Submitted:
12 January 2026
Posted:
13 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
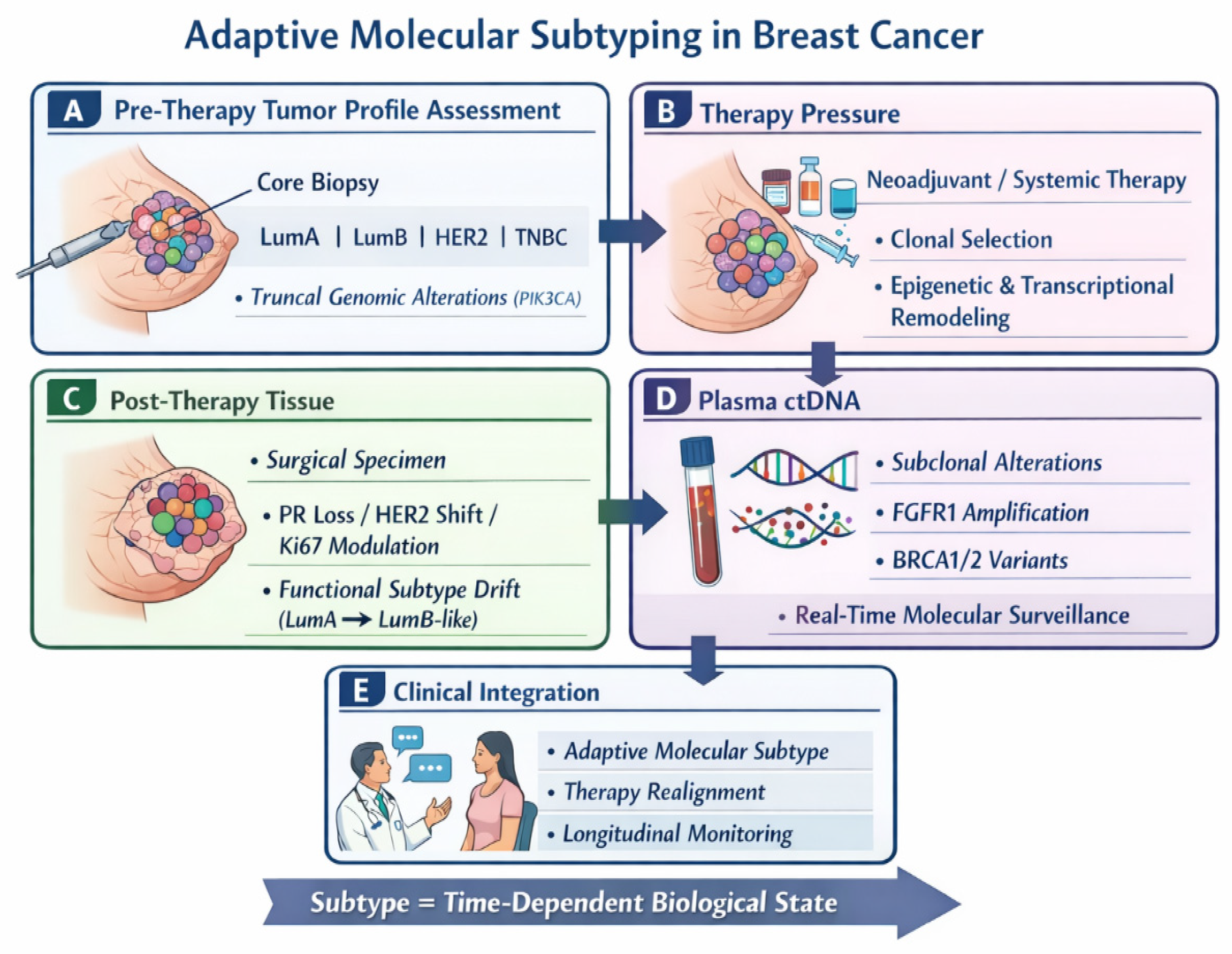
2. Materials and Methods
2.1. Study Design and Patient Cohort
- 1.
- Core biopsy specimens, representing diagnostic tissue obtained at initial presentation, were used for histopathological evaluation, immunohistochemistry (IHC), molecular subtype assignment, and tissue-based NGS when available.
