Submitted:
11 July 2023
Posted:
12 July 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. Clinical profiles of CAD patients and control subjects
2.2. Genotype frequencies of TS gene polymorphisms in CAD patients and control subjects
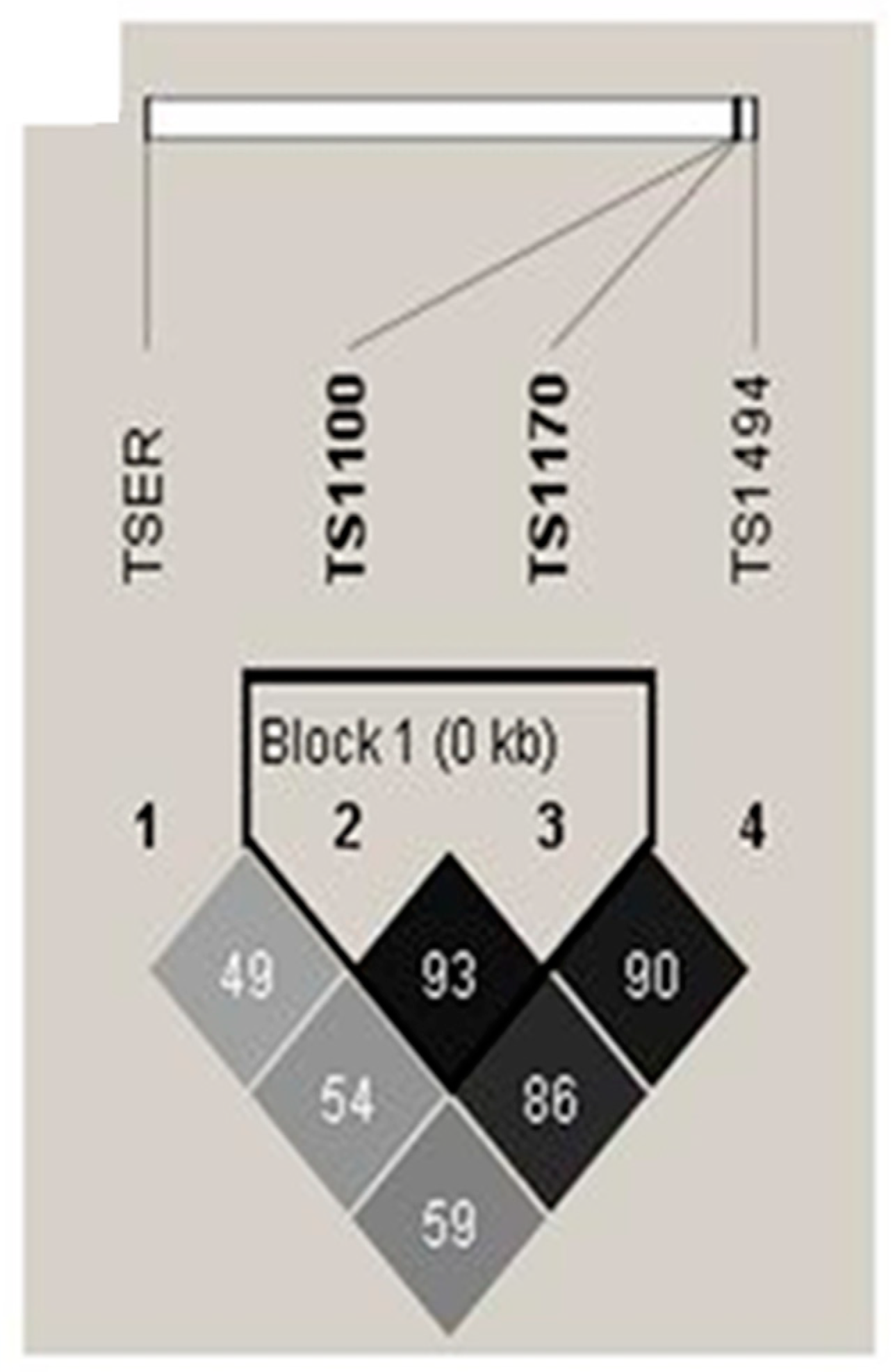
2.3. Haplotype and genotype combination analysis of TSER and TS 3’-UTR gene polymorphisms between CAD patients and control subjects
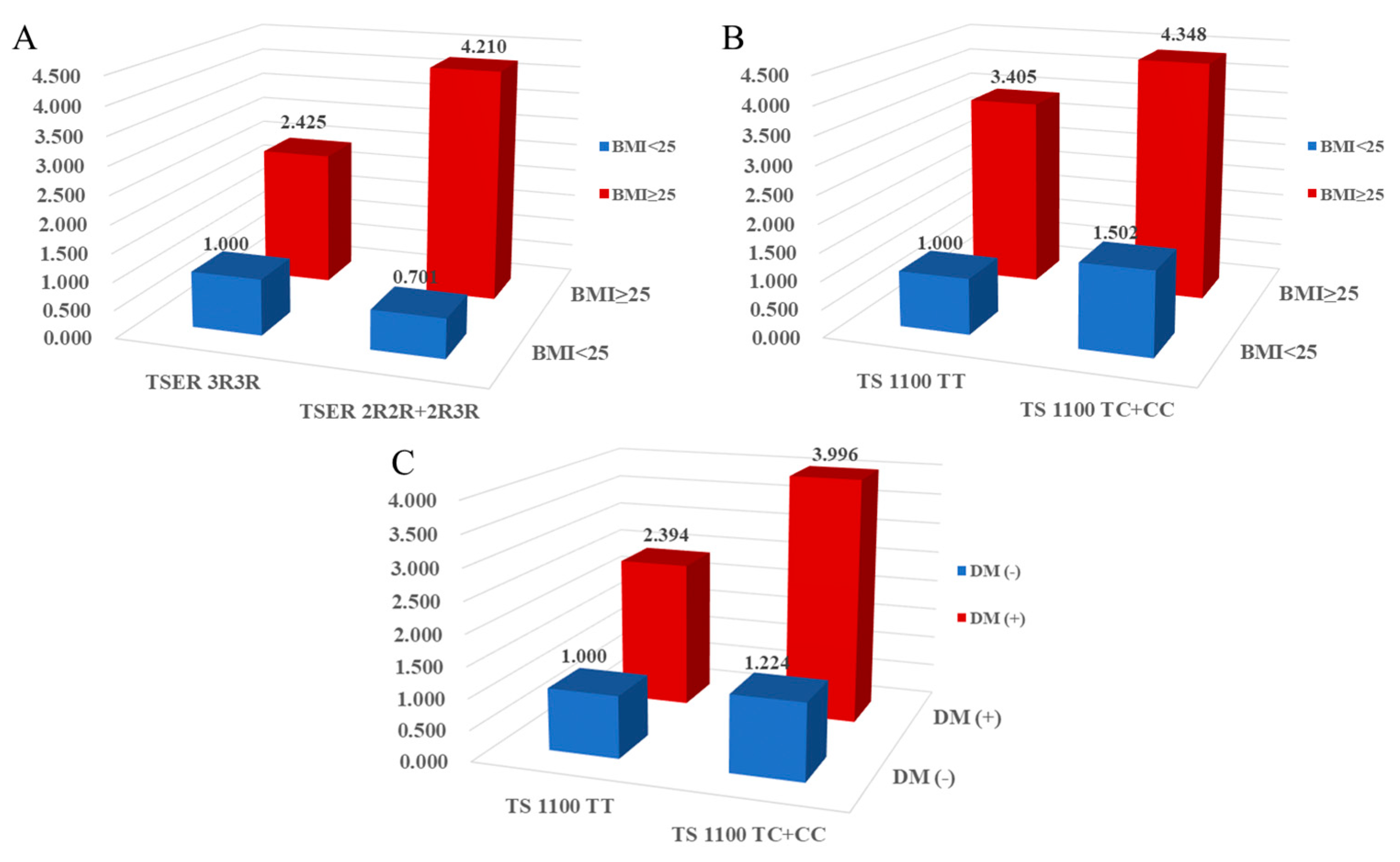
2.4. Combined effects between TS gene polymorphisms and environmental factors on CAD prevalence
2.5. Baseline characteristics and genotype frequencies in CAD patients and control subjects stratified by replication groups
3. Discussion
4. Materials and Methods
4.1. Study approval and population
4.2. Assessment of biochemical factors
4.3. Genotyping
4.4. Statistical analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Fernandez-Ortiz, A.; Fuster, V. Pathophysiology of coronary artery disease. Clin Geriatr Med 1996, 12, 1–21. [Google Scholar] [PubMed]
- Libby, P.; Theroux, P. Pathophysiology of coronary artery disease. Circulation 2005, 111, 3481–3488. [Google Scholar]
- McCullough, P.A. Coronary artery disease. Clin J Am Soc Nephrol 2007, 2, 611–616. [Google Scholar] [PubMed]
- Menees, D.S.; Bates, E.R. Evaluation of patients with suspected coronary artery disease. Coron Artery Dis 2010, 21, 386–390. [Google Scholar]
- Stolker, J.M.; Lim, M.J. Update in the management of coronary artery disease. Mo Med 2012, 109, 137–141. [Google Scholar] [PubMed]
- Perdoncin, E.; Duvernoy, C. Treatment of Coronary Artery Disease in Women. Methodist Debakey Cardiovasc J 2017, 13, 201–208. [Google Scholar] [PubMed]
- Huang, J.; Huang, S.; Li, J.; Li, M.; Gong, L.; Li, T.; Gu, L. CALM1 rs3179089 polymorphism might contribute to coronary artery disease susceptibility in Chinese male: a case-control study. Genes Genomics 2022, 44, 415–423. [Google Scholar] [PubMed]
- Liang, T.; Liang, A.; Zhang, X.; Wang, Q.; Wu, H.; He, J.; Jin, T. The association study between CYP20A1, CYP4F2, CYP2D6 gene polymorphisms and coronary heart disease risk in the Han population in southern China. Genes Genomics 2022, 44, 1125–1135. [Google Scholar]
- Lievers, K.J.; Kluijtmans, L.A.; Blom, H.J. Genetics of hyperhomocysteinaemia in cardiovascular disease. Ann Clin Biochem 2003, 40 (Pt 1), 46–59. [Google Scholar] [CrossRef]
- Kim, I.J.; Kim, S.H.; Cha, D.H.; Lim, S.W.; Moon, J.Y.; Kim, J.O.; Ryu, C.S.; Park, H.S.; Sung, J.H.; Kim, N.K. Association of COX2 -765G>C promoter polymorphism and coronary artery disease in Korean population. Genes Genomics 2019, 41, 1055–1062. [Google Scholar]
