Submitted:
19 February 2026
Posted:
23 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Literature Review and Theoretical Background
2.1. Era of Traditional CAD Systems
2.2. The Deep Learning Revolution
2.3. Foundational Milestones: CheXNet and CheXpert
2.4. Foundations and Applications of Federated Learning in Healthcare
2.4.1. Conceptual Origins and Theoretical Development
2.4.2. Unique Challenges in Healthcare Applications
2.4.3. Pioneering Applications in Medical Imaging
2.5. Privacy-Preserving Techniques in Medical AI
2.5.1. Practical Implementations and Trade-off Considerations
3. Methodology and System Design
3.1. Guiding Principles
3.2. Adaptive Aggregation Architecture
3.2.1. Confidence-Weighted Aggregation for Classification Components
3.2.2. Structure-Aware Aggregation for Segmentation Components
3.2.3. Semantic Aggregation for Report Generation Components
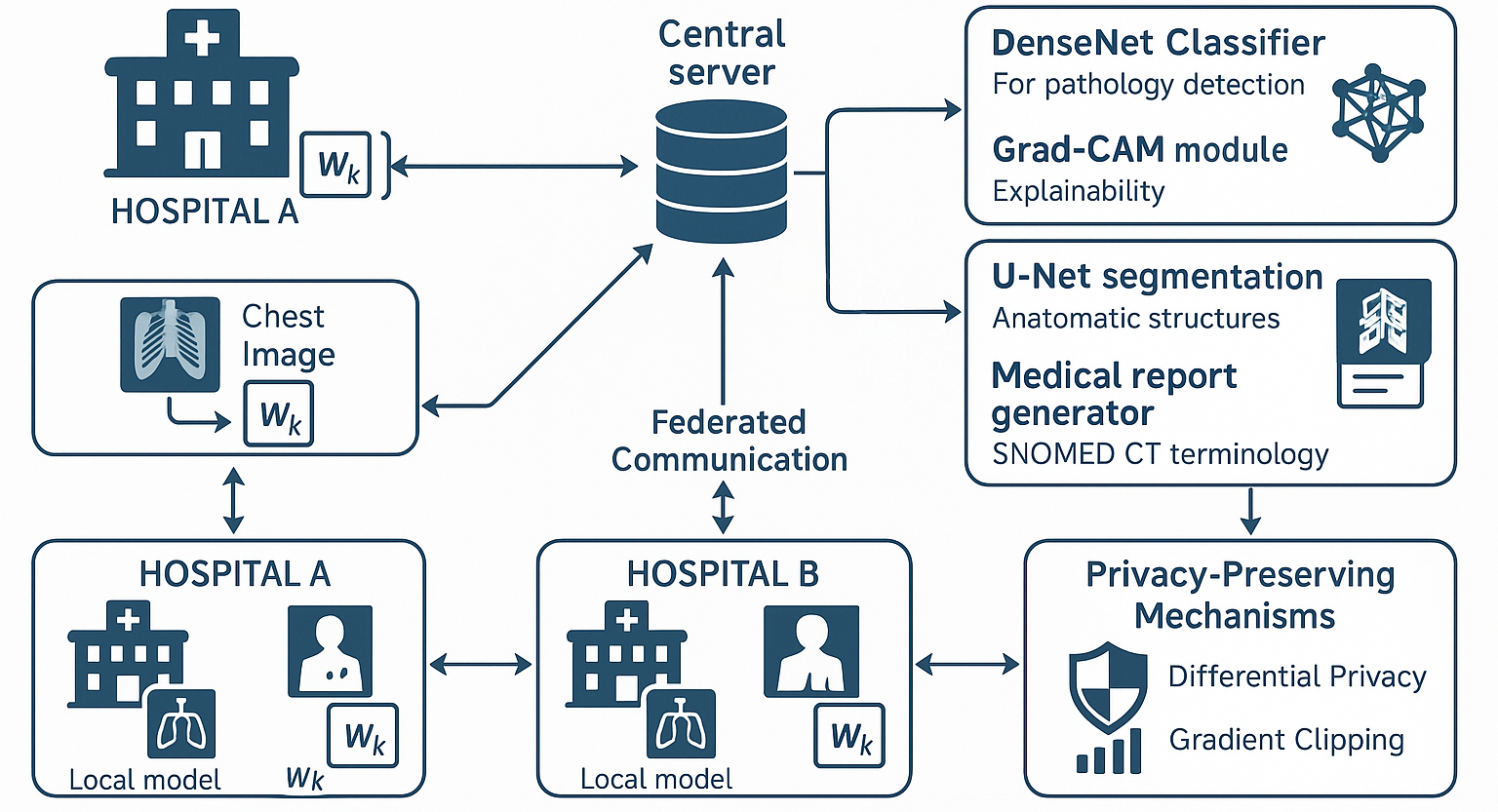
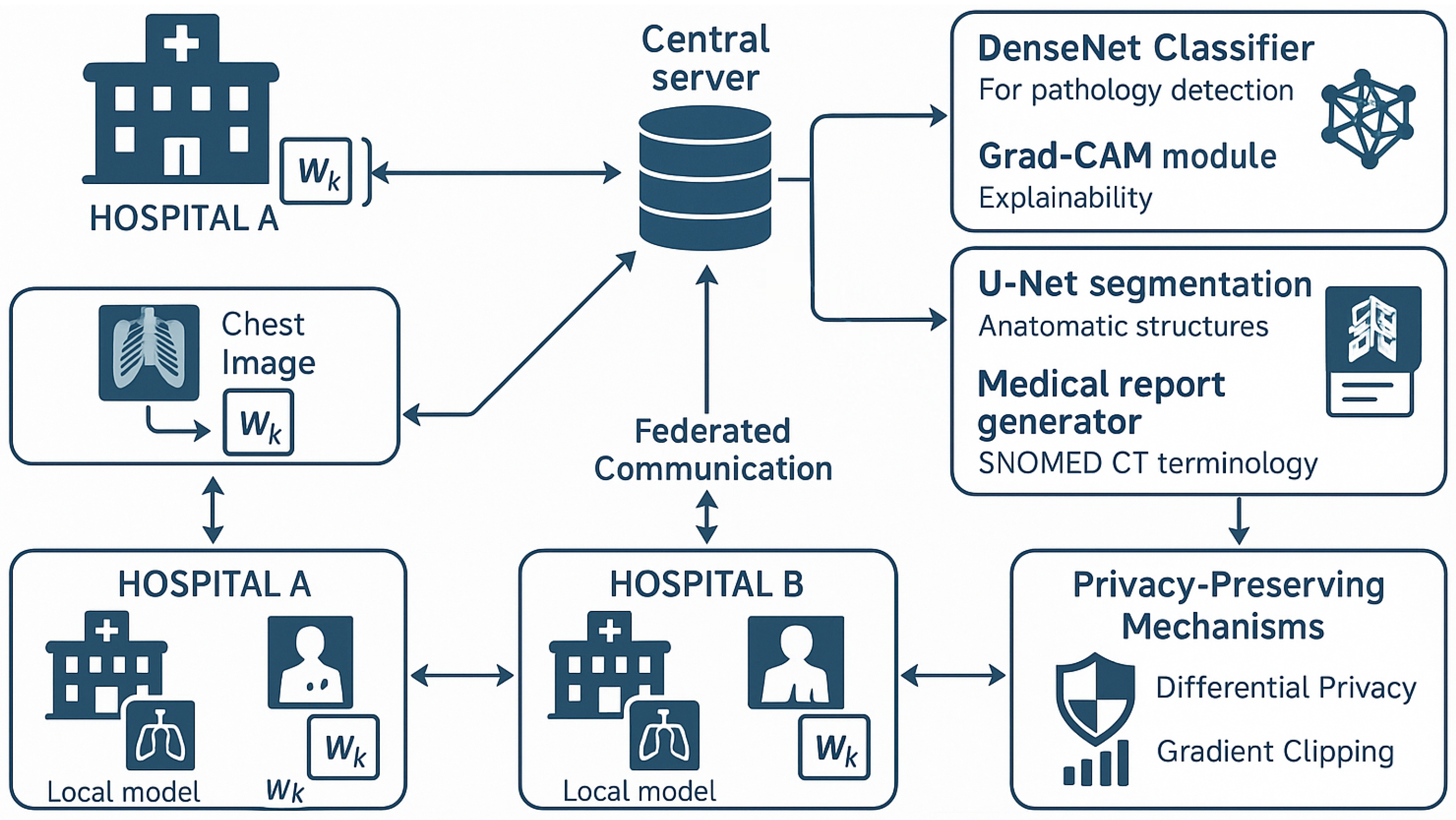
3.3. Architecture and Components of FedIHRAS
3.3.1. Local Node Architecture
3.3.2. Secure Communication Infrastructure