- 2.
- Plasma samples were analyzed using circulating tumor DNA (ctDNA)/cell-free DNA (cfDNA) NGS assays.
- 3.
- Surgical specimens (mastectomy samples), when available, were subjected to histopathological examination and immunophenotypic and molecular analyses to assess post-therapy changes.
2.2. Longitudinal Biomarker Change. For Paired Baseline–Surgical Comparisons, the Following Definitions Were Applied
- Receptor conversion: change in ER and/or PR status between core biopsy and surgical specimen;
- HER2 score variability: any change in HER2 IHC score between time points;
- Ki67 variation: absolute change in Ki67 expressed as percentage points
2.3. Tissue NGS and Plasma ctDNA Analysis
2.4. Statistical Analysis
2.5. Study Limitations
3. Results
3.1. Pre-Treatment Molecular and Immunophenotypic Profile
3.1.1. Histopathology Assessment (Histological Subtype and Grade)
3.1.2. Hormone Receptor Expression Pre-Treatment
| Biomarker | Paired Cases (n) | Concordant Cases n (%) | Discordant Cases n (%) | Direction of Change | Agreement Statistic | p-Value |
| Estrogen receptor (ER) | 18 | 18 (100%) | 0 (0%) | No gain or loss observed | Cohen’s κ = 1.00 | Not applicable |
| Progesterone receptor (PR) | 18 | 12 (66.7%) | 6 (33.3%) | PR loss only (ER⁺PR⁺ → ER⁺PR⁻) | Cohen’s κ = 0.44 (moderate) | McNemar p = 0.031 |
| HER2 IHC score (0–3+) | 18 | 12 (66.7%) | 6 (33.3%) | Both upward and downward shifts | Weighted κ = 0.52 | Not tested* |
| Ki67 (%) | 18 | 4 (22.2%) | 14 (77.8%) | ↓ in 10 (55.6%); ↑ in 4 (22.2%) | — | Wilcoxon p = 0.018 |
3.1.3. Pre-Treatment Ki67
3.2. Pre-Treatment Genomic Alterations
3.2.1. PIK3CA Pathogenic Variants
3.2.2. AKT and Downstream Signaling Alterations
3.2.3. TP53, BRCA1/2, and MYC Alterations
3.3. Plasma ctDNA Analysis
3.3.1. Genomic Alterations Detected Exclusively in Plasma Samples
3.3.2. ctDNA Interpretive Constraints: Variant Classification and Assay Validity