- Biselli, P.M.; Guerzoni, A.R.; de Godoy, M.F.; Eberlin, M.N.; Haddad, R.; Carvalho, V.M.; Vannucchi, H.; Pavarino-Bertelli, E.C.; Goloni-Bertollo, E.M. Genetic polymorphisms involved in folate metabolism and concentrations of methylmalonic acid and folate on plasma homocysteine and risk of coronary artery disease. J Thromb Thrombolysis 2010, 29, 32–40. [Google Scholar]
- Trinh, B.N.; Ong, C.N.; Coetzee, G.A.; Yu, M.C.; Laird, P.W. Thymidylate synthase: a novel genetic determinant of plasma homocysteine and folate levels. Hum Genet 2002, 111, 299–302. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Schmitz, J.C.; Lin, X.; Tai, N.; Yan, W.; Farrell, M.; Bailly, M.; Chen, T.; Chu, E. Thymidylate synthase as a translational regulator of cellular gene expression. Biochim Biophys Acta 2002, 1587, 174–182. [Google Scholar] [CrossRef] [PubMed]
- Tai, N.; Schmitz, J.C.; Liu, J.; Lin, X.; Bailly, M.; Chen, T.M.; Chu, E. Translational autoregulation of thymidylate synthase and dihydrofolate reductase. Front Biosci 2004, 9, 2521–2526. [Google Scholar] [CrossRef] [PubMed]
- Farnham, P.J.; Slansky, J.E.; Kollmar, R. The role of E2F in the mammalian cell cycle. Biochim Biophys Acta 1993, 1155, 125–131. [Google Scholar]
- Chen, J.J.; Roberson, P.K.; Schell, M.J. The false discovery rate: a key concept in large-scale genetic studies. Cancer Control 2010, 17, 58–62. [Google Scholar]
- Mei, S.; Karimnezhad, A.; Forest, M.; Bickel, D.R.; Greenwood, C.M.T. The performance of a new local false discovery rate method on tests of association between coronary artery disease (CAD) and genome-wide genetic variants. PLoS One 2017, 12, e0185174. [Google Scholar]
- White, H.; Boden-Albala, B.; Wang, C.; Elkind, M.S.; Rundek, T.; Wright, C.B.; Sacco, R.L. Ischemic stroke subtype incidence among whites, blacks, and Hispanics: the Northern Manhattan Study. Circulation 2005, 111, 1327–1331. [Google Scholar] [CrossRef]
- Hartmann, A.; Rundek, T.; Mast, H.; Paik, M.C.; Boden-Albala, B.; Mohr, J.P.; Sacco, R.L. Mortality and causes of death after first ischemic stroke: the Northern Manhattan Stroke Study. Neurology 2001, 57, 2000–2005. [Google Scholar] [CrossRef]
- Touze, E.; Varenne, O.; Chatellier, G.; Peyrard, S.; Rothwell, P.M.; Mas, J.L. Risk of myocardial infarction and vascular death after transient ischemic attack and ischemic stroke: a systematic review and meta-analysis. Stroke 2005, 36, 2748–2755. [Google Scholar] [CrossRef]
- Hoshino, A.; Nakamura, T.; Enomoto, S.; Kawahito, H.; Kurata, H.; Nakahara, Y.; Ijichi, T. Clinical utility of evaluating intracranial artery stenosis and silent brain infarction to predict the presence of subclinical coronary artery disease in ischemic stroke patients. Intern Med 2008, 47, 1775–1781. [Google Scholar] [PubMed]
