Submitted:
12 July 2023
Posted:
13 July 2023
You are already at the latest version
Abstract
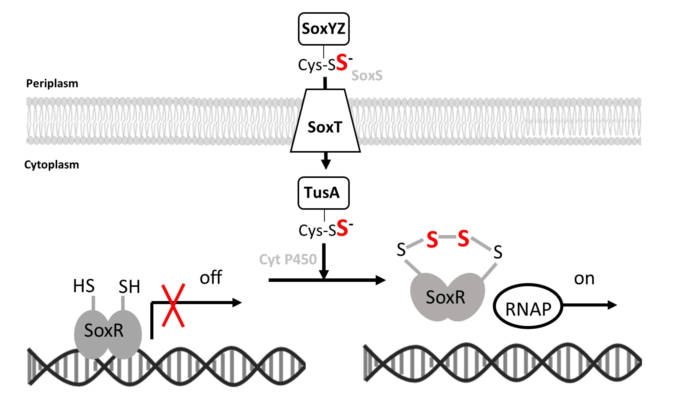
Keywords:
1. Introduction
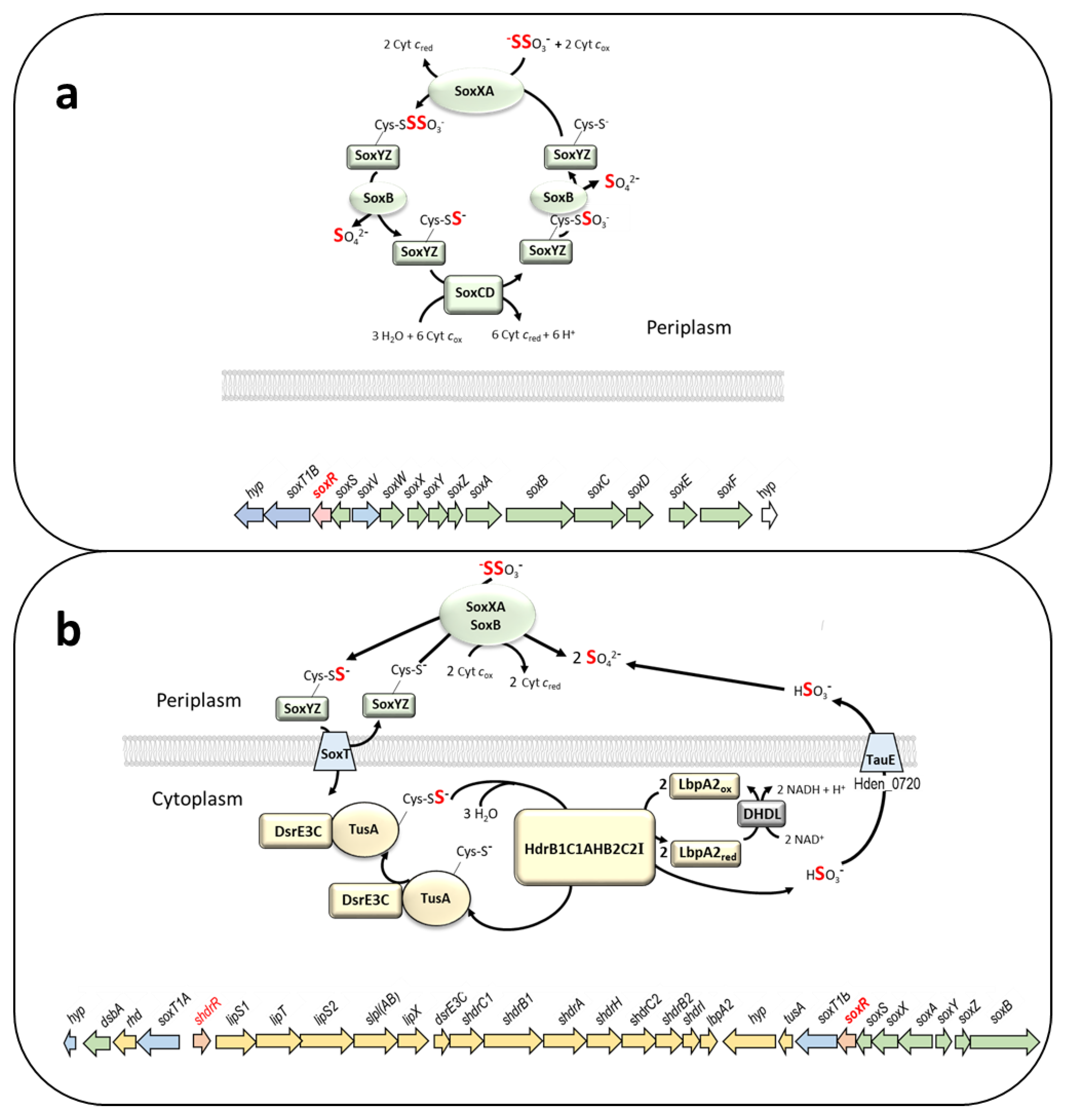
2. Materials and Methods
2.1. Bacterial strains, plasmids, primers, and growth conditions
2.2. Recombinant DNA techniques
2.3. Construction of plasmid for deletion of soxR in H. denitrificans
2.4. Characterization of phenotypes, quantification of sulfur compounds and protein content
2.5. RNA preparation