3.4. Phenotypic/Genomic Drift Between Core Biopsy and Resection Specimens: Evidence of Molecular Drift/Plasticity and Implications for Resistance/Therapy Response
3.4.1. Hormone Receptor Conversion
3.4.2. HER2 Variability
3.4.3. Ki67 Variability
3.4.4. Molecular Subtypes. Luminal A to Functional Luminal B: Proliferation-Driven Drift
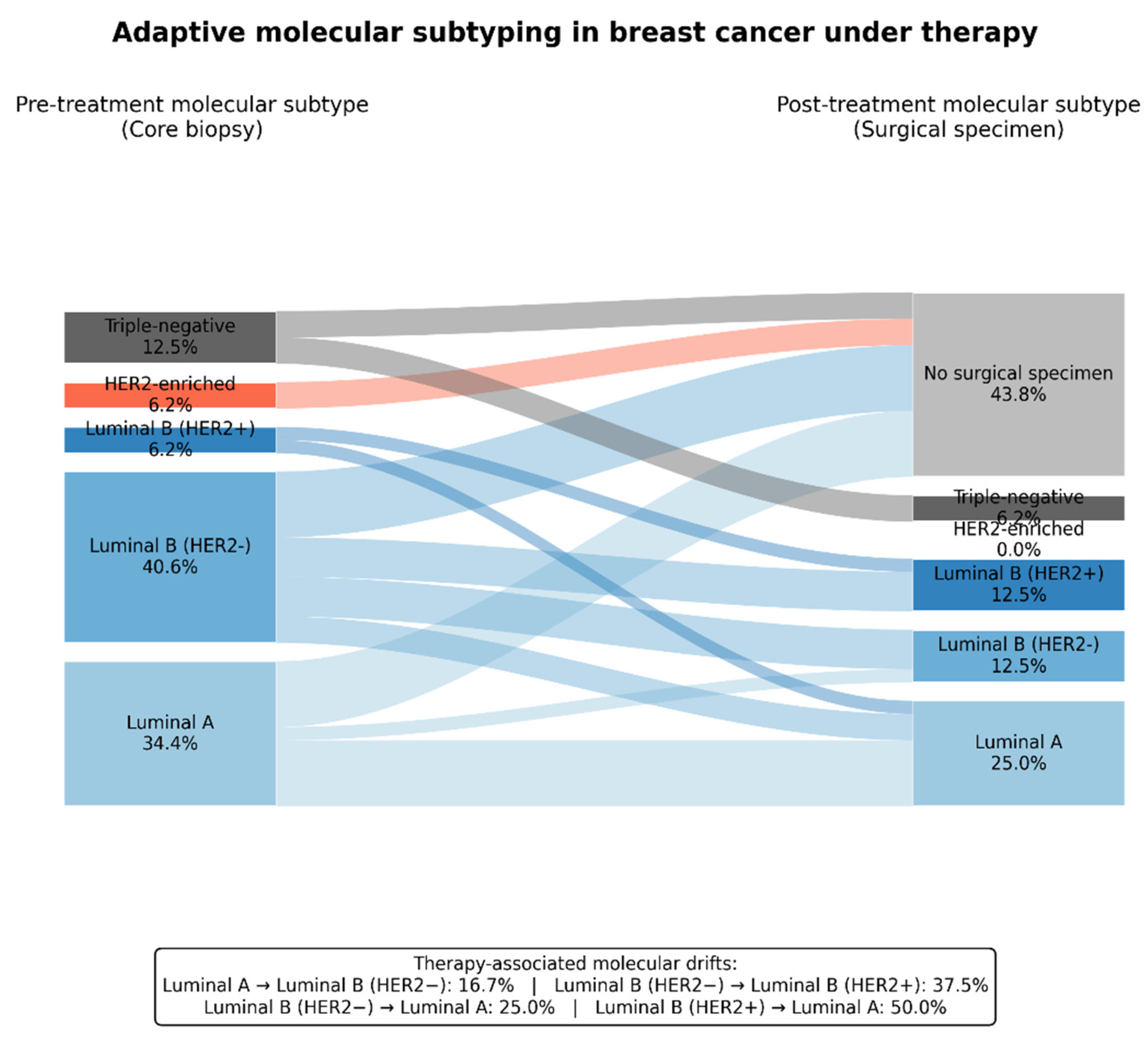
3.4.5. HER2-Positive / HER2-Enriched Tumors: A High-Proliferation, Instability-Associated Phenotype
3.5. PI3K/AKT Pathway Alterations and Endocrine Resistance Biology
3.5.1. FGFR Amplification
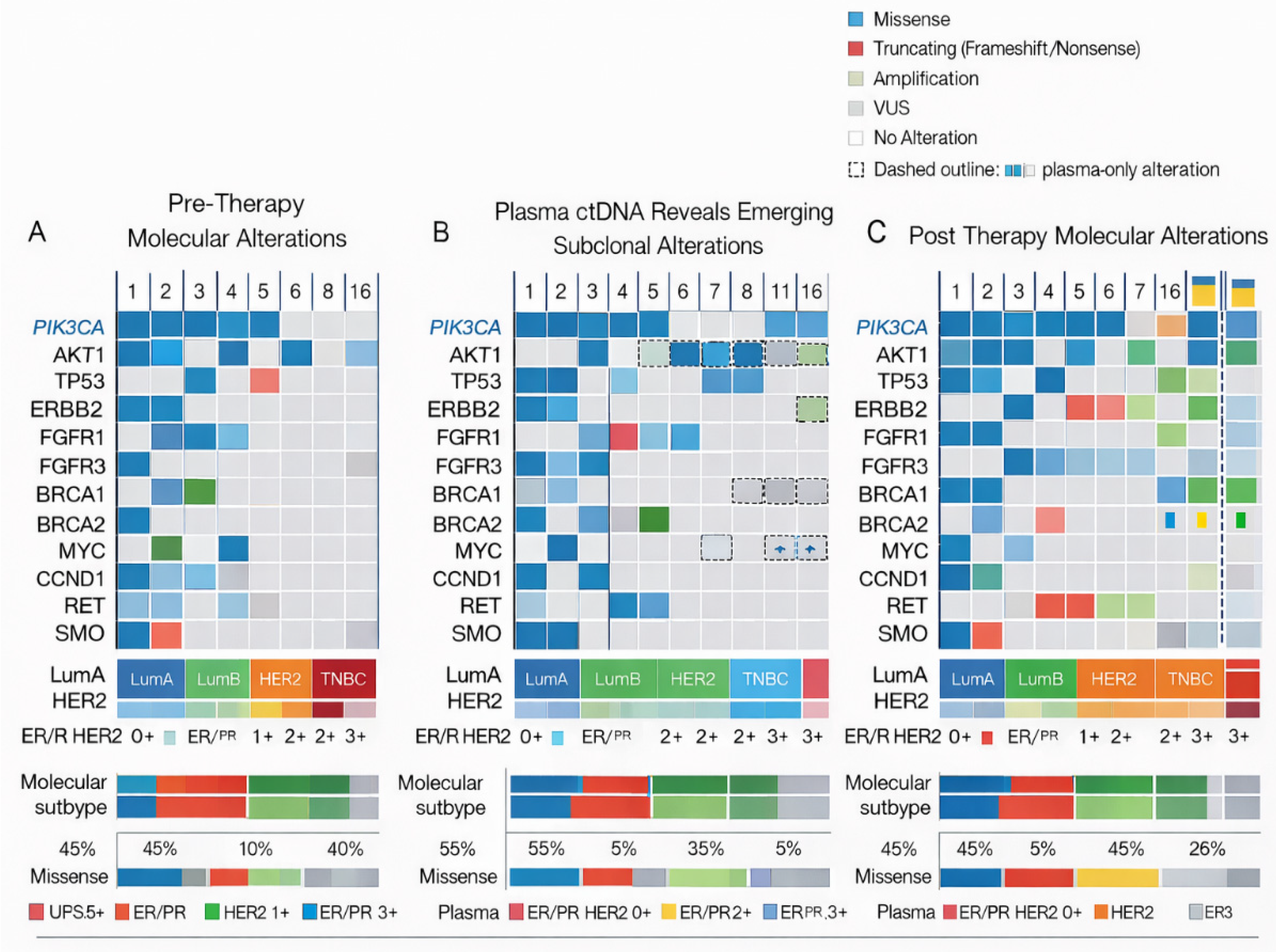
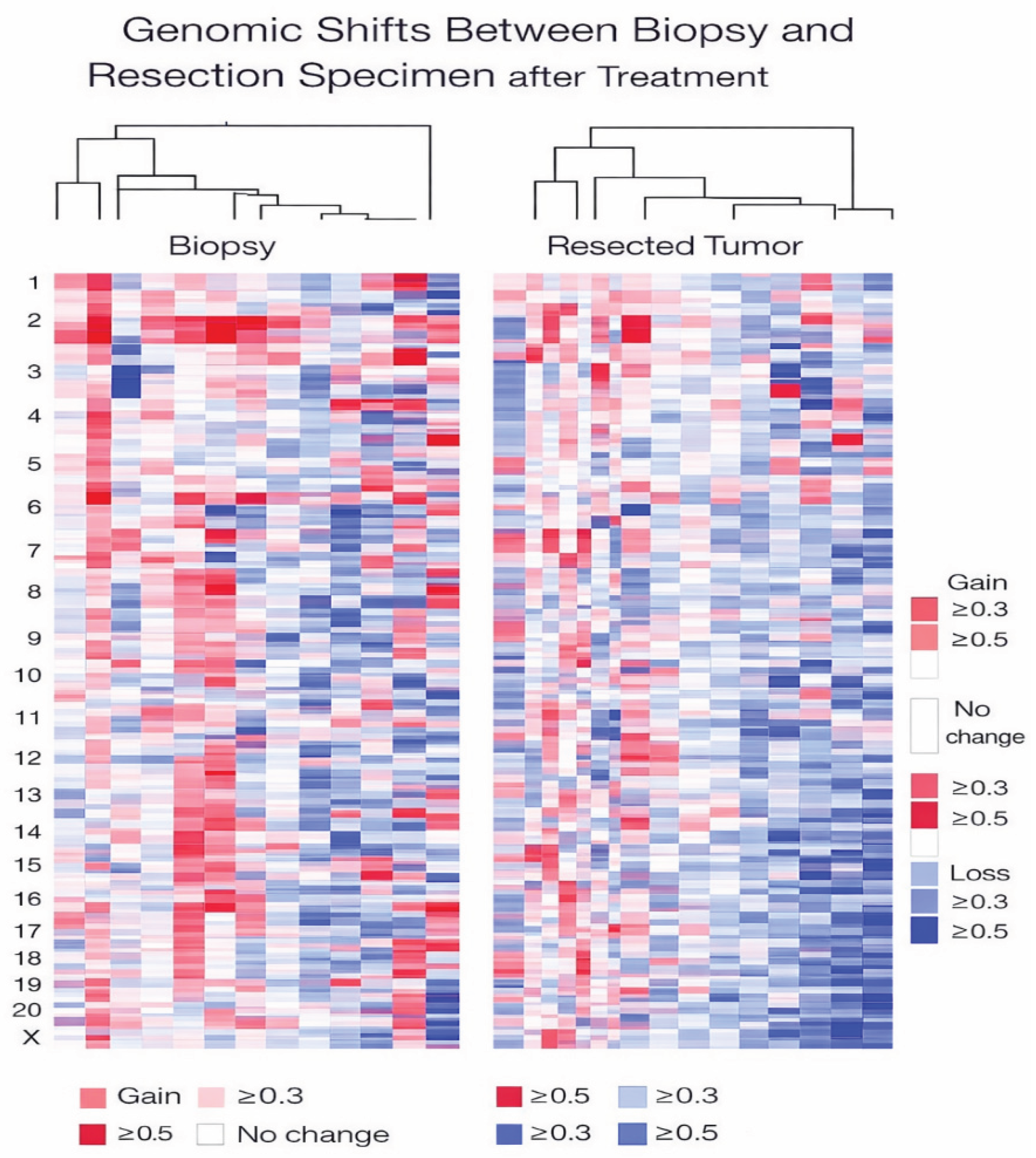
3.5.2. TP53 Alterations and High-Risk Biological States
4. Discussion
4.1. Molecular Plasticity Under Therapeutic Pressure
4.2. Adaptive Molecular Subtyping as a Clinically Relevant Framework
4.3. Genomic Drivers of Adaptation and Resistance
4.4. Complementary Value of Liquid Biopsy in Longitudinal Assessment
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- (Miron), A.-I.I.; Atasiei, D.-I.; Ionescu, R.-T.; Ultimescu, F.; Barnonschi, A.-A.; Anghel, A.-V.; Anghel, C.-A.; Antone-Iordache, I.-L.; Mitre, R.; Bobolocu, A.M.; et al. Prediction of Subclinical and Clinical Multiple Organ Failure Dysfunction in Breast Cancer Patients—A Review Using AI Tools. Cancers 2024, 16, 381. [CrossRef]
- Ultimescu, F.; Hudita, A.; Popa, D.E.; Olinca, M.; Muresean, H.A.; Ceausu, M.; Stanciu, D.I.; Ginghina, O.; Galateanu, B. Impact of Molecular Profiling on Therapy Management in Breast Cancer. J. Clin. Med. 2024, 13, 4995. [CrossRef]
- Hoogstraat, M.; Lips, E.H.; Mayayo-Peralta, I.; Mulder, L.; Kristel, P.; van der Heijden, I.; Annunziato, S.; van Seijen, M.; Nederlof, P.M.; Sonke, G.S.; et al. Comprehensive characterization of pre- and post-treatment samples of breast cancer reveal potential mechanisms of chemotherapy resistance. npj Breast Cancer 2022, 8, 1–12. [CrossRef]