- Hammad, H.; Sarkar, M.; Gupta, N.; Ardalan, B.; Subbarayan, P.R. The presence of three repeats in the 5' UTR region of thymidylate synthase (TS) is associated with increased TS mRNA expression in cultured human cancer cell lines in vitro. Oncol Rep 2012, 27, 246–249. [Google Scholar] [PubMed]
- Ho, V.; Massey, T.E.; King, W.D. Effects of methionine synthase and methylenetetrahydrofolate reductase gene polymorphisms on markers of one-carbon metabolism. Genes Nutr 2013, 8, 571–580. [Google Scholar]
- Kim, O.J.; Hong, S.P.; Ahn, J.Y.; Hong, S.H.; Hwang, T.S.; Kim, S.O.; Yoo, W.; Oh, D.; Kim, N.K. Influence of combined methionine synthase (MTR 2756A > G) and methylenetetrahydrofolate reductase (MTHFR 677C > T) polymorphisms to plasma homocysteine levels in Korean patients with ischemic stroke. Yonsei Med J 2007, 48, 201–209. [Google Scholar] [PubMed]
- Brevik, A.; Vollset, S.E.; Tell, G.S.; Refsum, H.; Ueland, P.M.; Loeken, E.B.; Drevon, C.A.; Andersen, L.F. Plasma concentration of folate as a biomarker for the intake of fruit and vegetables: the Hordaland Homocysteine Study. Am J Clin Nutr 2005, 81, 434–439. [Google Scholar] [PubMed]
- McCully, K.S. Vascular pathology of homocysteinemia: implications for the pathogenesis of arteriosclerosis. Am J Pathol 1969, 56, 111–128. [Google Scholar]
- Joachim, E.; Goldenberg, N.A.; Bernard, T.J.; Armstrong-Wells, J.; Stabler, S.; Manco-Johnson, M.J. The methylenetetrahydrofolate reductase polymorphism (MTHFR c.677C>T) and elevated plasma homocysteine levels in a U.S. pediatric population with incident thromboembolism. Thromb Res 2013, 132, 170–174. [Google Scholar]
- Park, S.Y.; An, S.A.; Lee, H.B.; Kim, Y.; Kim, N.K.; Kim, S.H.; Kim, O.J.; Oh, S.H. Different impact of hyperhomocysteinemia on cerebral small vessel ischemia and cervico-cerebral atherosclerosis in non-stroke individuals. Thromb Res 2013, 131, e12-6. [Google Scholar] [CrossRef]
- Nazki, F.H.; Sameer, A.S.; Ganaie, B.A. Folate: metabolism, genes, polymorphisms and the associated diseases. Gene 2014, 533, 11–20. [Google Scholar]
- Blom, H.J.; Smulders, Y. Overview of homocysteine and folate metabolism. With special references to cardiovascular disease and neural tube defects. J Inherit Metab Dis 2011, 34, 75–81. [Google Scholar]
- Boni, V.; Bitarte, N.; Cristobal, I.; Zarate, R.; Rodriguez, J.; Maiello, E.; Garcia-Foncillas, J.; Bandres, E. miR-192/miR-215 influence 5-fluorouracil resistance through cell cycle-mediated mechanisms complementary to its post-transcriptional thymidilate synthase regulation. Mol Cancer Ther 2010, 9, 2265–2275. [Google Scholar] [PubMed]
