Submitted:
26 January 2026
Posted:
05 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
Cell Culture
N-HCC25 Cell Line
Primary Murine Macrophage Isolation
CRISPR/Cas9-Mediated KO of Nrf2 and Keap1
Fluorescent Labeling of Cells
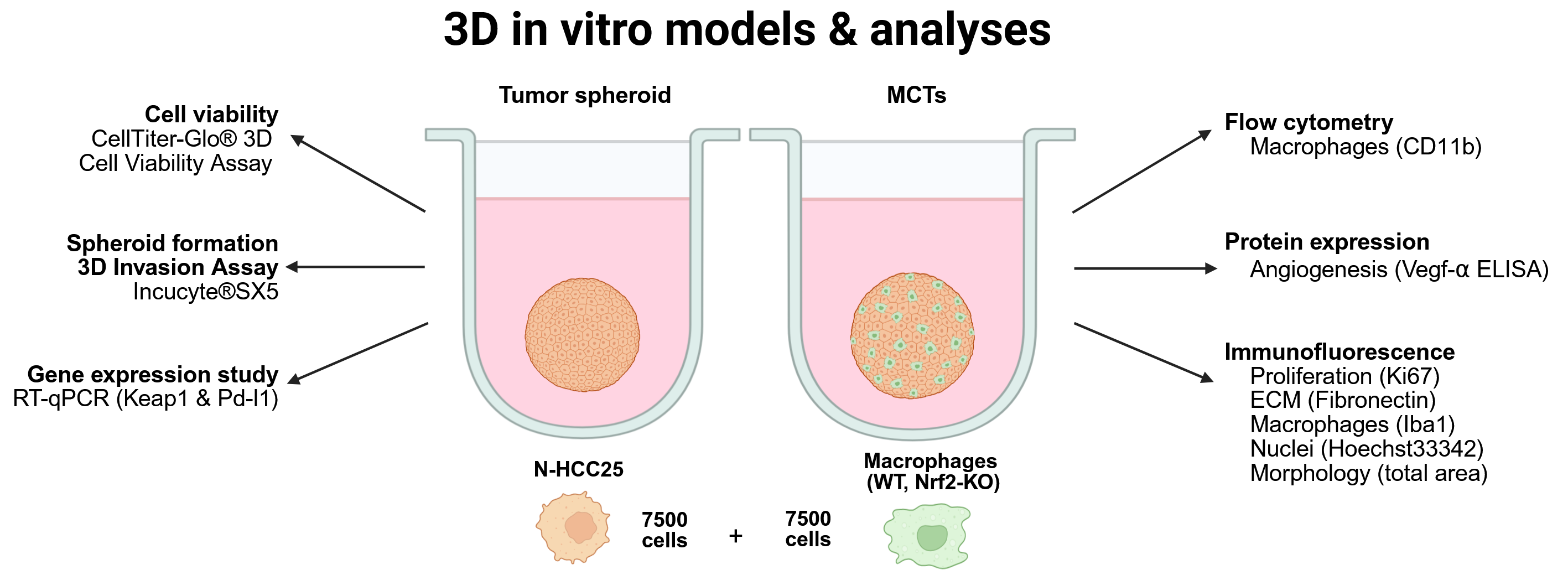
Experimental In Vitro Models (see Fig. A1)
Conditioned Media = Tumor Spheroid and Macrophages CM Model (Fig. A1A)
Co-Culture Models
Transwell = Spheroid in Well and BMDMs in Transwell (Fig. A1B)
Transwell Tower = Spheroids Inside the Transwell, BMDMs on the Bottom (Fig. A1C)
Spheroid and Multicellular Tumor Spheroid (MCT) model (Fig. A1D)
Readouts
Spheroid Morphology
Viability Assay
Invasions Assay
ELISA
Cryo-Sectioning and IF Staining of Spheroids
Flow Cytometry
RNA Isolation, cDNA Synthesis and RT-qPCR
Statistics
3. Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bray, F; Laversanne, M; Sung, H; Ferlay, J; Siegel RLS, I.; Jemal, A. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: a cancer journal for clinicians 2024, 74(3), 229–263. [Google Scholar] [CrossRef] [PubMed]
- Ferlay, J; Ervik, M; Lam, F; Laversanne, M; Colombet, M; Mery, L; Piñeros, M; Znaor, A; Soerjomataram, I; Bray, F. Global Cancer Observatory: Cancer Today (version 1.1). Lyon, France: International Agency for Research on Cancer, 2024. [Google Scholar]
- Filho, AM; Laversanne, M; Ferlay, J; Colombet, M; Piñeros, M; Znaor, A; Parkin, DM; Soerjomataram, I; Bray, F. The GLOBOCAN 2022 cancer estimates: Data sources, methods, and a snapshot of the cancer burden worldwide. 2025, 156(7), 1336–1346. [Google Scholar] [CrossRef] [PubMed]