- Miron, A.-I.; Anghel, A.-V.; Barnonschi, A.-A.; Mitre, R.; Liscu, H.-D.; Găinariu, E.; Pătru, R.; Coniac, S. Real-World Outcomes of CDK4/6 Inhibitors Treatment in Metastatic Breast Cancer in Romania. Diagnostics 2023, 13, 1938. [CrossRef]
- Khan, A.; Sisodiya, S.; Aftab, M.; Tanwar, P.; Hussain, S.; Gupta, V. Mechanisms and Therapeutic Strategies for Endocrine Resistance in Breast Cancer: A Comprehensive Review and Meta-Analysis. Cancers 2025, 17, 1653. [CrossRef]
- Dowling, G.P.; Daly, G.R.; Hehir, C.M.; AlRawasdeh, M.M.; Calpin, G.G.; Almasri, S.; Toomey, S.; Young, L.S.; Hennessey, B.T.; Hill, A.D. Prognostic significance of receptor conversion following neoadjuvant therapy in breast cancer: a systematic review & meta-analysis. Breast 2025, 82, 104516. [CrossRef]
- Ferenczi, Á.; Lantos, T.; Cserni, G. Temporality of change in Ki67 labelling following neoadjuvant systemic chemotherapy of breast cancer—the common drop often subsides with time. Virchows Arch. 2025, 1–10. [CrossRef]
- Cheung, A.M.; Wang, D.; Quintayo, M.A.; Yerofeyeva, Y.; Spears, M.; Bartlett, J.M.S.; Stein, L.; Bayani, J.; Yaffe, M.J. Intra-tumoral spatial heterogeneity in breast cancer quantified using high-dimensional protein multiplexing and single cell phenotyping. Breast Cancer Res. 2025, 27, 1–18. [CrossRef]
- Schettini, F.; Nucera, S.; Brasó-Maristany, F.; De Santo, I.; Pascual, T.; Bergamino, M.; Galván, P.; Conte, B.; Seguí, E.; Fructuoso, I.G.; et al. Unraveling the clinicopathological and molecular changes induced by neoadjuvant chemotherapy and endocrine therapy in hormone receptor-positive/HER2-low and HER2-0 breast cancer. ESMO Open 2024, 9, 103619. [CrossRef]