- Wu, C.; Gong, Y.; Sun, A.; Zhang, Y.; Zhang, C.; Zhang, W.; Zhao, G.; Zou, Y.; Ge, J. The human MTHFR rs4846049 polymorphism increases coronary heart disease risk through modifying miRNA binding. Nutr Metab Cardiovasc Dis 2013, 23, 693–698. [Google Scholar] [CrossRef]
- Duan, L.; Liu, C.; Hu, J.; Liu, Y.; Wang, J.; Chen, G.; Li, Z.; Chen, H. Epigenetic mechanisms in coronary artery disease: The current state and prospects. Trends Cardiovasc Med 2018, 28, 311–319. [Google Scholar]
- Labbaf, A.; Ghaedi, H.; Alipoor, B.; Omrani, M.D.; Kazerouni, F.; Shanaki, M.; Ghaffarzadeh, M.; Pashaiefar, H.; Rahimipour, A. The pre-mir-499 Variant rs3746444 May Contribute to Coronary Artery Disease Susceptibility: a Case-Control and Meta-Analysis Study. Clin Lab 2017, 63, 587–595. [Google Scholar] [CrossRef]
- Yuan, M.; Zhan, Q.; Duan, X.; Song, B.; Zeng, S.; Chen, X.; Yang, Q.; Xia, J. A functional polymorphism at miR-491-5p binding site in the 3'-UTR of MMP-9 gene confers increased risk for atherosclerotic cerebral infarction in a Chinese population. Atherosclerosis 2013, 226, 447–452. [Google Scholar] [CrossRef]
- Navalgund, A.A.; Alifimoff, J.K.; Jakymec, A.J.; Bleyaert, A.L. Amiodarone-induced sinus arrest successfully treated with ephedrine and isoproterenol. Anesth Analg 1986, 65, 414–416. [Google Scholar]
- Jenh, C.H.; Rao, L.G.; Johnson, L.F. Regulation of thymidylate synthase enzyme synthesis in 5-fluorodeoxyuridine-resistant mouse fibroblasts during the transition from the resting to growing state. J Cell Physiol 1985, 122, 149–154. [Google Scholar]
- Ayusawa, D.; Shimizu, K.; Koyama, H.; Kaneda, S.; Takeishi, K.; Seno, T. Cell-cycle-directed regulation of thymidylate synthase messenger RNA in human diploid fibroblasts stimulated to proliferate. J Mol Biol 1986, 190, 559–567. [Google Scholar]
- Jenh, C.H.; Geyer, P.K.; Baskin, F.; Johnson, L.F. Thymidylate synthase gene amplification in fluorodeoxyuridine-resistant mouse cell lines. Mol Pharmacol 1985, 28, 80–85. [Google Scholar] [PubMed]
- Jenh, C.H.; Geyer, P.K.; Johnson, L.F. Control of thymidylate synthase mRNA content and gene transcription in an overproducing mouse cell line. Mol Cell Biol 1985, 5, 2527–2532. [Google Scholar] [PubMed]
- Ayusawa, D.; Koyama, H.; Shimizu, K.; Kaneda, S.; Takeishi, K.; Seno, T. Induction, by thymidylate stress, of genetic recombination as evidenced by deletion of a transferred genetic marker in mouse FM3A cells. Mol Cell Biol 1986, 6, 3463–3469. [Google Scholar] [PubMed]
- Linke, A.; Erbs, S.; Hambrecht, R. Effects of exercise training upon endothelial function in patients with cardiovascular disease. Front Biosci 2008, 13, 424–432. [Google Scholar] [PubMed]