- Estes, C; Razavi, H; Loomba, R; Younossi, Z; Sanyal, AJ. Modeling the epidemic of nonalcoholic fatty liver disease demonstrates an exponential increase in burden of disease. Hepatology 2018, 67(1), 123–133. [Google Scholar] [CrossRef] [PubMed]
- Connell, LC; Harding, JJ; Abou-Alfa, GK. Advanced Hepatocellular Cancer: the Current State of Future Research. Current treatment options in oncology 2016, 17(8), 43. [Google Scholar] [CrossRef]
- Lv, B; Wang, Y; Ma, D; Cheng, W; Liu, J; Yong, T; Chen, H; Wang, C. Immunotherapy: Reshape the Tumor Immune Microenvironment. Front Immunol 2022, 13, 844142. [Google Scholar] [CrossRef]
- Binnewies, M; Roberts, EW; Kersten, K; Chan, V; Fearon, DF; Merad, M; Coussens, LM; Gabrilovich, DI; Ostrand-Rosenberg, S; Hedrick, C.C; et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nature medicine 2018, 24(5), 541–550. [Google Scholar] [CrossRef]
- Martinez, FO; Sica, A; Mantovani, A; Locati, M. Macrophage activation and polarization. Frontiers in bioscience: a journal and virtual library 2008, 13, 453–461. [Google Scholar] [CrossRef]
- Boutilier, AJ; Elsawa, SF. Macrophage Polarization States in the Tumor Microenvironment. International journal of molecular sciences 2021, 22(13). [Google Scholar] [CrossRef]
- Yamamoto, M; Kensler, TW; Motohashi, H. The KEAP1-NRF2 System: a Thiol-Based Sensor-Effector Apparatus for Maintaining Redox Homeostasis. Physiological reviews 2018, 98(3), 1169–1203. [Google Scholar] [CrossRef]
- Rojo de la Vega, M; Chapman, E; Zhang, DD. NRF2 and the Hallmarks of Cancer . Cancer cell 2018, 34(1), 21–43. [Google Scholar] [CrossRef]
- Schaer, DJ; Schulthess-Lutz, N; Baselgia, L; Hansen, K; Buzzi, RM; Humar, R; Dürst, E; Vallelian, F. Hemorrhage-activated NRF2 in tumor-associated macrophages drives cancer growth, invasion, and immunotherapy resistance. The Journal of clinical investigation 2023, 134(3). [Google Scholar] [CrossRef] [PubMed]
