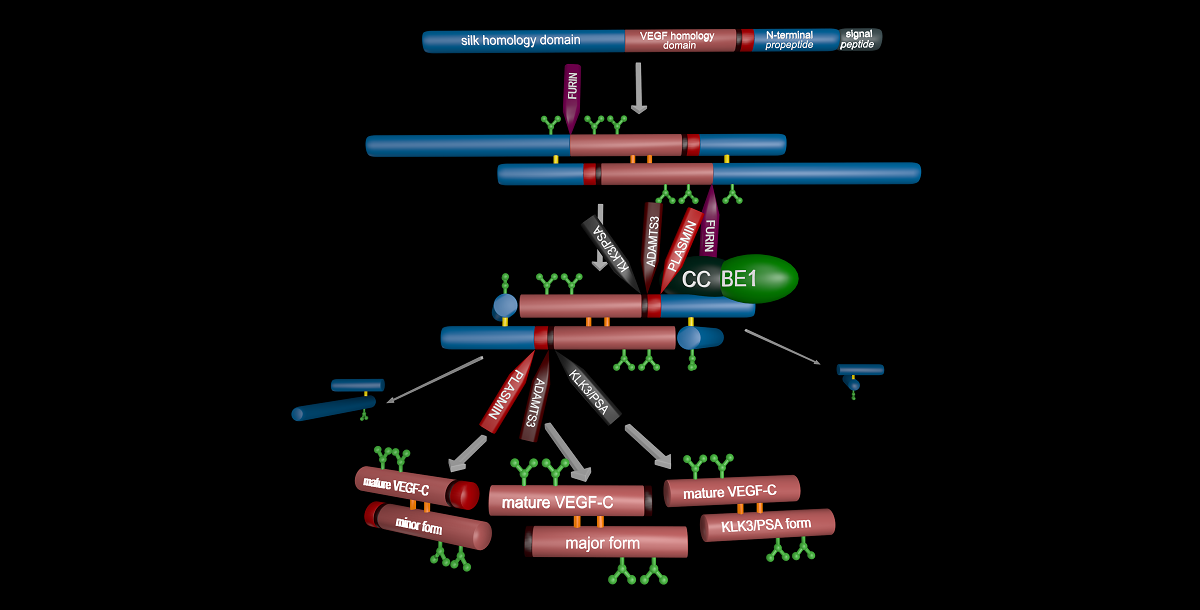
Specific proteolytic cleavages turn on, modify, or turn off the activity of vascular endothelial growth factors (VEGFs). Proteolysis is most prominent among the lymphangiogenic VEGF-C and VEGF-D, which are synthesized as precursors that need to undergo enzymatic removal of their C- and N-terminal propeptides before they can activate their receptors. The activating cleavage of VEGF-C is mediated by at least five different proteases: plasmin, ADAMTS3, prostate-specific antigen, cathepsin D, and thrombin. All of these proteases except for ADAMTS3 can also activate VEGF-D. Processing by different proteases results in distinct forms of the "mature" growth factors, which differ in affinity and receptor activation potential. The “default” VEGF-C-activating enzyme ADAMTS3 does not activate VEGF-D and therefore, VEGF-C and VEGF-D do function in different contexts. VEGF-C itself is also regulated in different contexts by different proteases. During embryonic development, ADAMTS3 activates VEGF-C. In contrast, thrombin and plasmin likely activate VEGF-C/-D during tissue injury-induced lymphangiogenesis, and PSA and cathepsin D perhaps during tumor-associated pathological lymphangiogenesis. Additionally, cathepsin D from saliva might activate latent VEGF-C/-D upon wound licking, thereby accelerating healing. Similar to tyrosine kinase receptors and VEGFs themselves, these activating proteases could be targeted to modulate angiogenesis and lymphangiogenesis in relevant diseases.