- Erbs, S.; Linke, A.; Hambrecht, R. Effects of exercise training on mortality in patients with coronary heart disease. Coron Artery Dis 2006, 17, 219–225. [Google Scholar] [PubMed]
- Erbs, S.; Mobius-Winkler, S.; Linke, A.; Adams, V.; Doll, N.; Gielen, S.; Gummert, J.F.; Mohr, F.W.; Schuler, G.; Hambrecht, R. Both T-786C and G894T polymorphism of endothelial nitric oxide synthase affect in-vitro endothelium-dependent relaxation of internal mammary artery rings from patients with coronary artery disease. Eur J Cardiovasc Prev Rehabil 2006, 13, 826–831. [Google Scholar] [CrossRef]
- Park, D.W.; Yun, S.C.; Lee, S.W.; Kim, Y.H.; Lee, C.W.; Hong, M.K.; Kim, J.J.; Choo, S.J.; Song, H.; Chung, C.H.; Lee, J.W.; Park, S.W.; Park, S.J. Long-term mortality after percutaneous coronary intervention with drug-eluting stent implantation versus coronary artery bypass surgery for the treatment of multivessel coronary artery disease. Circulation 2008, 117, 2079–2086. [Google Scholar] [CrossRef]
- Jing, J.; Su, L.; Zeng, Y.; Tang, X.; Wei, J.; Wang, L.; Zhou, L. Variants in 9p21 Predicts Severity of Coronary Artery Disease in a Chinese Han Population. Ann Hum Genet 2016, 80, 274–281. [Google Scholar]
- Bae, J.; Kim, I.J.; Hong, S.H.; Sung, J.H.; Lim, S.W.; Cha, D.H.; Cho, Y.W.; Oh, D.; Kim, N.K. Association of endothelial nitric oxide synthase polymorphisms with coronary artery disease in Korean individuals with or without diabetes mellitus. Exp Ther Med 2010, 1, 719–724. [Google Scholar] [CrossRef]
- Park, H.S.; Sung, J.H.; Ryu, C.S.; Lee, J.Y.; Ko, E.J.; Kim, I.J.; Kim, N.K. The Synergistic Effect of Plasminogen Activator Inhibitor-1 (PAI-1) Polymorphisms and Metabolic Syndrome on Coronary Artery Disease in the Korean Population. J Pers Med 2020, 10. [Google Scholar] [CrossRef]
- Pullarkat, S.T.; Stoehlmacher, J.; Ghaderi, V.; Xiong, Y.P.; Ingles, S.A.; Sherrod, A.; Warren, R.; Tsao-Wei, D.; Groshen, S.; Lenz, H.J. Thymidylate synthase gene polymorphism determines response and toxicity of 5-FU chemotherapy. Pharmacogenomics J 2001, 1, 65–70. [Google Scholar]
- Villafranca, E.; Okruzhnov, Y.; Dominguez, M.A.; Garcia-Foncillas, J.; Azinovic, I.; Martinez, E.; Illarramendi, J.J.; Arias, F.; Martinez Monge, R.; Salgado, E.; Angeletti, S.; Brugarolas, A. Polymorphisms of the repeated sequences in the enhancer region of the thymidylate synthase gene promoter may predict downstaging after preoperative chemoradiation in rectal cancer. J Clin Oncol 2001, 19, 1779–1786. [Google Scholar]
- Ulrich, C.M.; Bigler, J.; Velicer, C.M.; Greene, E.A.; Farin, F.M.; Potter, J.D. Searching expressed sequence tag databases: discovery and confirmation of a common polymorphism in the thymidylate synthase gene. Cancer Epidemiol Biomarkers Prev 2000, 9, 1381–1385. [Google Scholar] [PubMed]