- Dufau, I; Frongia, C; Sicard, F; Dedieu, L; Cordelier, P; Ausseil, F; Ducommun, B; Valette, A. Multicellular tumor spheroid model to evaluate spatio-temporal dynamics effect of chemotherapeutics: application to the gemcitabine/CHK1 inhibitor combination in pancreatic cancer. BMC cancer 2012, 12, 15. [Google Scholar] [CrossRef] [PubMed]
- Pingitore, P; Sasidharan, K; Ekstrand, M; Prill, S; Lindén, D; Romeo, S. Human Multilineage 3D Spheroids as a Model of Liver Steatosis and Fibrosis. International journal of molecular sciences 2019, 20(7). [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S; Spagnoli, GC; Martin, I; Ploegert, S; Demougin, P; Heberer, M; Reschner, A. Three-dimensional culture of melanoma cells profoundly affects gene expression profile: a high density oligonucleotide array study. Journal of cellular physiology 2005, 204(2), 522–531. [Google Scholar] [CrossRef]
- Kim, H; Phung, Y; Ho, M. Changes in global gene expression associated with 3D structure of tumors: an ex vivo matrix-free mesothelioma spheroid model. PloS one 2012, 7(6), e39556. [Google Scholar] [CrossRef]
- Díaz, L; Zambrano, E; Flores, ME; Contreras, M; Crispín, JC; Alemán, G; Bravo, C; Armenta, A; Valdés, VJ; Tovar, A.; et al. Ethical Considerations in Animal Research: The Principle of 3R’s. Revista de investigacion clinica; organo del Hospital de Enfermedades de la Nutricion 2020, 73(4), 199–209. [Google Scholar] [CrossRef]
