc. 2011, 133, 17386–17394. [Google Scholar] [CrossRef]
- Abe, R.; Jeong, H. J.; Arakawa, D.; Dong, J.; Ohashi, H.; Kaigome, R.; Saiki, F.; Yamane, K.; Takagi, H.; Ueda, H. Ultra Q-bodies: quench-based antibody probes that utilize dye-dye interactions with enhanced antigen-dependent fluorescence. Sci. Rep. 2014, 4, 4640. [Google Scholar] [CrossRef]
- Dong, J.; Ueda, H. Recent Advances in Quenchbody, a Fluorescent Immunosensor. Sensors 2021, 21. [Google Scholar] [CrossRef]
- Jeong, H. J. Quenchbodies That Enable One-Pot Detection of Antigens: A Structural Perspective. Bioengineering (Basel) 2023, 10. [Google Scholar] [CrossRef]
- Huynh Nhat, K. P.; Watanabe, T.; Yoshikoshi, K.; Hohsaka, T. Antibody-based fluorescent and fluorescent ratiometric indicators for detection of phosphotyrosine. J. Biosci. Bioeng. 2016, 122, 146–154. [Google Scholar] [CrossRef]
- Yoshinari, T.; Ohashi, H.; Abe, R.; Kaigome, R.; Ohkawa, H.; Sugita-Konishi, Y. Development of a rapid method for the quantitative determination of deoxynivalenol using Quenchbody. Anal. Chim. Acta 2015, 888, 126. [Google Scholar] [CrossRef]
- Tsujikawa, K.; Saiki, F.; Yamamuro, T.; Iwata, Y. T.; Abe, R.; Ohashi, H.; Kaigome, R.; Yamane, K.; Kuwayama, K.; Kanamori, T.; Inoue, H. Development of a novel immunoassay for herbal cannabis using a new fluorescent antibody probe, "Ultra Quenchbody". Forensic Sci. Int. 2016, 266, 541–548. [Google Scholar] [CrossRef] [PubMed]
- Marmé, N.; Knemeyer, J. P.; Sauer, M.; Wolfrum, J. Inter- and intramolecular fluorescence quenching of organic dyes by tryptophan. Bioconjugate Chem. 2003, 14, 1133–1139. [Google Scholar] [CrossRef]
- Vaiana, A. C.; Neuweiler, H.; Schulz, A.; Wolfrum, J.; Sauer, M.; Smith, J. C. Fluorescence quenching of dyes by tryptophan: interactions at atomic detail from combination of experiment and computer simulation. J. Am. Chem. Soc. 2003, 125, 14564–14572. [Google Scholar] [CrossRef] [PubMed]
- Lim, S. L.; Ichinose, H.; Shinoda, T.; Ueda, H. Noncompetitive detection of low molecular weight peptides by open sandwich immunoassay. Anal. Chem. 2007, 79, 6193–6200. [Google Scholar] [CrossRef]
- Ohashi, H.; Matsumoto, T.; Jeong, H. J.; Dong, J.; Abe, R.; Ueda, H. Insight into the Working Mechanism of Quenchbody: Transition of the Dye around Antibody Variable Region That Fluoresces upon Antigen Binding. Bioconjug Chem. 2016, 27, 2248–2253. [Google Scholar] [CrossRef]
- Li, J.; Zhang, H.; Yang, C.; Li, Y.; Dai, Z. An overview of osteocalcin progress. J Bone Miner Metab 2016, 34, 367–379. [Google Scholar] [CrossRef]
- Teplyakov, A.; Obmolova, G.; Malia, T. J.; Luo, J.; Gilliland, G. L. Structural evidence for a constrained conformation of short CDR-L3 in antibodies. Proteins 2014, 82, 1679–1683. [Google Scholar] [CrossRef]
- Laskowski, R. A.; Swindells, M. B. LigPlot+: multiple ligand-protein interaction diagrams for drug discovery. J. Chem. Inf. Model 2011, 51, 2778–2786. [Google Scholar] [CrossRef]
- Kabat, E. A.; Wu, T. T.; Perry, H. M.; Gottesman, K. S.; Foeller, C. , Sequences of Proteins of Immunological Interest., 5th ed.; U.S. Government Printing Office: Bethesda, MD, USA, 1991. [Google Scholar]
- Grosdidier, A.; Zoete, V.; Michielin, O. SwissDock, a protein-small molecule docking web service based on EADock DSS. Nucleic Acids Res. 2011, 39, W270–277. [Google Scholar] [CrossRef]
- Grosdidier, A.; Zoete, V.; Michielin, O. Fast docking using the CHARMM force field with EADock DSS. J. Comput. Chem. 2011, 32, 2149–2159. [Google Scholar] [CrossRef]
- Moretti, R.; Lyskov, S.; Das, R.; Meiler, J.; Gray, J. J. Web-accessible molecular modeling with Rosetta: The Rosetta Online Server that Includes Everyone (ROSIE). Protein Sci. 2018, 27, 259–268. [Google Scholar] [CrossRef] [PubMed]
- Hush, N. S.; Paddon-Row, M. N.; Cotsaris, E.; Oevering, H.; Verhoeven, J. W.; Heppener, M. Distance dependence of photoinduced electron transfer through non-conjugated bridges. Chem Phys Lett 1985, 117, 8–11. [Google Scholar] [CrossRef]