- Lenz, H.J.; Anderson, W.F.; Hall, F.L.; Gordon, E.M. Clinical protocol. Tumor site specific phase I evaluation of safety and efficacy of hepatic arterial infusion of a matrix-targeted retroviral vector bearing a dominant negative cyclin G1 construct as intervention for colorectal carcinoma metastatic to liver. Hum Gene Ther 2002, 13, 1515–1537. [Google Scholar] [PubMed]
- Jeon, Y.J.; Cho, S.H.; Kim, E.J.; Ryu, C.S.; Park, H.S.; Kim, J.W.; Lee, J.Y.; An, H.J.; Kim, N.K. 3'-UTR Polymorphisms in Thymidylate Synthase with Colorectal Cancer Prevalence and Prognosis. J Pers Med 2021, 11. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Jeong, C.K.; Hong, S.P.; Chong, S.Y.; Oh, D.; Hwang, S.G.; Ahn, D.H.; Kim, S.; Han, J.H.; Kim, N.K. [Clinical significance of thymidylate synthase and methylenetetrahydrofolate reductase gene polymorphism in Korean patients with gastric cancer]. Korean J Gastroenterol 2005, 46, 32–38. [Google Scholar]
- Yim, D.J.; Kim, O.J.; An, H.J.; Kang, H.; Ahn, D.H.; Hwang, S.G.; Oh, D.; Kim, N.K. Polymorphisms of thymidylate synthase gene 5'- and 3'-untranslated region and risk of gastric cancer in Koreans. Anticancer Res 2010, 30, 2325–2330. [Google Scholar]
- Kang, S.Y.; Lee, S.J.; Hong, S.H.; Chung, Y.K.; Oh, H.S.; Kim, S.W.; Yim, D.J.; Kim, N.K. Polymorphisms of 5,10-methylenetetrahydrofolate reductase and thymidylate synthase in squamous cell carcinoma and basal cell carcinoma of the skin. Mol Med Rep 2010, 3, 741–747. [Google Scholar]
| Characteristic | Controls | CAD patients | P |
|---|---|---|---|
| (n=427) | (n=424) | ||
| Male (n, %) | 153 (35.8) | 153 (36.1) | 0.375 |
| Age (years, mean ± SD) | 61.44±11.52 | 62.55±10.26 | 0.136 |
| Hypertension (n, %) | 171 (40.0) | 226 (57.4) | <0.0001 |
| Diabetes mellitus (n, %) | 51 (11.9) | 119 (30.2) | <0.0001 |
| Hyperlipidemia (n, %) | 97 (22.7) | 116 (29.4) | 0.028 |
| Smoking (n, %) | 136 (31.9) | 101 (25.6) | 0.049 |
| Body mass index (kg/cm2, mean ± SD) | 24.26±3.34 | 25.21±3.12 | <0.001 |
| Fasting blood sugar (mg/dL, mean ± SD) | 111.98±33.93 | 141.23±63.78 | <0.001 |
| HbA1c (%, mean ± SD) | 6.08±1.16 | 6.67±2.99 | 0.048 |
| HDL-C (mg/dl, mean ± SD) | 47.02±13.90 | 43.83±11.58 | 0.005 |
| LDL-C (mg/dl, mean ± SD) | 121.31±43.32 | 111.87±38.20 | 0.012 |
| Total cholesterol (mg/dl, mean ± SD) | 194.02±38.14 | 185.98±45.06 | 0.006 |
| Triglyceride (mg/dl, mean ± SD) | 142.16±86.73 | 155.52±97.04 | 0.037 |
| Hcy (μmol/L, mean ± SD) | 9.77±3.91 | 9.92±5.09 | 0.620 |
| Folate (nmol/L, mean ± SD) | 9.14±7.94 | 8.69±9.73 | 0.467 |