3.3.3. Multi-Layer Privacy Protection Framework
4. Validation Methodology and Experimental Setup
4.1. Comprehensive Experimental Design and Validation Protocol
4.2. Dataset Composition and Multi-Institutional Simulation
4.3. Adaptive Federated Training Protocol and Aggregation Strategies
5. Experimental Results and Performance Analysis
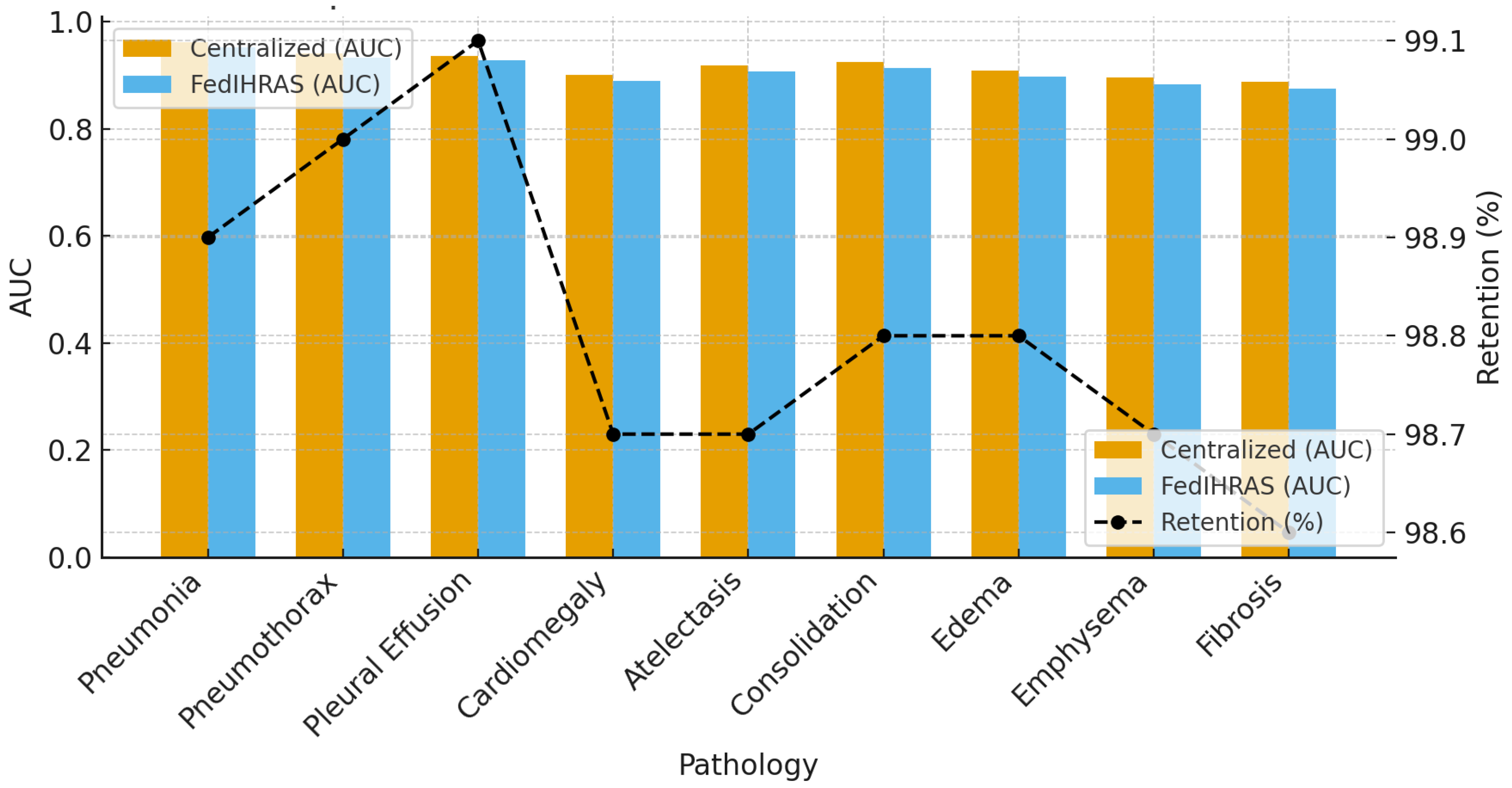
5.1. Pathology Classification Performance Analysis
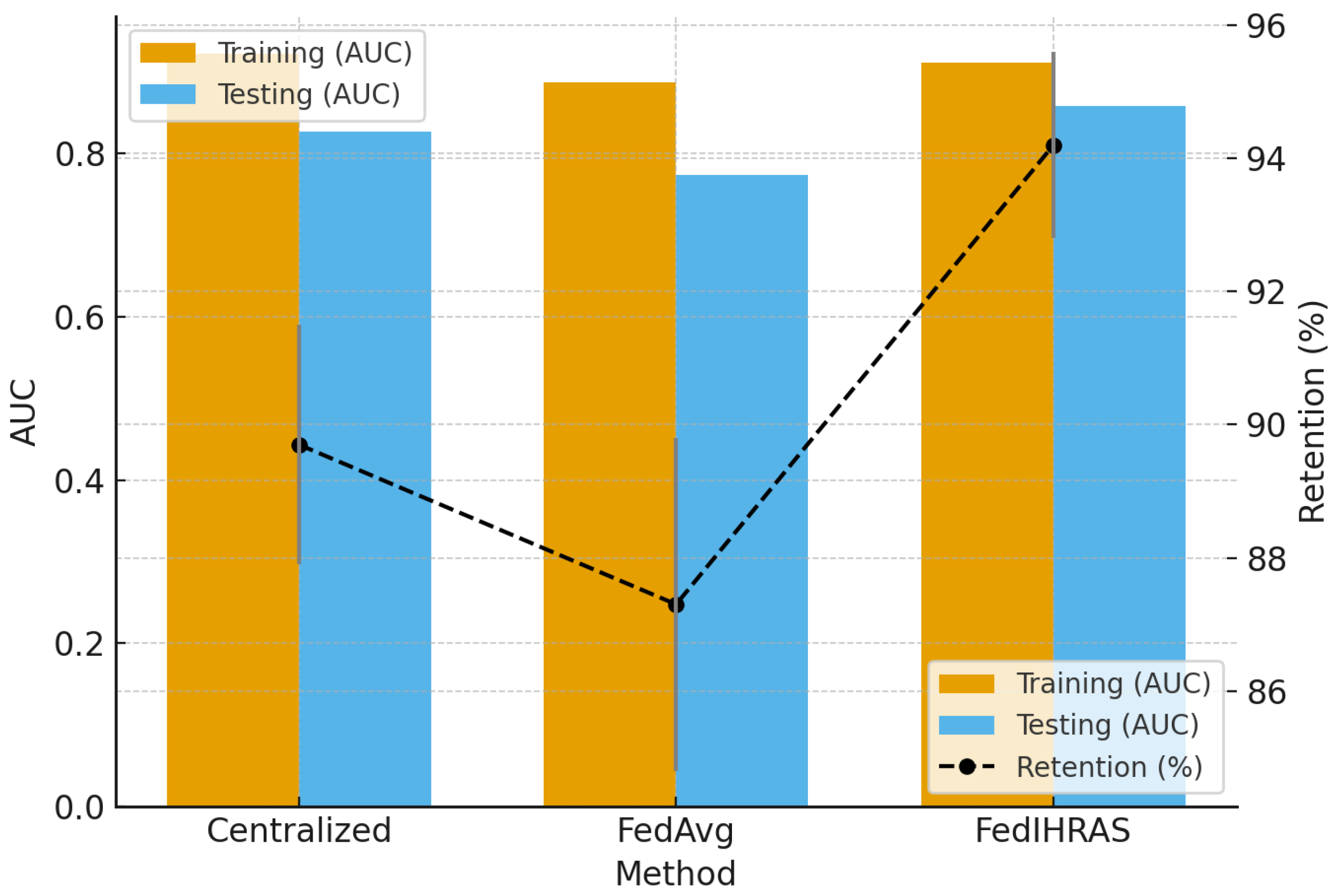
5.2. Cross-Institutional Generalization and Robustness Analysis