- Dong, J.; Jeong, H. J.; Ueda, H. Preparation of Quenchbodies by protein transamination reaction. J. Biosci. Bioeng. 2016, 122, 125–130. [Google Scholar] [CrossRef]
- Stewart, P. D. S.; Kolek, S. A.; Briggs, R. A.; Chayen, N. E.; Baldock, P. F. M. Random Microseeding: A Theoretical and Practical Exploration of Seed Stability and Seeding Techniques for Successful Protein Crystallization. Cryst. Growth Des. 2011, 11, 3432–3441. [Google Scholar] [CrossRef]
- Otwinowski, Z.; Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 1997, 276, 307–326. [Google Scholar] [CrossRef]
- Emsley, P.; Lohkamp, B.; Scott, W. G.; Cowtan, K. Features and development of Coot. Acta Crystallogr. D 2010, 66, 486–501. [Google Scholar] [CrossRef]
- Murshudov, G. N.; Skubak, P.; Lebedev, A. A.; Pannu, N. S.; Steiner, R. A.; Nicholls, R. A.; Winn, M. D.; Long, F.; Vagin, A. A. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D 2011, 67, 355–367. [Google Scholar] [CrossRef]
- Agirre, J.; Atanasova, M.; Bagdonas, H.; Ballard, C. B.; Basle, A.; Beilsten-Edmands, J.; Borges, R. J.; Brown, D. G.; Burgos-Marmol, J. J.; Berrisford, J. M.; Bond, P. S.; Caballero, I.; Catapano, L.; Chojnowski, G.; Cook, A. G.; Cowtan, K. D.; Croll, T. I.; Debreczeni, J. E.; Devenish, N. E.; Dodson, E. J.; Drevon, T. R.; Emsley, P.; Evans, G.; Evans, P. R.; Fando, M.; Foadi, J.; Fuentes-Montero, L.; Garman, E. F.; Gerstel, M.; Gildea, R. J.; Hatti, K.; Hekkelman, M. L.; Heuser, P.; Hoh, S. W.; Hough, M. A.; Jenkins, H. T.; Jimenez, E.; Joosten, R. P.; Keegan, R. M.; Keep, N.; Krissinel, E. B.; Kolenko, P.; Kovalevskiy, O.; Lamzin, V. S.; Lawson, D. M.; Lebedev, A. A.; Leslie, A. G. W.; Lohkamp, B.; Long, F.; Maly, M.; McCoy, A. J.; McNicholas, S. J.; Medina, A.; Millan, C.; Murray, J. W.; Murshudov, G. N.; Nicholls, R. A.; Noble, M. E. M.; Oeffner, R.; Pannu, N. S.; Parkhurst, J. M.; Pearce, N.; Pereira, J.; Perrakis, A.; Powell, H. R.; Read, R. J.; Rigden, D. J.; Rochira, W.; Sammito, M.; Sanchez Rodriguez, F.; Sheldrick, G. M.; Shelley, K. L.; Simkovic, F.; Simpkin, A. J.; Skubak, P.; Sobolev, E.; Steiner, R. A.; Stevenson, K.; Tews, I.; Thomas, J. M. H.; Thorn, A.; Valls, J. T.; Uski, V.; Uson, I.; Vagin, A.; Velankar, S.; Vollmar, M.; Walden, H.; Waterman, D.; Wilson, K. S.; Winn, M. D.; Winter, G.; Wojdyr, M.; Yamashita, K. The CCP4 suite: integrative software for macromolecular crystallography. Acta Crystallogr. D 2023, 79, 449–461. [Google Scholar] [CrossRef] [PubMed]
- Laskowski, R. A.; Macarthur, M. W.; Moss, D. S.; Thornton, J. M. Procheck - a Program to Check the Stereochemical Quality of Protein Structures. J. Appl. Crystallogr. 1993, 26, 283–291. [Google Scholar] [CrossRef]
- Williams, C. J.; Headd, J. J.; Moriarty, N. W.; Prisant, M. G.; Videau, L. L.; Deis, L. N.; Verma, V.; Keedy, D. A.; Hintze, B. J.; Chen, V. B.; Jain, S.; Lewis, S. M.; Arendall, W. B., 3rd; Snoeyink, J.; Adams, P. D.; Lovell, S. C.; Richardson, J. S.; Richardson, D. C. MolProbity: More and better reference data for improved all-atom structure validation. Protein Sci. 2018, 27, 293–315. [Google Scholar] [CrossRef]
- Pettersen, E. F.; Goddard, T. D.; Huang, C. C.; Couch, G. S.; Greenblatt, D. M.; Meng, E. C.; Ferrin, T. E. UCSF Chimera--a visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef] [PubMed]
- Pettersen, E. F.; Goddard, T. D.; Huang, C. C.; Meng, E. C.; Couch, G. S.; Croll, T. I.; Morris, J. H.; Ferrin, T. E. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 2021, 30, 70–82. [Google Scholar] [CrossRef] [PubMed]
- Combs, S. A.; Deluca, S. L.; Deluca, S. H.; Lemmon, G. H.; Nannemann, D. P.; Nguyen, E. D.; Willis, J. R.; Sheehan, J. H.; Meiler, J. Small-molecule ligand docking into comparative models with Rosetta. Nat. Protoc. 2013, 8, 1277–1298. [Google Scholar] [CrossRef] [PubMed]
- DeLuca, S.; Khar, K.; Meiler, J. Fully Flexible Docking of Medium Sized Ligand Libraries with RosettaLigand. PLoS One 2015, 10, e0132508. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