| Genotypes | Controls | CAD | AOR (95% CI)* | Pa | FDR-P |
|---|---|---|---|---|---|
| (n=427) | (n=424) | ||||
| TSER | |||||
| 3R3R | 296 (69.3) | 301 (71.0) | 1.000 (reference) | ||
| 2R3R | 123 (28.8) | 115 (27.1) | 0.934 (0.677-1.288) | 0.676 | 0.676 |
| 2R2R | 8 (1.9) | 8 (1.9) | 1.139 (0.401-3.236) | 0.807 | 0.807 |
| Additive model | 0.964 (0.725-1.281) | 0.800 | 0.800 | ||
| Dominant model | 0.945 (0.691-1.293) | 0.724 | 0.724 | ||
| Recessive model | 1.141 (0.398-3.271) | 0.806 | 0.826 | ||
| HWE-P | 0.240 | 0.431 | |||
| TS 1100T>C | |||||
| TT | 217 (50.8) | 194 (45.8) | 1.000 (reference) | ||
| TC | 177 (41.5) | 189 (44.6) | 1.333 (0.989-1.798) | 0.060 | 0.240 |
| CC | 33 (7.7) | 41 (9.7) | 1.344 (0.786-2.298) | 0.281 | 0.562 |
| Additive model | 1.249 (1.000-1.562) | 0.050 | 0.100 | ||
| Dominant model | 1.350 (1.014-1.797) | 0.040 | 0.158 | ||
| Recessive model | 1.230 (0.738-2.052) | 0.427 | 0.826 | ||
| HWE-P | 0.709 | 0.607 | |||
| TS 1170A>G | |||||
| AA | 211 (49.4) | 234 (55.2) | 1.000 (reference) | ||
| AG | 183 (42.9) | 172 (40.6) | 0.832 (0.618-1.121) | 0.227 | 0.454 |
| GG | 33 (7.7) | 18 (4.2) | 0.464 (0.242-0.889) | 0.021 | 0.084 |
| Additive model | 0.763 (0.602-0.966) | 0.025 | 0.100 | ||
| Dominant model | 0.773 (0.580-1.030) | 0.079 | 0.158 | ||
| Recessive model | 0.532 (0.288-0.983) | 0.044 | 0.176 | ||
| HWE-P | 0.439 | 0.084 | |||
| TS 1494ins/del | |||||
| 0bp0bp | 195 (45.7) | 211 (49.8) | 1.000 (reference) | ||
| 0bp6bp | 193 (45.2) | 174 (41.0) | 0.907 (0.673-1.223) | 0.522 | 0.676 |
| 6bp6bp | 39 (9.1) | 39 (9.2) | 0.878 (0.521-1.480) | 0.625 | 0.807 |
| Additive model | 0.934 (0.749-1.164) | 0.542 | 0.723 | ||
| Dominant model | 0.908 (0.683-1.208) | 0.508 | 0.677 | ||
| Recessive model | 0.946 (0.574-1.557) | 0.826 | 0.826 | ||
| HWE-P | 0.372 | 0.717 |
| Haplotype | Control (2n=854) | CAD (2n=848) | OR (95% CI) | Pa | FDR-P |
|---|---|---|---|---|---|
| TSER/TS1100/1170/1494 | |||||
| 3R-T-A-0bp | 0.3554 | 0.3916 | 1.000 (reference) | ||
| 3R-T-G-0bp | 0.2780 | 0.2133 | 0.699 (0.546-0.896) | 0.006 | 0.039 |
| 3R-C-A-0bp | 0.0024 | 0.0512 | 19.690 (4.727-81.990) | <0.0001 | 0.001 |
| TSER/TS1100/1170 | |||||
| 3R-T-A | 0.3781 | 0.4148 | 1.000 (Reference) | ||
| 3R-T-G | 0.2794 | 0.2132 | 0.695 (0.544 - 0.888) | 0.003 | 0.021 |
| 2R-T-A | 0.048 | 0.0307 | 0.582 (0.348 - 0.973) | 0.033 | 0.116 |
| TSER/TS1100/1494 | |||||
| 3R-T-0bp | 0.6336 | 0.6049 | 1.000 (Reference) | ||
| 3R-C-0bp | 0.0037 | 0.0550 | 16.522 (5.110 - 53.414) | <0.0001 | 0.001 |
| TSER/TS1170/1494 | |||||
| 3R-A-0bp | 0.3592 | 0.4487 | 1.000 (Reference) | ||
| 3R-A-6bp | 0.1965 | 0.1792 | 0.731 (0.560 - 0.954) | 0.021 | 0.053 |
