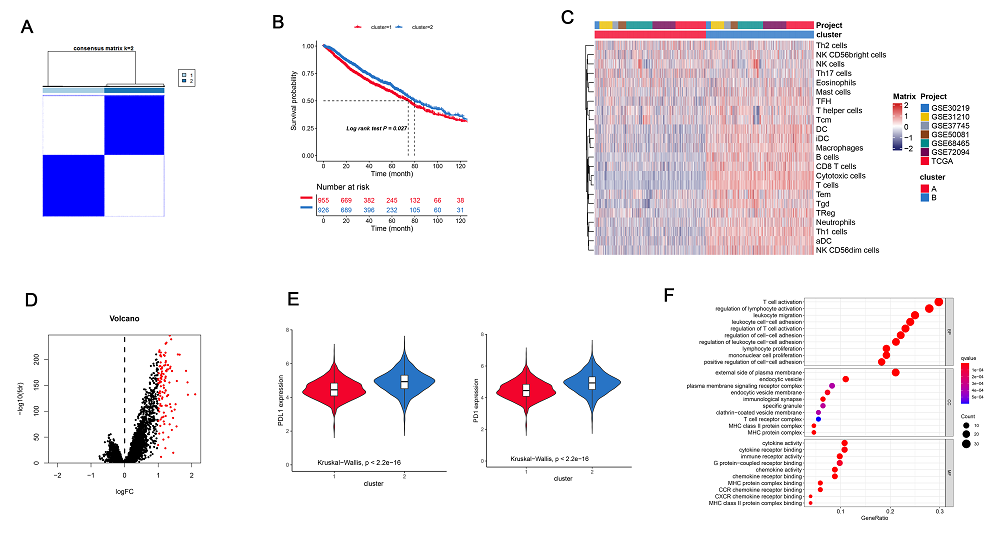
In a recent study, the PD-1 inhibitor has been widely used in clinical trials and shown to improve various cancers. However, PD-1/PD-L1 inhibitors showed a low response rate and showed to be effective for a small number of cancer patients. Thus, it is important to identify key genes, which can enhance the PD-1/PD-L1 response for promoting immunotherapy. Here, we used ssGSEA and unsupervised clustering analysis to identify three clusters to show different immune cell infiltration status, prognosis, and biological action. The cluster C showed a better survival rate, high immune cells infiltration, and immunotherapy effect enriched in a variety of immune active pathways, including T and B cell signal receptors. Besides, it showed more immune subtypes C2 and C3. Further, we used WGCNA analysis to confirm the cluster C correlated genes. The red module highly correlated with cluster C for 111 genes which were enriched in a variety of immune-related pathways. To pick candidate genes in SD/PD and CR/PR patients, we used the Least Absolute Shrinkage and SVM-RFE algorithms. In conclusion, our LASSO analysis and SVM-RFE based research identified targets with better prognosis, activated immune-related pathways, and better immunotherapy. The KLRC3 was identified as the key gene which can efficiently respond to immunotherapy with greater efficacy and better prognosis.