- Denkert, C.; Rachakonda, S.; Karn, T.; Weber, K.; Martin, M.; Marmé, F.; Untch, M.; Bonnefoi, H.; Kim, S.-B.; Seiler, S.; et al. Dynamics of molecular heterogeneity in high-risk luminal breast cancer—From intrinsic to adaptive subtyping. Cancer Cell 2025, 43, 232–247.e4. [CrossRef]
- Denkert, C.; Rachakonda, K.; Filipits, M.; Weber, K.; Marmé, F.; Untch, M.; Witkiewicz, A.K.; Im, S.-A.; DeMichele, A.; Pehl, A.; et al. Relationship of adaptive subtyping and tumour heterogeneity of treatment response to neoadjuvant therapy in hormone receptor–positive HER2-negative early breast cancer: PENELOPE-B.. J. Clin. Oncol. 2024, 42, 566–566. [CrossRef]
- Aanestad, K.L.; Austdal, M.; Nordgård, O.; Mellgren, G.; Oltedal, S.; Austbø, M.L.; Vignes, Y.H.; Helland, T.; Jonsdottir, K.; Lende, T.H.; et al. Monitoring of circulating tumor DNA allows early detection of disease relapse in patients with operable breast cancer. Mol. Oncol. 2025. [CrossRef]
- Fuentes-Antrás, J.; Elliott, M.J.; Main, S.C.; Echelard, P.; Dou, A.; Bedard, P.L.; Amir, E.; Nadler, M.B.; Meti, N.; Gregorio, N.; et al. Personalized ctDNA monitoring in metastatic HR+/HER2− breast cancer patients during endocrine and CDK4/6 inhibitor therapy. npj Breast Cancer 2025, 11, 74. [CrossRef]
- Jørgensen, C.L.T.; Larsson, A.-M.; Forsare, C.; Aaltonen, K.; Jansson, S.; Bradshaw, R.; Bendahl, P.-O.; Rydén, L. PAM50 Intrinsic Subtype Profiles in Primary and Metastatic Breast Cancer Show a Significant Shift toward More Aggressive Subtypes with Prognostic Implications. Cancers 2021, 13, 1592. [CrossRef]
- Garcia-Recio, S.; Hinoue, T.; Wheeler, G.L.; Kelly, B.J.; Garrido-Castro, A.C.; Pascual, T.; De Cubas, A.A.; Xia, Y.; Felsheim, B.M.; McClure, M.B.; et al. Multiomics in primary and metastatic breast tumors from the AURORA US network finds microenvironment and epigenetic drivers of metastasis. Nat. Cancer 2022, 4, 128–147. [CrossRef]
- Nolan, E.; Lindeman, G.J.; Visvader, J.E. Deciphering breast cancer: from biology to the clinic. Cell 2023, 186, 1708–1728. [CrossRef]