| 3R-G-0bp | 0.2782 | 0.2109 | 0.608 (0.475 - 0.777) | <0.0001 | 0.001 |
| 2R-A-0bp | 0.0355 | 0.0231 | 0.539 (0.300 - 0.967) | 0.033 | 0.053 |
| 2R-A-6bp | 0.1173 | 0.1037 | 0.711 (0.514 - 0.983) | 0.038 | 0.053 |
| 2R-G-6bp | 0.0002 | 0.0075 | 10.51 (0.589 - 187.400) | 0.037 | 0.053 |
| TS1100/1170/1494 | |||||
| T-A-0bp | 0.392 | 0.4153 | 1.000 (Reference) | ||
| T-G-0bp | 0.287 | 0.2326 | 0.765 (0.602 - 0.973) | 0.028 | 0.098 |
| C-A-0bp | 0.0024 | 0.0509 | 20.462 (4.918 - 85.133) | <0.0001 | 0.001 |
| TSER/TS 1170 | |||||
| 3R-A | 0.5578 | 0.6302 | 1.000 (Reference) | ||
| 3R-G | 0.2794 | 0.2153 | 0.683 (0.543 - 0.858) | 0.001 | 0.003 |
| 2R-A | 0.1506 | 0.1245 | 0.732 (0.551 - 0.974) | 0.031 | 0.031 |
| 2R-G | 0.0122 | 0.0300 | 2.228 (1.059 - 4.688) | 0.017 | 0.026 |
| TS 1100/1170 | |||||
| T-A | 0.4259 | 0.4452 | 1.000 (Reference) | ||
| T-G | 0.2896 | 0.2353 | 0.780 (0.616 - 0.987) | 0.038 | 0.057 |
| C-G | 0.0020 | 0.0100 | 3.852 (0.813 - 18.261) | 0.023 | 0.057 |
| TS 1100/1494 | |||||
| T-0bp | 0.6790 | 0.6478 | 1.000 (Reference) | ||
| C-0bp | 0.0037 | 0.0550 | 16.551 (5.122 - 53.488) | <0.0001 | 0.0003 |
| TS 1170/1494 | |||||
| A-0bp | 0.3949 | 0.4715 | 1.000 (Reference) | ||
| A-6bp | 0.3135 | 0.2832 | 0.754 (0.602 - 0.946) | 0.015 | 0.015 |
| G-0bp | 0.2878 | 0.2313 | 0.671 (0.530 - 0.851) | 0.001 | 0.003 |
| G-6bp | 0.0038 | 0.0140 | 3.370 (0.943 - 12.042) | 0.014 | 0.015 |
| Genotype combinations | Controls | CAD | AOR (95% CI) | P* |
|---|---|---|---|---|
| (n=427) | (n=424) | |||
| TSER/TS 1170A>G | ||||
| 3R3R/AA | 130 (30.4) | 164 (38.7) | 1.000 (reference) | |
| 3R3R/AG | 136 (31.9) | 122 (28.8) | 0.656 (0.461 - 0.934) | 0.020 |
| 3R3R/GG | 30 (7.0) | 15 (3.5) | 0.335 (0.162 - 0.691) | 0.003 |
| 2R3R/AA | 74 (17.3) | 65 (15.3) | 0.634 (0.405 - 0.990) | 0.045 |
| TS 1100T>C/TS 1170A>G | ||||
| TT/AA | 70 (16.4) | 86 (20.3) | 1.000 (reference) | |
| TT/AG | 114 (26.7) | 92 (21.7) | 0.593 (0.376 - 0.936) | 0.025 |
| TT/GG | 33 (7.7) | 16 (3.8) | 0.330 (0.153 - 0.708) | 0.004 |
| TS 1100T>C/TS 1494ins>del | ||||
| TT/0bp0bp | 194 (45.4) | 180 (42.5) | 1.000 (reference) | |
| TC/0bp0bp | 1 (0.2) | 21 (5.0) | 26.713 (3.462 - 206.115) | 0.002 |
| TS 1170A>G/TS 1494ins>del | ||||
| AA/0bp0bp | 57 (13.3) | 101 (23.8) | 1.000 (reference) | |
| AA/0bp6bp | 117 (27.4) | 97 (22.9) | 0.479 (0.303 - 0.755) | 0.002 |
| AA/6bp6bp | 37 (8.7) | 36 (8.5) | 0.454 (0.241 - 0.857) | 0.015 |
| AG/0bp0bp | 105 (24.6) | 94 (22.2) | 0.463 (0.292 - 0.734) | 0.001 |
| AG/0bp6bp | 76 (17.8) | 75 (17.7) | 0.539 (0.331 - 0.876) | 0.013 |
| GG/0bp0bp | 33 (7.7) | 16 (3.8) | 0.213 (0.098 - 0.462) | 0.0001 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