2.6. Expression studies based on RT-qPCR
2.7. Cloning, site-directed mutagenesis, overproduction, and purification of recombinant SoxR proteins
2.8. Electrophoretic mobility shift assays (EMSA)
2.9. Gel permeation chromatography
2.10. Preparation of polysulfides
2.11. Redox treatments, persulfuration reactions, MalPEG gel-shift assays and mass spectrometry
2.12. Distribution of Sox systems and SoxR: dataset generation and anaylsis
3. Results
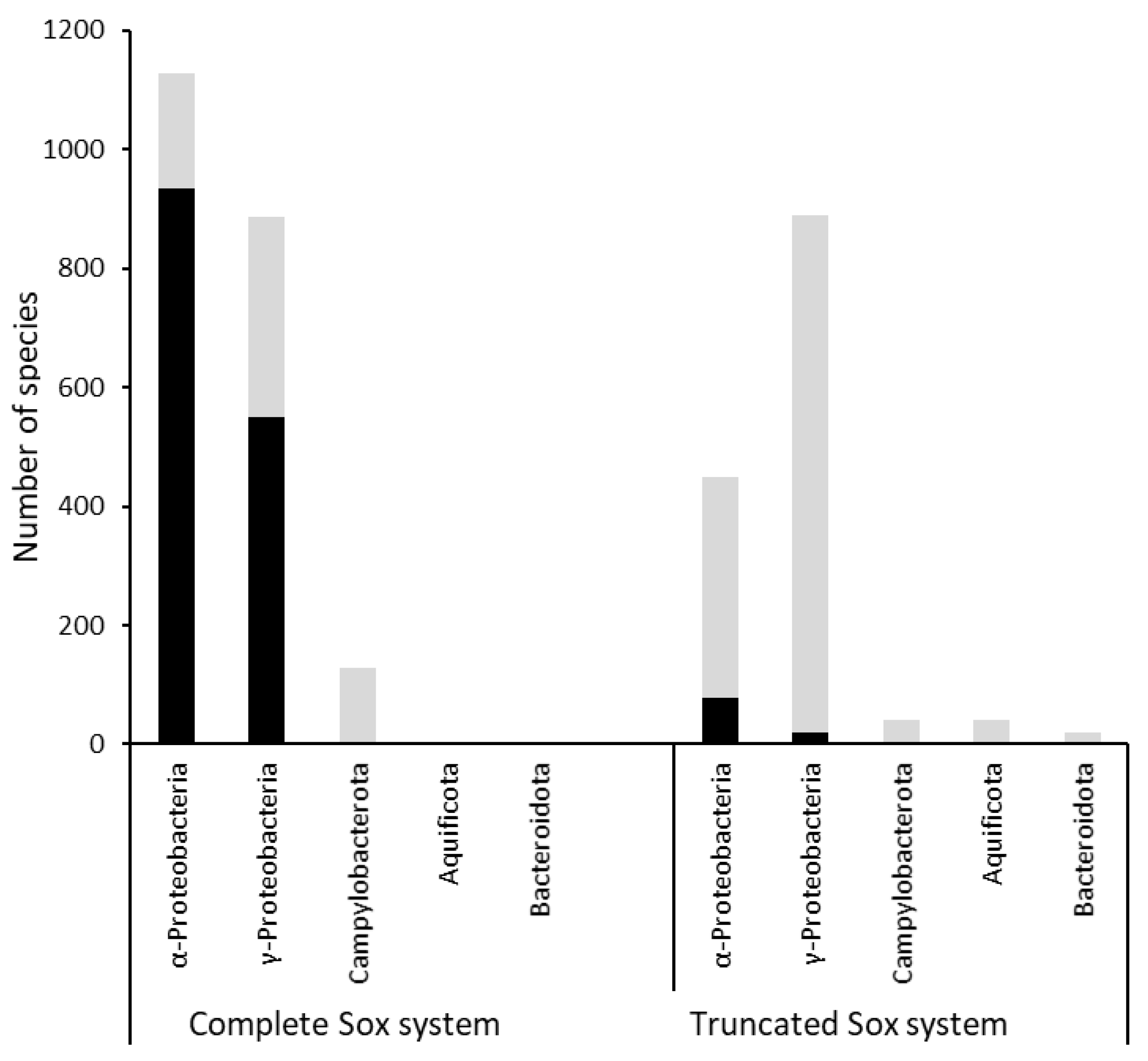
3.1. Distribution of Sox systems and the SoxR regulator