- Cardoso, F.; Paluch-Shimon, S.; Schumacher-Wulf, E.; Matos, L.; Gelmon, K.; Aapro, M.S.; Bajpai, J.; Barrios, C.H.; Bergh, J.; Bergsten-Nordström, E.; et al. 6th and 7th International consensus guidelines for the management of advanced breast cancer (ABC guidelines 6 and 7). Breast 2024, 76. [CrossRef]
- Comanescu, M.; Arsene, D.; Ardeleanu, C.; Bussolati, G. The mandate for a proper preservation in histopathological tissues. 2012, 53, 233–242.
- Wolff, A.C.; Somerfield, M.R.; Dowsett, M.; Hammond, M.E.H.; Hayes, D.F.; McShane, L.M.; Saphner, T.J.; Spears, P.A.; Allison, K.H. Human Epidermal Growth Factor Receptor 2 Testing in Breast Cancer. Arch. Pathol. Lab. Med. 2023, 147, 993–1000. [CrossRef]
- Goldhirsch, A.; Ingle, J.N.; Gelber, R.D.; Coates, A.S.; Thürlimann, B.; Senn, H.-J. Thresholds for therapies: highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2009. Ann. Oncol. 2009, 20, 1319–1329. [CrossRef]
- Lu, Z.; Wang, T.; Wang, L.; Ming, J. Research progress on estrogen receptor-positive/progesterone receptor-negative breast cancer. Transl. Oncol. 2025, 56, 102387. [CrossRef]
- Toli, M.A.; Liu, X.; Massa, D.; Lando, S.; Boman, C.; Tsiknakis, N.; Tranchell, C.; Papakonstantinou, A.; Fotia, G.; Vernieri, C.; et al. Prognostic significance of tumour Ki-67 dynamics during neoadjuvant treatment in patients with breast cancer: a population-based cohort study. Lancet Reg. Heal. - Eur. 2025, 58, 101432. [CrossRef]
- Bernhardt, M.; Weinhold, L.; Sanders, C.; Hommerding, O.; Lau, J.; Toma, M.; Tischler, V.; Schmid, M.; Zienkiewicz, T.; Hildenbrand, R.; et al. Peer-to-peer validation of Ki-67 scoring in a pathology quality circle as a tool to assess interobserver variability: are we better than we thought?. APMIS 2024, 132, 718–727. [CrossRef]