- Crouchet, E; Almeida, N; Durand, SC; Parnot, M; Oudot, MA; Giannone, F; Gadenne, C; Roehlen, N; Saviano, A; Felli, E.; et al. A patient-derived HCC spheroid system to model the tumor microenvironment and treatment response. JHEP reports: innovation in hepatology 2025, 7(2), 101252. [Google Scholar] [CrossRef]
- Han, S; Lim, JY; Cho, K; Lee, HW; Park, JY; Ro, SW; Kim, KS; Seo, HR; Kim, DY. : Anti-Cancer Effects of YAP Inhibitor (CA3) in Combination with Sorafenib against Hepatocellular Carcinoma (HCC) in Patient-Derived Multicellular Tumor Spheroid Models (MCTS). Cancers 2022, 14(11). [Google Scholar] [CrossRef]
- Ma, XL; Hu, B; Tang, WG; Xie, SH; Ren, N; Guo, L; Lu, RQ. : CD73 sustained cancer-stem-cell traits by promoting SOX9 expression and stability in hepatocellular carcinoma. Journal of hematology & oncology 2020, 13(1), 11. [Google Scholar] [CrossRef]
- Shimokawa, M; Yoshizumi, T; Itoh, S; Iseda, N; Sakata, K; Yugawa, K; Toshima, T; Harada, N; Ikegami, T; Mori, M. Modulation of Nqo1 activity intercepts anoikis resistance and reduces metastatic potential of hepatocellular carcinoma. Cancer science 2020, 111(4), 1228–1240. [Google Scholar] [CrossRef]
- Kroh, A; Walter, J; Schüler, H; Nolting, J; Eickhoff, R; Heise, D; Neumann, UP; Cramer, T; Ulmer, TF. Fragoulis A: A Newly Established Murine Cell Line as a Model for Hepatocellular Cancer in Non-Alcoholic Steatohepatitis. International journal of molecular sciences 2019, 20(22). [Google Scholar] [CrossRef] [PubMed]
- Zhang, X; Goncalves, R; Mosser, DM. The isolation and characterization of murine macrophages. Current protocols in immunology 2008, Chapter 14, 14.11.11–14.11.14. [Google Scholar] [CrossRef] [PubMed]
- Sanjana, NE; Shalem, O; Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat Methods 2014, 11(8), 783–784. [Google Scholar] [CrossRef] [PubMed]