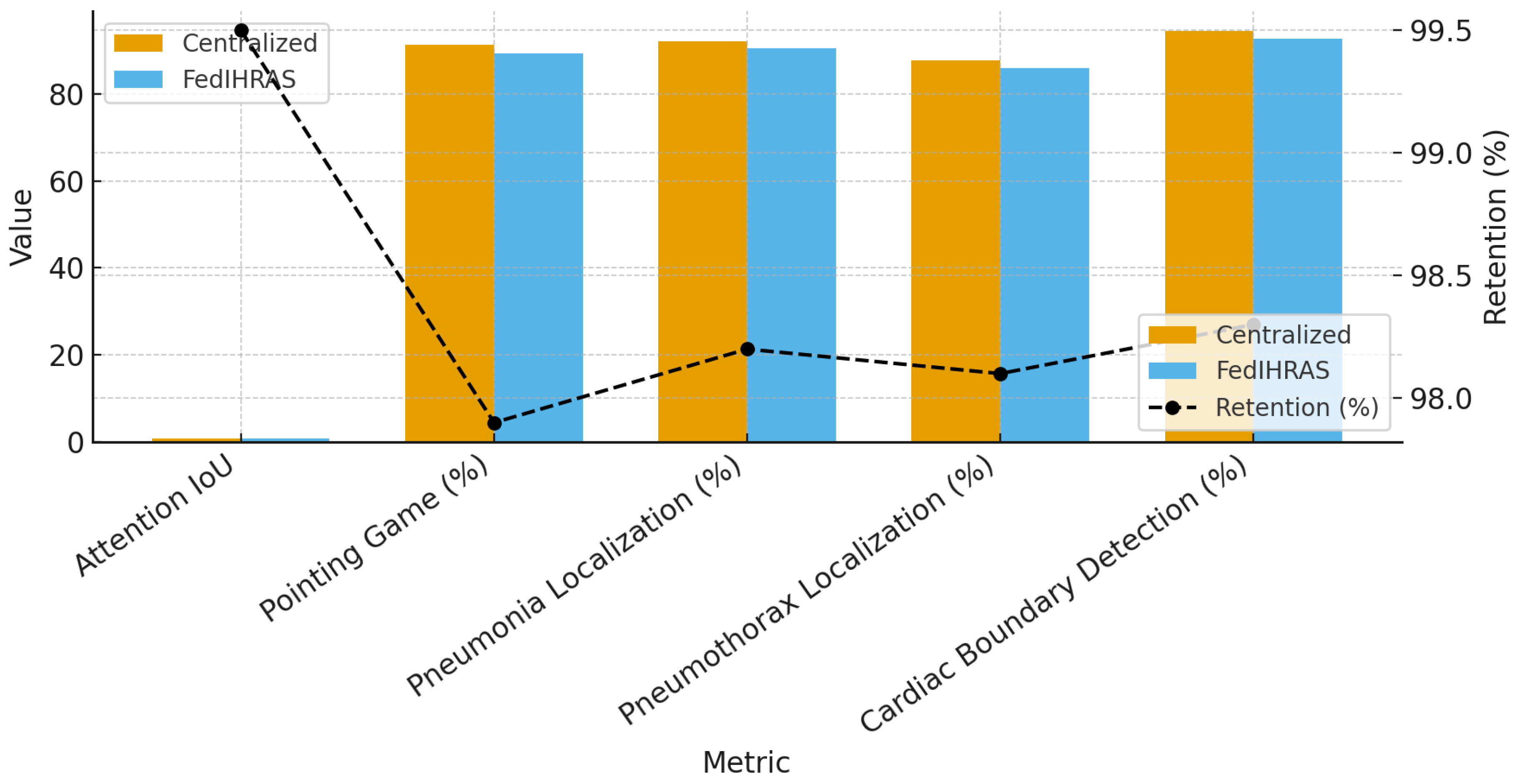
5.3. Preservation of Explainability and Clinical Interpretability (XAI)
5.4. Privacy Protection Effectiveness Analysis
5.5. Comprehensive Clinical Validation and Expert Evaluation
6. Discussion and Clinical Implications
6.1. Technical Contributions and Methodological Innovations
6.2. Clinical Impact and Healthcare Transformation
7. Conclusions
7.1. Limitations and Future Research Directions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rodrigues, G.A.P.; Serrano, A.L.M.; Bispo, G.D.; Filho, G.P.R.; Gonçalves, V.P.; Meneguette, R.I. IHRAS: Automated Medical Report Generation from Chest X-Rays via Classification, Segmentation, and LLMs. Bioengineering 2025, 12, 795. [Google Scholar] [CrossRef] [PubMed]
- Rao, A.; Kim, J.; Kamineni, M.; Pang, M.; Lie, W.; Dreyer, K.J.; Succi, M.D. Evaluating GPT as an adjunct for radiologic decision making: GPT-4 versus GPT-3.5 in a breast imaging pilot. Journal of the American College of Radiology 2023, 20, 990–997. [Google Scholar] [CrossRef] [PubMed]
- European Union. General Data Protection Regulation (GDPR), 2016.