- Boyaci, C.; Sun, W.; Hartman, J.; Ács, B. Global scoring method of Ki67 immunohistochemistry in breast cancer demonstrates improved concordance using real-world multi-institutional data. Breast Cancer Res. 2025, 27, 1–9. [CrossRef]
- Hao, C.; Wei, Y.; Meng, W.; Zhang, J.; Yang, X. PI3K/AKT/mTOR inhibitors for hormone receptor-positive advanced breast cancer. Cancer Treat. Rev. 2024, 132, 102861. [CrossRef]
- Hao, C.; Wei, Y.; Meng, W.; Zhang, J.; Yang, X. PI3K/AKT/mTOR inhibitors for hormone receptor-positive advanced breast cancer. Cancer Treat. Rev. 2024, 132, 102861. [CrossRef]
- Turner, N.C.; Oliveira, M.; Howell, S.J.; Dalenc, F.; Cortes, J.; Moreno, H.L.G.; Hu, X.; Jhaveri, K.; Krivorotko, P.; Loibl, S.; et al. Capivasertib in Hormone Receptor–Positive Advanced Breast Cancer. New Engl. J. Med. 2023, 388, 2058–2070. [CrossRef]
- Schagerholm, C.; Robertson, S.; Toosi, H.; Sifakis, E.G.; Hartman, J. PIK3CA mutations in endocrine-resistant breast cancer. Sci. Rep. 2024, 14, 1–13. [CrossRef]
- Turner, N.C.; Oliveira, M.; Howell, S.J.; Dalenc, F.; Cortes, J.; Moreno, H.L.G.; Hu, X.; Jhaveri, K.; Krivorotko, P.; Loibl, S.; et al. Capivasertib in Hormone Receptor–Positive Advanced Breast Cancer. New Engl. J. Med. 2023, 388, 2058–2070. [CrossRef]
- Li, J.; Chen, L.; Chen, X.; Huang, M.; Guo, W.; Chen, M.; Lin, Y.; Wang, Y.; Cai, W.; Qiu, Y.; et al. Clinical TP53 genetic testing is recommended for HER2-positive breast cancer patients aged 35 or younger. npj Genom. Med. 2025, 10, 1–7. [CrossRef]
- Zhou, T.; Zhang, J. Therapeutic advances and application of PARP inhibitors in breast cancer. Transl. Oncol. 2025, 57, 102410. [CrossRef]
- Amato, O.; Giannopoulou, N.; Ignatiadis, M. Circulating tumor DNA validity and potential uses in metastatic breast cancer. npj Breast Cancer 2024, 10, 1–7. [CrossRef]