- Wenzel, DM; Mackay, DR; Skalicky, JJ; Paine, EL; Miller, MS; Ullman, KS; Sundquist, WI. Comprehensive analysis of the human ESCRT-III-MIT domain interactome reveals new cofactors for cytokinetic abscission. eLife 2022, 11. [Google Scholar] [CrossRef]
- Wiznerowicz, M; Trono, D. Conditional suppression of cellular genes: lentivirus vector-mediated drug-inducible RNA interference. Journal of virology 2003, 77(16), 8957–8961. [Google Scholar] [CrossRef]
- Kang, YB; Rawat, S; Cirillo, J; Bouchard, M; Noh, HM. Layered long-term co-culture of hepatocytes and endothelial cells on a transwell membrane: toward engineering the liver sinusoid. Biofabrication 2013, 5(4), 045008. [Google Scholar] [CrossRef]
- Arganda-Carreras, I; Kaynig, V; Rueden, C; Eliceiri, KW; Schindelin, J; Cardona, A; Sebastian Seung, H. Trainable Weka Segmentation: a machine learning tool for microscopy pixel classification. Bioinformatics (Oxford, England) 2017, 33(15), 2424–2426. [Google Scholar] [CrossRef]
- Bustin, SA; Benes, V; Garson, JA; Hellemans, J; Huggett, J; Kubista, M; Mueller, R; Nolan, T; Pfaffl, MW; Shipley, G.L; et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 2009, 55(4), 611–622. [Google Scholar] [CrossRef]
- Ramakers, C; Ruijter, JM; Deprez, RH; Moorman, AF. Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neuroscience letters 2003, 339(1), 62–66. [Google Scholar] [CrossRef]
- Krumm, P; Böttcher, N; Ottermanns, R; Pufe, T; Fragoulis, A. BioMedStatX – Statistical workflows for reliable biomedical data analysis. MethodsX 2026, 16, 103776. [Google Scholar] [CrossRef]
- Arvanitakis, K; Koletsa, T; Mitroulis, I; Germanidis, G. Tumor-Associated Macrophages in Hepatocellular Carcinoma Pathogenesis, Prognosis and Therapy. Cancers 2022, 14(1). [Google Scholar] [CrossRef]