- McMahan, H.B.; Moore, E.; Ramage, D.; Hampson, S.; y Arcas, B.A. Communication-efficient learning of deep networks from decentralized data. In Proceedings of the Proceedings of the 20th International Conference on Artificial Intelligence and Statistics, 2017, Vol. 54, pp. 1273–1282.
- Li, T.; Sahu, A.K.; Talwalkar, A.; Smith, V. Federated learning: Challenges, methods, and future directions. IEEE Signal Processing Magazine 2020, 37, 50–60. [Google Scholar] [CrossRef]
- Vuokko, R.; Vakkuri, A.; Palojoki, S. Systematized nomenclature of medicine–clinical terminology (SNOMED CT) clinical use cases in the context of electronic health record systems: systematic literature review. JMIR medical informatics 2023, 11, e43750. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zeng, D.; Luo, J.; Fu, X.; Chen, G.; Xu, Z.; King, I. A survey of trustworthy federated learning: Issues, solutions, and challenges. ACM Transactions on Intelligent Systems and Technology 2024, 15, 1–47. [Google Scholar] [CrossRef]
- Wang, S.; Hosseinalipour, S.; Aggarwal, V.; Brinton, C.G.; Love, D.J.; Su, W.; Chiang, M. Toward cooperative federated learning over heterogeneous edge/fog networks. IEEE Communications Magazine 2023, 61, 54–60. [Google Scholar] [CrossRef]
- Pierre, K.; Haneberg, A.G.; Kwak, S.; Peters, K.R.; Hochhegger, B.; Sananmuang, T.; Tunlayadechanont, P.; Tighe, P.J.; Mancuso, A.; Forghani, R. Applications of artificial intelligence in the radiology roundtrip: process streamlining, workflow optimization, and beyond. In Proceedings of the Seminars in Roentgenology. Elsevier, 2023, Vol. 58, pp. 158–169.
- Doi, K. Computer-aided diagnosis in medical imaging: Historical review, current status and future potential. Computerized Medical Imaging and Graphics 2007, 31, 198–211. [Google Scholar] [CrossRef] [PubMed]
- Yin, F.F.; Giger, M.L.; Doi, K.; Metz, C.E.; Vyborny, C.J.; Schmidt, R.A. Computerized detection of masses in digital mammograms: Analysis of bilateral subtraction images. Medical physics 1991, 18, 955–963. [Google Scholar] [CrossRef] [PubMed]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet classification with deep convolutional neural networks. In Proceedings of the Advances in Neural Information Processing Systems, 2012, Vol. 25, pp. 1097–1105.