- de Jager, V.D.; Giacomini, P.; Fairley, J.A.; Toledo, R.A.; Patton, S.J.; Joosse, S.A.; Koch, C.; Deans, Z.C.; Agelaki, S.; Andersen, C.L.; et al. Reporting of molecular test results from cell-free DNA analyses: expert consensus recommendations from the 2023 European Liquid Biopsy Society ctDNA Workshop. EBioMedicine 2025, 114, 105636. [CrossRef]
- Bergamino, M.A.; López-Knowles, E.; Morani, G.; Tovey, H.; Kilburn, L.; Schuster, E.F.; Alataki, A.; Hills, M.; Xiao, H.; Holcombe, C.; et al. HER2-enriched subtype and novel molecular subgroups drive aromatase inhibitor resistance and an increased risk of relapse in early ER+/HER2+ breast cancer. EBioMedicine 2022, 83, 104205. [CrossRef]
- Nierengarten, M.B. FDA approves capivasertib with fulvestrant for breast cancer. Cancer 2024, 130, 835–836. [CrossRef]
- Formisano, L.; Lu, Y.; Servetto, A.; Hanker, A.B.; Jansen, V.M.; Bauer, J.A.; Sudhan, D.R.; Guerrero-Zotano, A.L.; Croessmann, S.; Guo, Y.; et al. Aberrant FGFR signaling mediates resistance to CDK4/6 inhibitors in ER+ breast cancer. Nat. Commun. 2019, 10, 1373. [CrossRef]
- Li, J.; Chen, L.; Chen, X.; Huang, M.; Guo, W.; Chen, M.; Lin, Y.; Wang, Y.; Cai, W.; Qiu, Y.; et al. Clinical TP53 genetic testing is recommended for HER2-positive breast cancer patients aged 35 or younger. npj Genom. Med. 2025, 10, 1–7. [CrossRef]
- Nik-Zainal, S.; Van Loo, P.; Wedge, D.C.; Alexandrov, L.B.; Greenman, C.D.; Lau, K.W.; Raine, K.; Jones, D.; Marshall, J.; Ramakrishna, M.; et al. The Life History of 21 Breast Cancers. Cell 2012, 149, 994–1007. [CrossRef]
- Esparza-López, J.; Escobar-Arriaga, E.; Soto-Germes, S.; Ibarra-Sánchez, M.d.J. Breast Cancer Intra-Tumor Heterogeneity: One Tumor, Different Entities. 2017, 69, 66–76. [CrossRef]
- Xu, M.; Yan, Y.; Chen, Y.; Chen, X.; Gong, K.; Fu, F. Expression and Subtype Discordance Between Core Needle Biopsy and Surgical Specimen in Breast Cancer. J. Surg. Res. 2025, 307, 42–52. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