- Cheng, K; Cai, N; Zhu, J; Yang, X; Liang, H; Zhang, W. Tumor-associated macrophages in liver cancer: From mechanisms to therapy. Cancer communications (London, England) 2022, 42(11), 1112–1140. [Google Scholar] [CrossRef] [PubMed]
- Xu, R; Huang, H; Zhang, Z; Wang, FS. The role of neutrophils in the development of liver diseases. Cellular & molecular immunology 2014, 11(3), 224–231. [Google Scholar] [CrossRef]
- Vallelian, F; Buzzi, RM; Pfefferlé, M; Yalamanoglu, A; Dubach, IL; Wassmer, A; Gentinetta, T; Hansen, K; Humar, R; Schulthess, N.; et al. Heme-stress activated NRF2 skews fate trajectories of bone marrow cells from dendritic cells towards red pulp-like macrophages in hemolytic anemia. Cell death and differentiation 2022, 29(8), 1450–1465. [Google Scholar] [CrossRef]
- Tsui, YM; Chan, LK; Ng, I.O. Cancer stemness in hepatocellular carcinoma: mechanisms and translational potential. British journal of cancer 2020, 122(10), 1428–1440. [Google Scholar] [CrossRef]
- Nath, S; Devi, GR. Three-dimensional culture systems in cancer research: Focus on tumor spheroid model. Pharmacology & therapeutics 2016, 163, 94–108. [Google Scholar] [CrossRef]
- Xiong, T; Wang, K. Reconstructing the hepatocellular carcinoma microenvironment: the current status and challenges of 3D culture technology. Discover oncology 2025, 16(1), 506. [Google Scholar] [CrossRef]
- Khawar, IA; Park, JK; Jung, ES; Lee, MA; Chang, S; Kuh, HJ. : Three Dimensional Mixed-Cell Spheroids Mimic Stroma-Mediated Chemoresistance and Invasive Migration in hepatocellular carcinoma. Neoplasia 2018, 20(8), 800–812. [Google Scholar] [CrossRef]