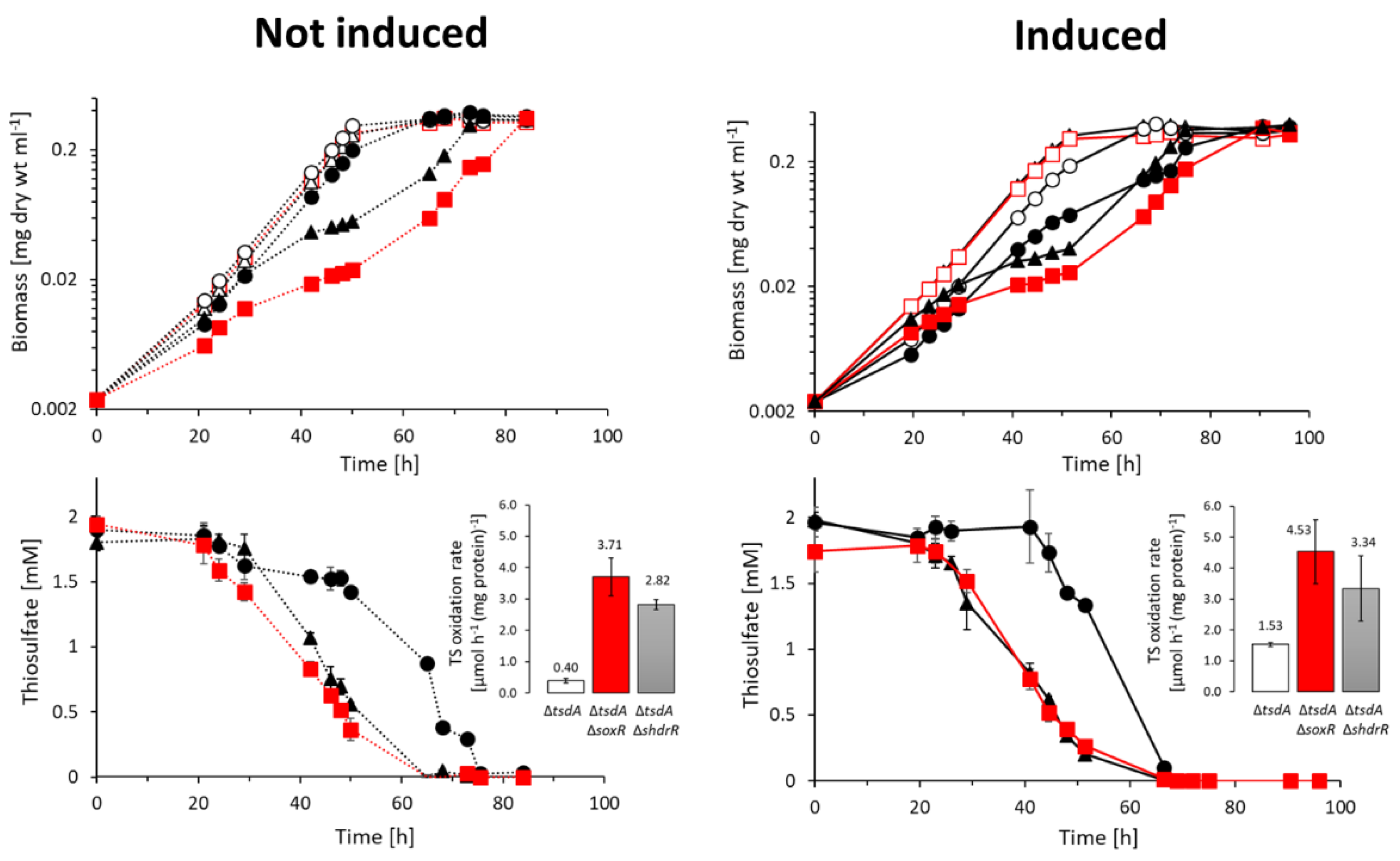
3.2. Genetic evidence for SoxR function in H. denitrificans
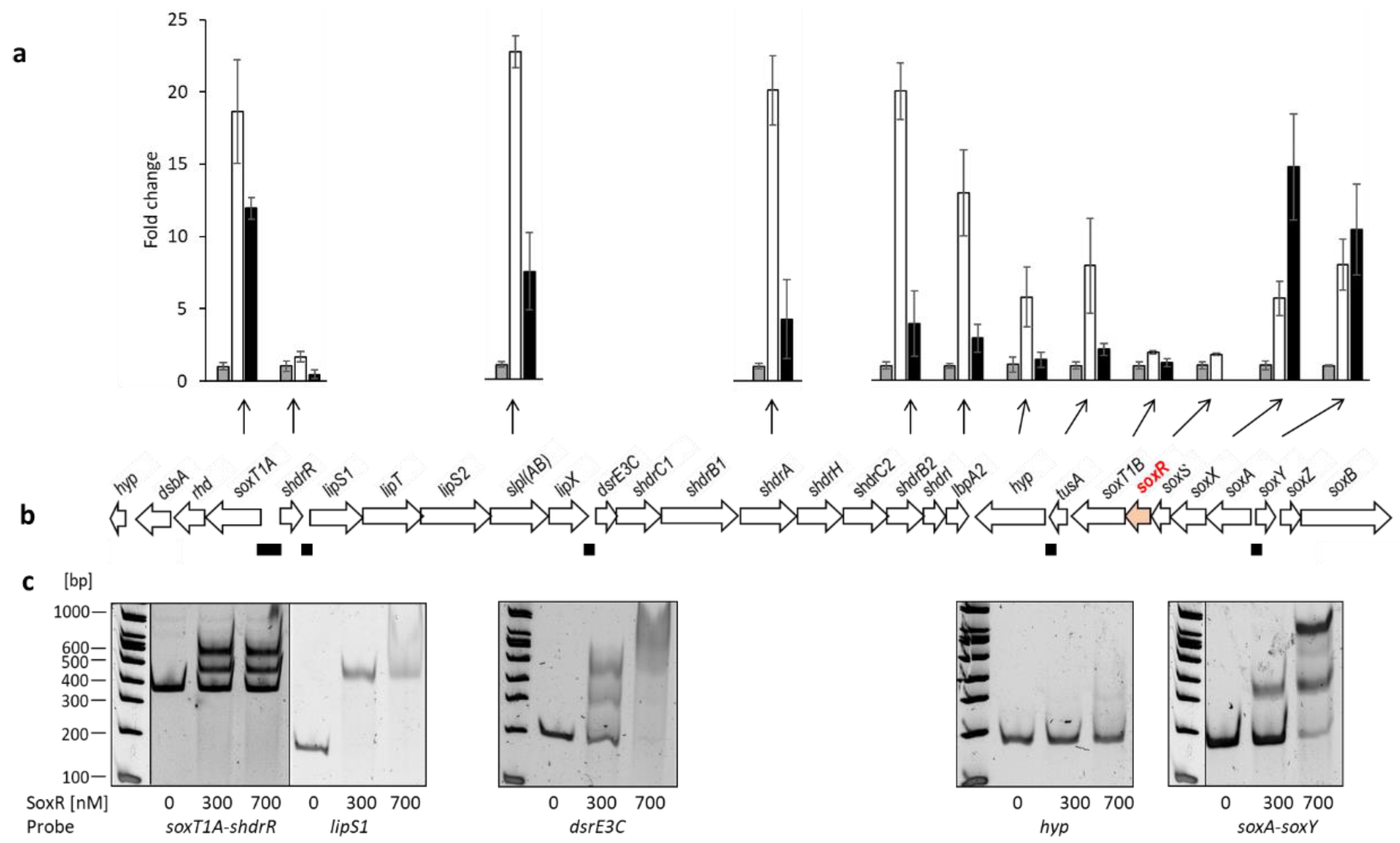
3.3. Identification of genes controlled by SoxR by RT-qPCR for different H. denitrificans strains
3.3. Identification of SoxR target sites by EMSA
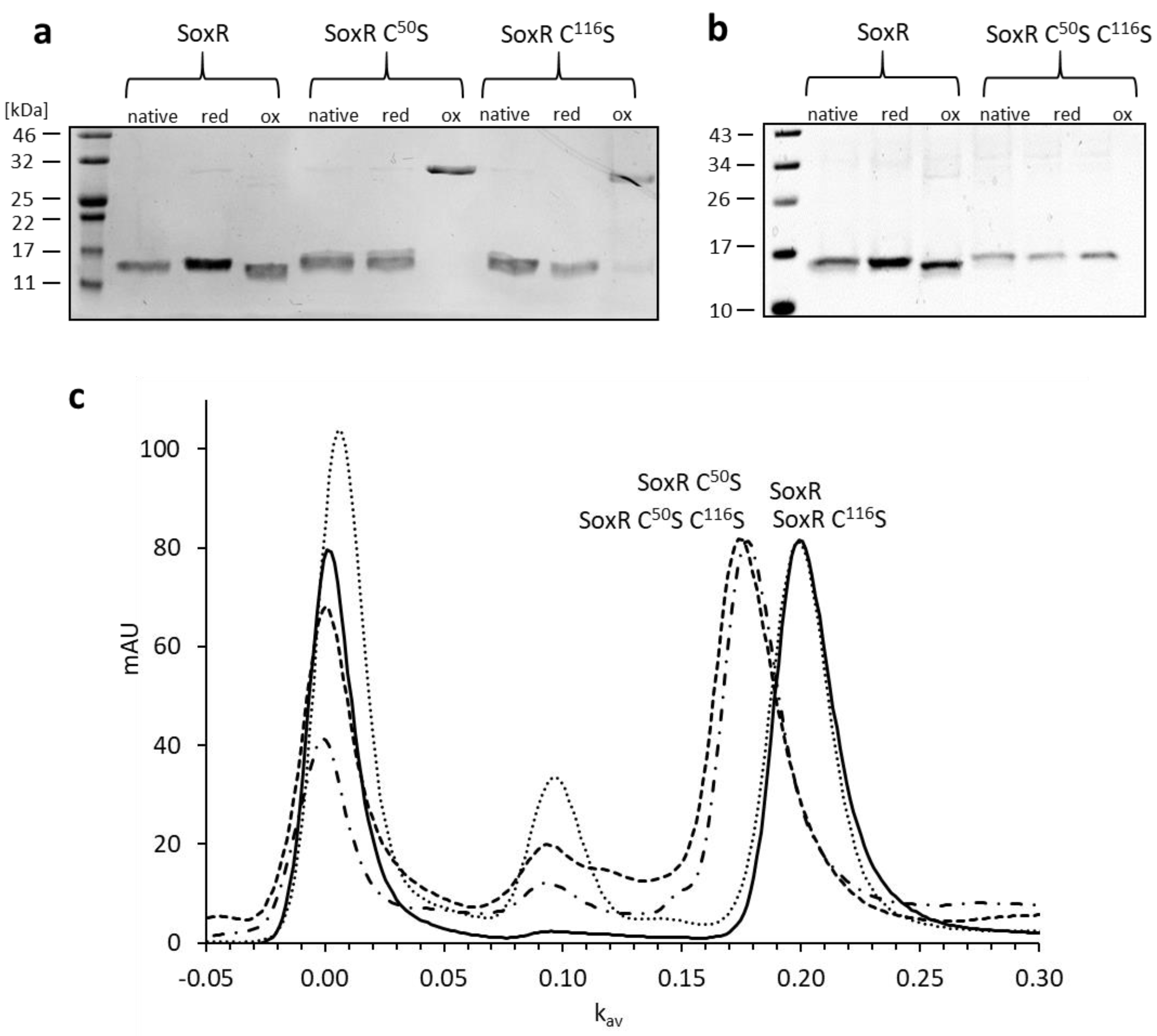
3.4. Properties of the SoxR protein
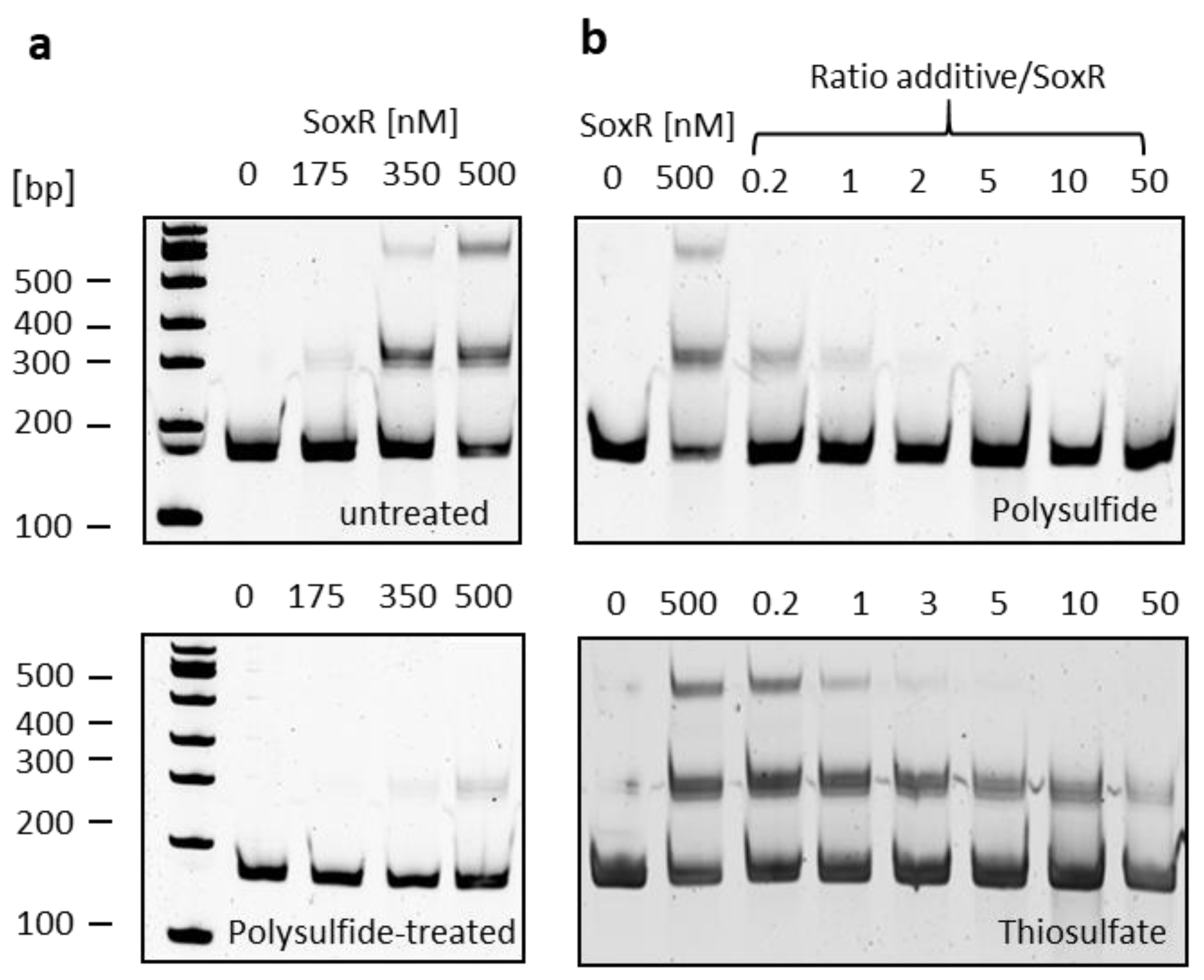
3.5. SoxR binding properties
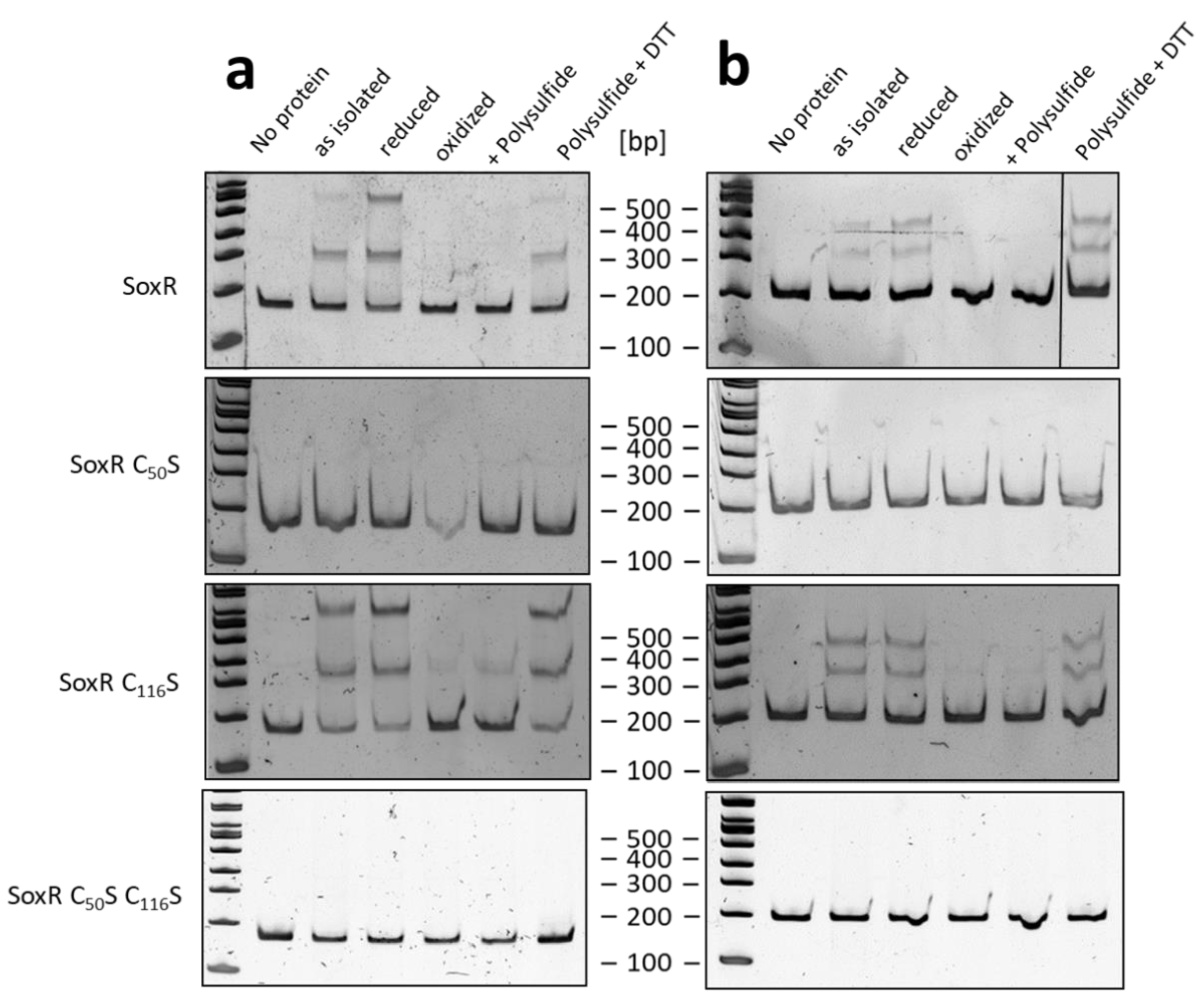
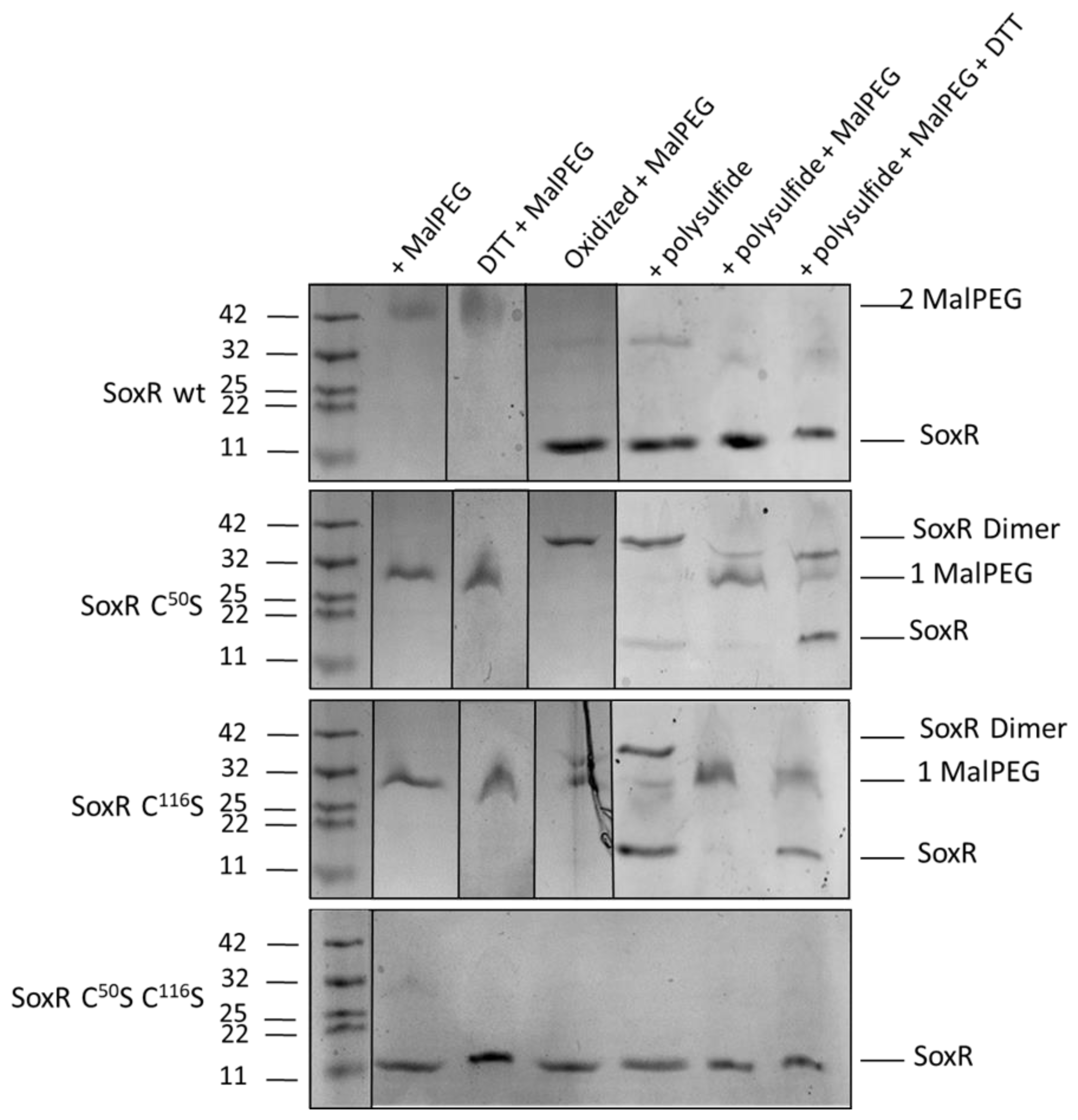
