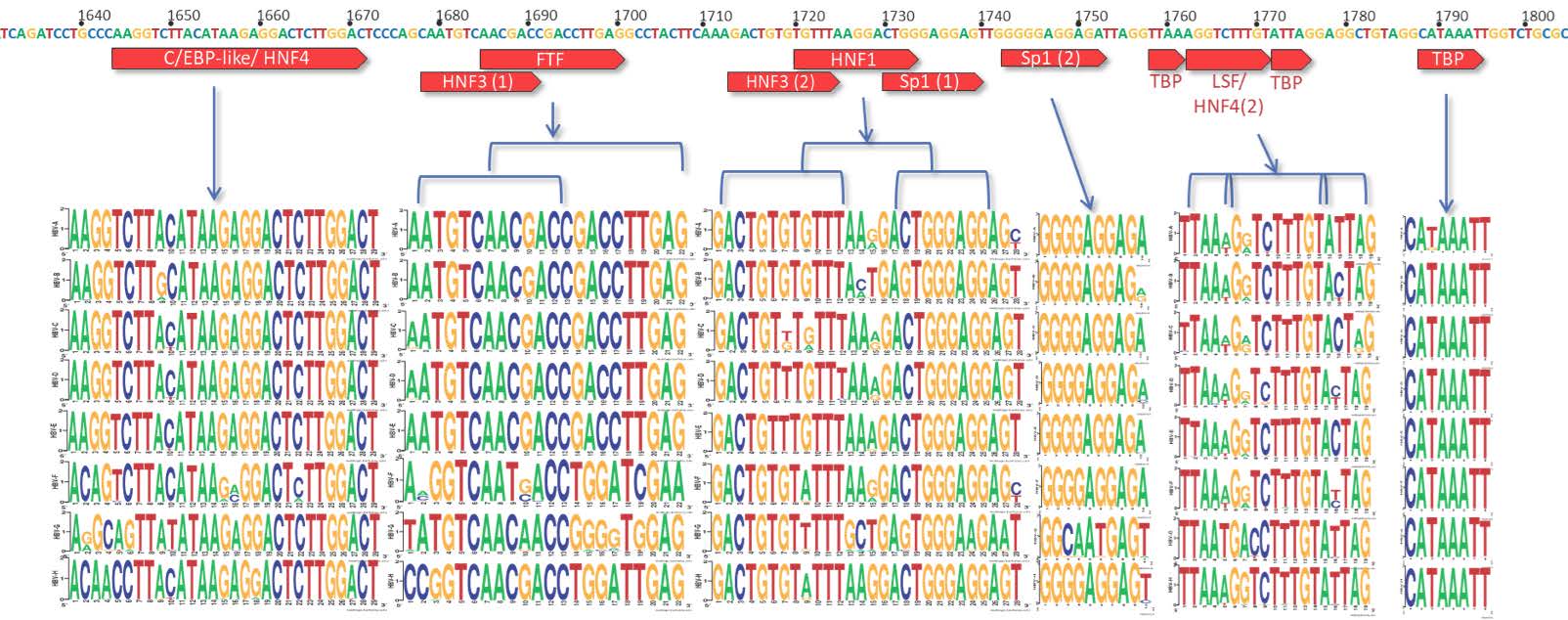
Over 250 million people are infected chronically with hepatitis B virus (HBV), the leading cause of liver cancer worldwide. HBV persists due in part to its compact, stable minichromosome, the covalently-closed, circular DNA (cccDNA), which resides in the hepatocytes’ nuclei. Current therapies target downstream replication products, however, a true virological cure will require targeting the cccDNA. Finding targets on such a small, compact genome is challenging. For HBV, to remain replication-competent, it needs to maintain nucleotide fidelity in key regions, such as the promoter regions, to ensure that it can continue to utilize the necessary host proteins. HBVdb (HBV database) is a repository of HBV sequences spanning all genotypes (A-H) amplified from clinical samples, and hence implying an extensive collection of replication-competent viruses. Here, we analyzed the HBV sequences from HBVdb using bioinformatics tools to comprehensively assess the HBV core and X promoter regions amongst the nearly 70,000 HBV sequences for highly-conserved nucleotides and variant frequencies. Notably, there is a high degree of nucleotide conservation within specific segments of these promoter regions highlighting their importance in potential host protein-viral interactions and thus the virus’ viability. Such findings may have key implications for designing antivirals to target these areas.