- Kweider, N; Fragoulis, A; Rosen, C; Pecks, U; Rath, W; Pufe, T; Wruck, CJ. Interplay between vascular endothelial growth factor (VEGF) and nuclear factor erythroid 2-related factor-2 (Nrf2): implications for preeclampsia. The Journal of biological chemistry 2011, 286(50), 42863–42872. [Google Scholar] [CrossRef]
- Tao, Z; Li, P; Tang, Y; Yang, W; Li, Y; Yang, J; Tian, J; Zhang, Y; Zou, Y; Xu, B.; et al. Dexmedetomidine Promotes Angiogenesis After Ischemic Stroke Through the NRF2/HO-1/VEGF Pathway. Neurochemical research 2025, 50(2), 138. [Google Scholar] [CrossRef]
- Kobayashi, EH; Suzuki, T; Funayama, R; Nagashima, T; Hayashi, M; Sekine, H; Tanaka, N; Moriguchi, T; Motohashi, H; Nakayama, K.; et al. Nrf2 suppresses macrophage inflammatory response by blocking proinflammatory cytokine transcription. Nature communications 2016, 7, 11624. [Google Scholar] [CrossRef]
- Wang, L; He, C. Nrf2-mediated anti-inflammatory polarization of macrophages as therapeutic targets for osteoarthritis. Front Immunol 2022, 13, 967193. [Google Scholar] [CrossRef]
- Giraud, J; Chalopin, D; Blanc, JF; Saleh, M. Hepatocellular Carcinoma Immune Landscape and the Potential of Immunotherapies. Front Immunol 2021, 12, 655697. [Google Scholar] [CrossRef]
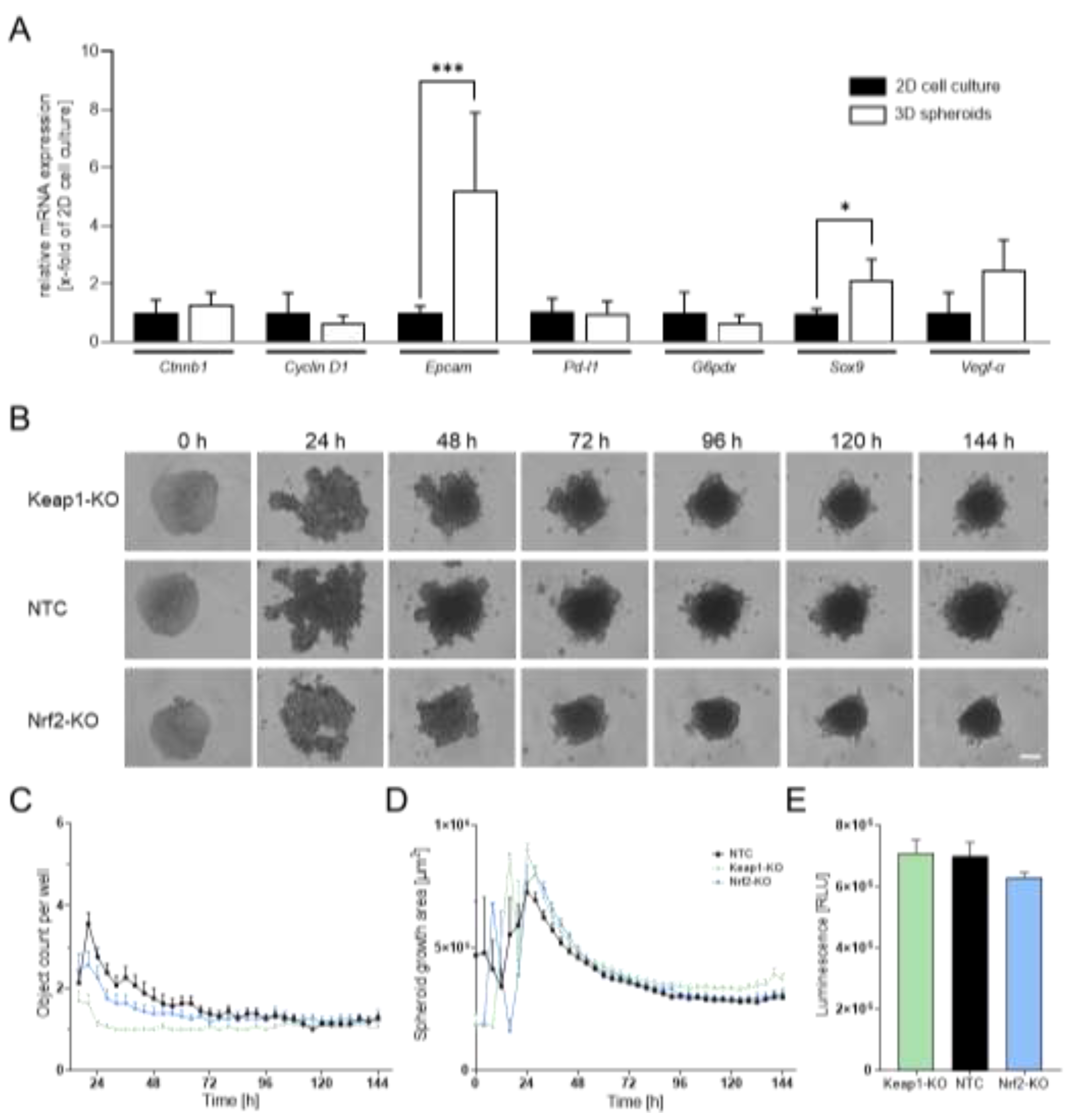
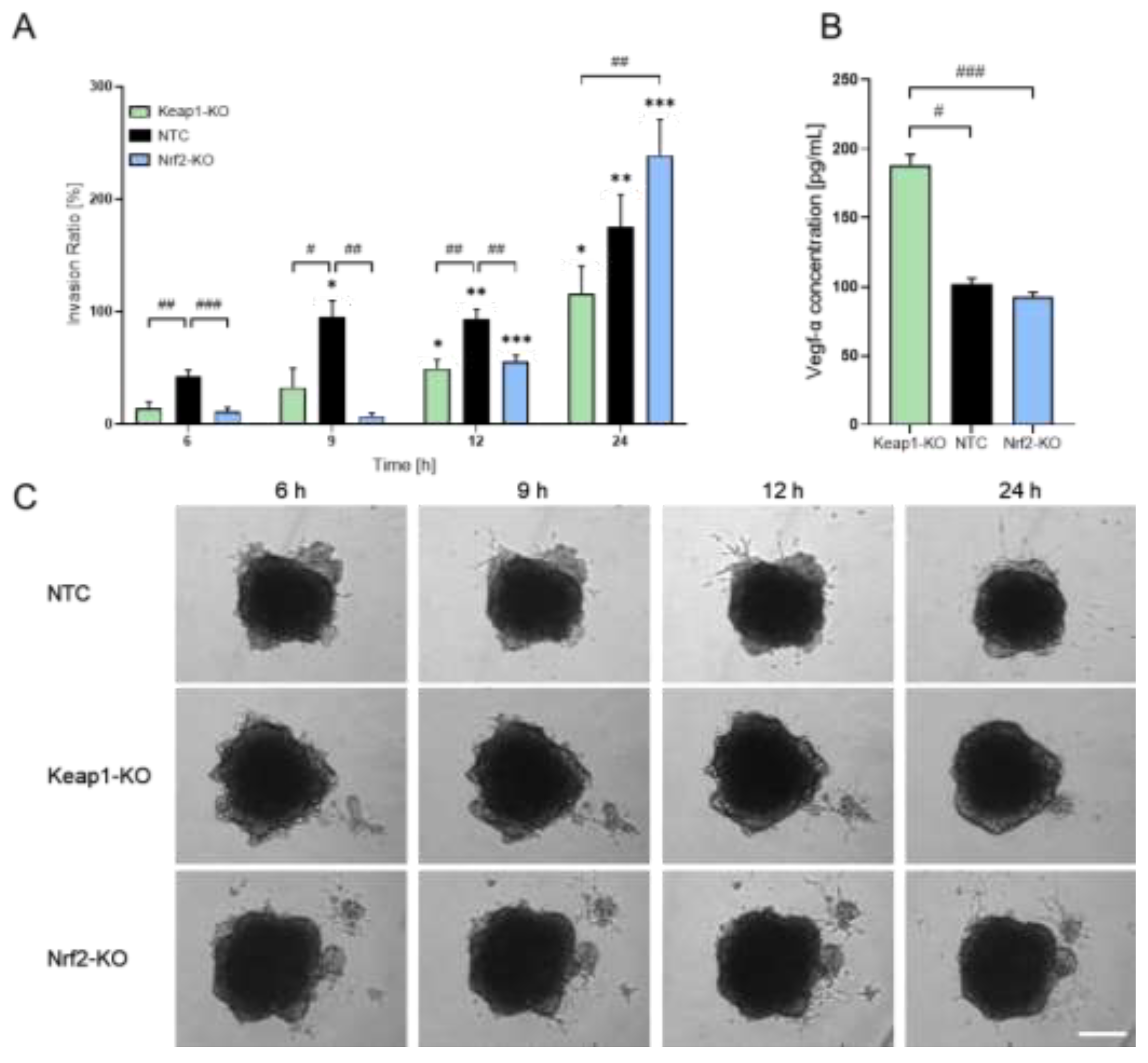
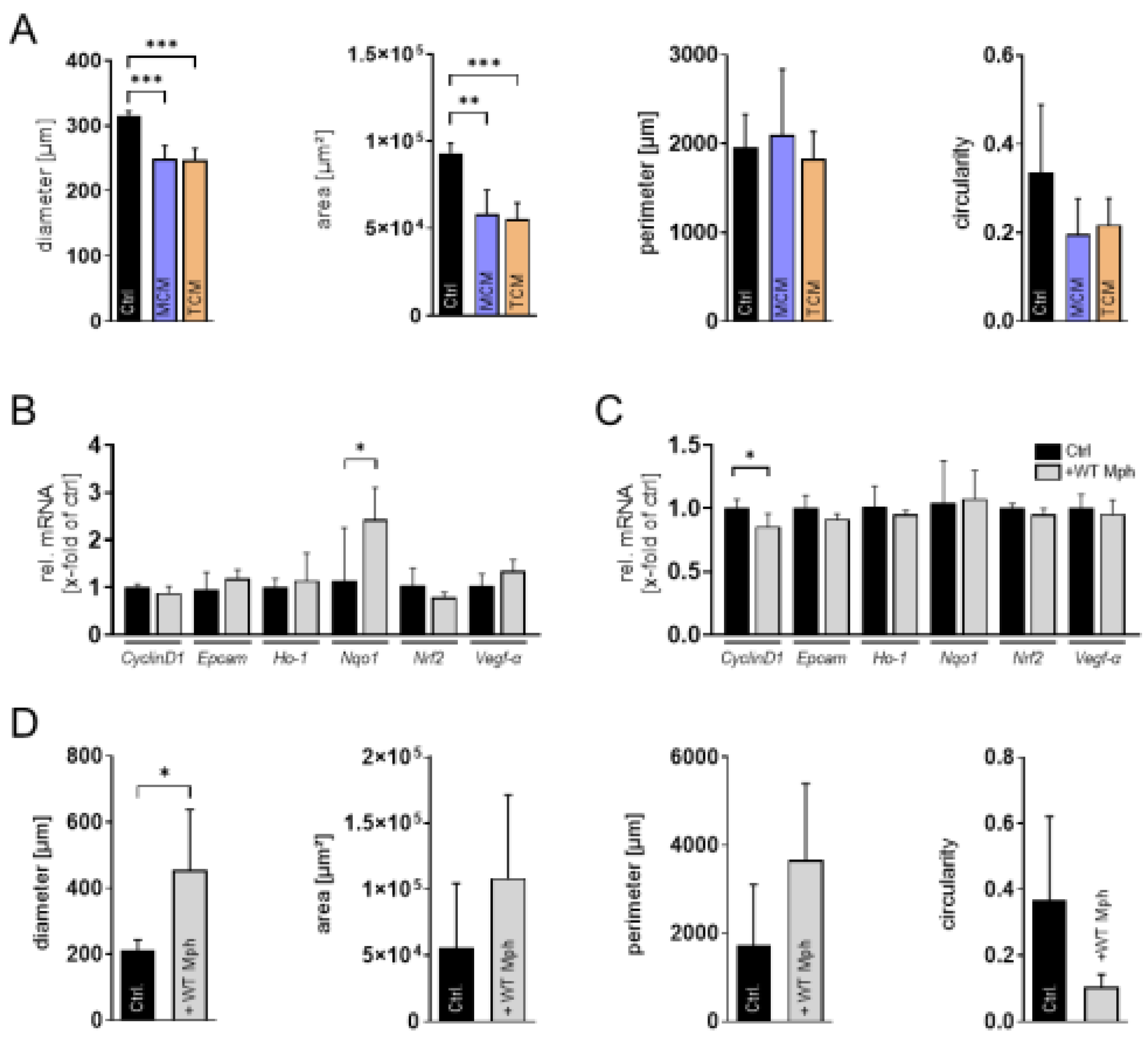
| Ref. genes | Sequence Accession Number | Direction | Sequence [5’-3’] |
TA [°C] | TM [°C] | Amplicon length [bp] |
|---|---|---|---|---|---|---|
| Cul4a | NM_001363450.1 | Forward Reverse |
TGATGCAGGACAGGGAGGTTC CCACACAGGCAATCAACGGT |
59 | 79.4 | 123 |
| Eef2 | NM_007907.2 | Forward Reverse |
CACAATCAAATCCACCGCC ATGGCCTGGAGAGTCGATGA |