3.6. Redox state and modification of SoxR
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Friedrich, C.G.; Rother, D.; Bardischewsky, F.; Quentmeier, A.; Fischer, J. Oxidation of reduced inorganic sulfur compounds by bacteria: emergence of a common mechanism? Appl. Environ. Microbiol. 2001, 67, 2873–2882. [Google Scholar] [CrossRef] [PubMed]
- Dahl, C. A biochemical view on the biological sulfur cycle. In Environmental technologies to treat sulfur pollution: principles and engineering, 2 ed; Lens, P. (ed.), IWA Publishing: London, 2020; pp. 55–96. [Google Scholar]
- Appia-Ayme, C.; Little, P.J.; Matsumoto, Y.; Leech, A.P.; Berks, B.C. Cytochrome complex essential for photosynthetic oxidation of both thiosulfate and sulfide in Rhodovulum sulfidophilum. J. Bacteriol. 2001, 183, 6107–6118. [Google Scholar] [CrossRef] [PubMed]
- Grabarczyk, D.B.; Berks, B.C. Intermediates in the Sox sulfur oxidation pathway are bound to a sulfane conjugate of the carrier protein SoxYZ. PLoS One 2017, 12, e0173395. [Google Scholar] [CrossRef]
- Sauvé, V.; Roversi, P.; Leath, K.J.; Garman, E.F.; Antrobus, R.; Lea, S.M.; Berks, B.C. Mechanism for the hydrolysis of a sulfur-sulfur bond based on the crystal structure of the thiosulfohydrolase SoxB. J. Biol. Chem. 2009, 284, 21707–21718. [Google Scholar] [CrossRef] [PubMed]
- Grabarczyk, D.B.; Chappell, P.E.; Johnson, S.; Stelzl, L.S.; Lea, S.M.; Berks, B.C. Structural basis for specificity and promiscuity in a carrier protein/enzyme system from the sulfur cycle. Proc. Natl. Acad. Sci. USA 2015, 112, E7166–E7175. [Google Scholar] [CrossRef]
- Zander, U.; Faust, A.; Klink, B.U.; de Sanctis, D.; Panjikar, S.; Quentmeier, A.; Bardischewsky, F.; Friedrich, C.G.; Scheidig, A.J. Structural basis for the oxidation of protein-bound sulfur by the sulfur cycle molybdohemo-enzyme sulfane dehydrogenase SoxCD. J. Biol. Chem. 2010; 286, 8349. [Google Scholar]
- Dahl, C.; Friedrich, C.G.; Kletzin, A. Sulfur oxidation in prokaryotes. In Encyclopedia of Life Sciences, John Wiley & Sons, Ltd.: Chichester, 2008.
