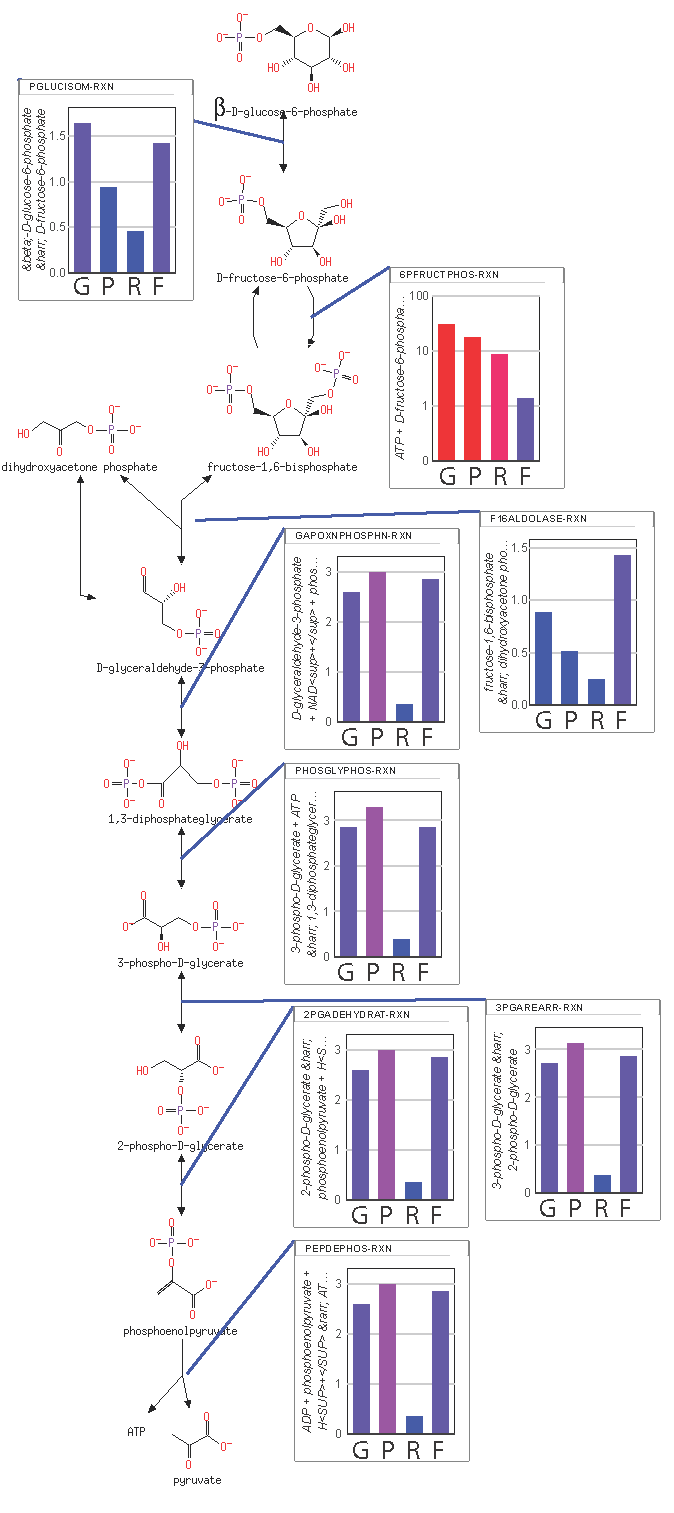
We report the application of a recently proposed approach for modeling biological systems using a maximum entropy production rate principle in lieu of having in vivo rate constants. The method is applied in four steps: (1) a new ODE-based optimization approach based on Marcelin’s 1910 mass action equation is used to obtain the maximum entropy distribution, (2) the predicted metabolite concentrations are compared to those generally expected from experiment using a loss function from which post-translational regulation of enzymes is inferred, (3) the system is re-optimized with the inferred regulation from which rate constants are determined from the metabolite concentrations and reaction fluxes, and finally (4) a full ODE-based, mass action simulation with rate parameters and allosteric regulation is obtained. From the last step, the power characteristics and resistance of each reaction can be determined. The method is applied to the central metabolism of Neurospora crassa and the flow of material through the three competing pathways of upper glycolysis, the non-oxidative pentose phosphate pathway, and the oxidative pentose phosphate pathway are evaluated as a function of the NADP/NADPH ratio. It is predicted that regulation of phosphofructokinase (PFK) and flow through the pentose phosphate pathway are essential for preventing an extreme level of fructose 1, 6-bisphophate accumulation. Such an extreme level of fructose 1,6-bisphophate would otherwise result in a glassy cytoplasm with limited diffusion, dramatically decreasing the entropy and energy production rate and, consequently, biological competitiveness.