60 | 80.7 | 122 |
| Hprt | NM_013556.2 | Forward Reverse |
TCAGTCAACGGGGGACATAAA GGGGCTGTACTGCTTAACCAG |
61 | 76.1 | 142 |
| Ywhaz | NM_011740.3 | Forward Reverse |
GAAAAGTTCTTGATCCCCAATGC TGTGACTGGTCCACAATTCCTT |
62 | 79.2 | 134 |
| Target genes | Sequence Accession Number | Direction | Sequence [5’-3’] |
TA [°C] | TM [°C] | bp |
|---|---|---|---|---|---|---|
| Arg1 | NM_007482.3 | Forward Reverse |
AAGGACAGCCTCGAGGAGGGGTAG TGGACCTCTCCCACCACACCA |
60.5 | 82 | 214 |
| Ccnd1 | NM_007631.2 | Forward Reverse |
GCGTACCCTGACACCAATCT CACAGACCTCCAGCATCCAG |
60 | 83 | 160 |
| Ctnnb1 | NM_001165902.1 | Forward Reverse |
CTAGCTGGTGGACTGCAGAAA TTCAGCACTCTGCTTGTGGT |
59 | 79.7 | 212 |
| Epcam | NM_008532.2 | Forward Reverse |
CATTTGCTCCAAACTGGCGT TTGTTCTGGATCGCCCCTTC |
60 | 79.5 | 125 |
| G6pdx | NM_008062.3 | Forward Reverse |
GGGAAGAGTTGTACCAGGGTG TCTTCAGGTAGAAGGCCATCCC |
59 | 80.1 | 142 |
| Keap1 | NM_001110305.1 | Forward Reverse |
GGCAGGACCAGTTGAACAGT CATAGCCTCCGAGGACGTAG |
59 | 87.5 | 138 |
| Nos2 | NM_010927.4 | Forward Reverse |
ACCCTAAGAGTCACCAAAATGGC TTGATCCTCACATACTGTGCACG |
60.5 | 82 | 118 |
| Nqo1 | NM_008706 | Forward Reverse |
AGAGAGTGCTCGTAGCAGGAT CTACCCCCAGTGGTGATAGAAA |
61.5 | 78 | 103 |
| Nrf2 | NM_010902.4 | Forward Reverse |
CCCAGCAGGACATGGATTTGA AGCTCATAGTCCTTCTGTCGC |
60 | 77.1 | 106 |
| Pd-l1 | NM_021893.3 | Forward Reverse |
CTCGCCTGCAGATAGTTCCC AGCCGTGATAGTAAACGCCC |
60 | 77.4 | 94 |
| Sox9 | NM_011448.4 | Forward Reverse |
GTGAAGAACGGACAAGCGGA GATTGCCCAGAGTGCTCGC |
60 | 84 | 148 |
| Vegf-α | NM_001110268.1 | Forward Reverse |
GCAGATGTGAATGCAGACCAAA GCGTGGTGGTGACATGGTTA |
59 | 83 | 154 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