- Maity, A.; Nair, T.R.; Mehta, S.; Prakasam, P. Automatic lung parenchyma segmentation using a deep convolutional neural network from chest X-rays. Biomedical Signal Processing and Control 2022, 73, 103398. [Google Scholar] [CrossRef]
- Santomartino, S.M.; Hafezi-Nejad, N.; Parekh, V.S.; Yi, P.H. Performance and usability of code-free deep learning for chest radiograph classification, object detection, and segmentation. Radiology: Artificial Intelligence 2023, 5, e220062. [Google Scholar] [CrossRef] [PubMed]
- Nicolson, A.; Dowling, J.; Koopman, B. Improving chest X-ray report generation by leveraging warm starting. Artificial intelligence in medicine 2023, 144, 102633. [Google Scholar] [CrossRef] [PubMed]
- Debelee, T.G. Skin lesion classification and detection using machine learning techniques: a systematic review. Diagnostics 2023, 13, 3147. [Google Scholar] [CrossRef] [PubMed]
- Rajpurkar, P.; et al. CheXNet: Radiologist-level pneumonia detection on chest X-rays with deep learning. arXiv preprint arXiv:1711.05225 2017. arXiv:1711.05225.
- Fanni, S.C.; Marcucci, A.; Volpi, F.; Valentino, S.; Neri, E.; Romei, C. Artificial intelligence-based software with CE mark for chest X-ray interpretation: Opportunities and challenges. Diagnostics 2023, 13, 2020. [Google Scholar] [CrossRef] [PubMed]
- Irvin, J.; et al. CheXpert: A large chest radiograph dataset with uncertainty labels and expert comparison. In Proceedings of the Proceedings of the AAAI Conference on Artificial Intelligence, 2019, Vol. 33, pp. 590–597.
- Akhter, Y.; Singh, R.; Vatsa, M. AI-based radiodiagnosis using chest X-rays: A review. Frontiers in big data 2023, 6, 1120989. [Google Scholar] [CrossRef] [PubMed]
- Torab-Miandoab, A.; Samad-Soltani, T.; Jodati, A.; Rezaei-Hachesu, P. Interoperability of heterogeneous health information systems: a systematic literature review. BMC medical informatics and decision making 2023, 23, 18. [Google Scholar]
- Sheller, M.J.; et al. Federated learning in medicine: facilitating multi-institutional collaborations without sharing patient data. Scientific Reports 2020, 10, 12598. [Google Scholar] [CrossRef]
- Dou, Q.; et al. Federated deep learning for detecting COVID-19 lung abnormalities in CT: a privacy-preserving multinational validation study. NPJ Digital Medicine 2021, 4, 60. [Google Scholar] [CrossRef]
- Naz, S.; Phan, K.T.; Chen, Y.P.P. A comprehensive review of federated learning for COVID-19 detection. International Journal of Intelligent Systems 2022, 37, 2371–2392. [Google Scholar] [CrossRef] [PubMed]
- Rajpoot, R.; Gour, M.; Jain, S.; Semwal, V.B. Integrated ensemble CNN and explainable AI for COVID-19 diagnosis from CT scan and X-ray images. Scientific Reports 2024, 14, 24985. [Google Scholar] [CrossRef]
- Dwork, C.; McSherry, F.; Nissim, K.; Smith, A. Calibrating noise to sensitivity in private data analysis. In Proceedings of the Theory of Cryptography Conference, 2006, pp. 265–284.
- Abadi, M.; et al. Deep learning with differential privacy. In Proceedings of the Proceedings of the 2016 ACM SIGSAC Conference on Computer and Communications Security, 2016, pp. 308–318.
| Attack | FedIHRAS | Baseline | Reduction (%) |
|---|---|---|---|
| Inference Attacks (Metric: Success Rate %) | |||
| Membership Inference | 52.3 | 78.4 | 33.3 |
| Property Inference | 48.7 | 71.2 | 31.6 |
| Reconstruction Attacks (Metric: SSIM) | |||
| Model Inversion | 0.147 | 0.634 | 76.8 |
| Gradient Leakage | 0.089 | 0.523 | 83.0 |
| Attribute Recovery | 0.203 | 0.745 | 72.8 |
| Pathology | Acc. (%) | Sens. (%) | Spec. (%) |
|---|---|---|---|
| Pneumonia | 94.6 | 92.8 | 96.4 |
| Pneumothorax | 93.2 | 91.5 | 94.9 |
| Pleural Effusion | 95.1 | 93.7 | 96.5 |
| Cardiomegaly | 91.8 | 89.4 | 94.2 |
| Atelectasis | 92.5 | 90.1 | 94.9 |
| Average | 94.3 | 91.5 | 95.4 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