- Koch, T.; Dahl, C. A novel bacterial sulfur oxidation pathway provides a new link between the cycles of organic and inorganic sulfur compounds. ISME J. 2018, 12, 2479–2491. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Koch, J.; Flegler, W.; Garcia Ruiz, L.; Hager, N.; Ballas, A.; Tanabe, T.S.; Dahl, C. A metabolic puzzle: consumption of C1 compounds and thiosulfate in Hyphomicrobium denitrificans XT. Biochim. Biophys. Acta - Bioenergetics 2022, 1864, 148932. [Google Scholar] [CrossRef] [PubMed]
- Cao, X.; Koch, T.; Steffens, L.; Finkensieper, J.; Zigann, R.; Cronan, J.E.; Dahl, C. Lipoate-binding proteins and specific lipoate-protein ligases in microbial sulfur oxidation reveal an atpyical role for an old cofactor. eLife 2018, 7, e37439. [Google Scholar] [CrossRef]
- Appia-Ayme, C.; Berks, B.C. SoxV, an orthologue of the CcdA disulfide transporter, is involved in thiosulfate oxidation in Rhodovulum sulfidophilum and reduces the periplasmic thioredoxin SoxW. Biochem. Biophys. Res. Commun. 2002, 296, 737–741. [Google Scholar] [CrossRef]
- Friedrich, C.G.; Quentmeier, A.; Bardischewsky, F.; Rother, D.; Orawski, G.; Hellwig, P.; Fischer, J. Redox control of chemotrophic sulfur oxidation of Paracoccus pantotrophus. In Microbial sulfur metabolism; Dahl, C., Ed.; Friedrich, C. G. (eds.), Springer: Berlin Heidelberg, 2008; pp. 139–150. [Google Scholar]
- Orawski, G.; Bardischewsky, F.; Quentmeier, A.; Rother, D.; Friedrich, C.G. The periplasmic thioredoxin SoxS plays a key role in activation in vivo of chemotrophic sulfur oxidation of Paracoccus pantotrophus. Microbiology 2007, 153, 1081–1086. [Google Scholar] [CrossRef]
- Carius, Y.; Rother, D.; Friedrich, C.G.; Scheidig, A.J. The structure of the periplasmic thiol-disulfide oxidoreductase SoxS from Paracoccus pantotrophus indicates a triple Trx/Grx/DsbC functionality in chemotrophic sulfur oxidation. Acta Crystallogr. D Biol. Cryst. 2009, 65, 229–240. [Google Scholar] [CrossRef] [PubMed]
- Rother, D.; Ringk, J.; Friedrich, C.G. Sulfur oxidation of Paracoccus pantotrophus: The sulfur-binding protein SoxYZ is the target of the periplasmic thiol-disulfide oxidoreductase SoxS. Microbiology 2008, 154, 1980–1988. [Google Scholar] [CrossRef] [PubMed]
- Neutzling, O.; Pfleiderer, C.; Trüper, H.G. Dissimilatory sulphur metabolism in phototrophic “non-sulphur” bacteria. J. Gen. Microbiol. 1985, 131, 791–798. [Google Scholar] [CrossRef]
- Kappler, U.; Enemark, J.H. Sulfite-oxidizing enzymes. J. Biol. Inorg. Chem. 2014, 20, 253–264. [Google Scholar] [CrossRef]
- Kappler, U.; Schwarz, G. The sulfite oxidase family of molybdenum enzymes. In Molybdenum and tungsten enzymes; Hille, R., Schulzke, C., Eds.; Kirk, M. L. (eds.), Royal Society of Chemistry: Cambridge, 2016; pp. 240–273. [Google Scholar]
- Meng, Z.; Qin, G.; Zhang, B.; Bai, J. DNA damaging effects of sulfur dioxide derivatives in cells from various organs of mice. Mutagenesis 2004, 19, 465–468. [Google Scholar] [CrossRef]
- Yi, H.; Liu, J.; Zheng, K. Effect of sulfur dioxide hydrates on cell cycle, sister chromatid exchange, and micronuclei in barley. Ecotoxicol. Environ. Saf. 2005, 62, 421–426. [Google Scholar] [CrossRef]
- Ozturk, O.H.; Kucukatay, V.; Yonden, Z.; Agar, A.; Bagci, H.; Delibas, N. Expressions of N-methyl-D-aspartate receptors NR2A and NR2B subunit proteins in normal and sulfite-oxidase deficient rat’s hippocampus: effect of exogenous sulfite ingestion. Arch. Toxicol. 2006, 80, 671–679. [Google Scholar] [CrossRef]
- Kao, Y.T.; Tan, C.; Song, S.H.; Ozturk, N.; Li, J.; Wang, L.; Sancar, A.; Zhong, D. Ultrafast dynamics and anionic active states of the flavin cofactor in cryptochrome and photolyase. J. Am. Chem. Soc. 2008, 130, 7695–7701. [Google Scholar] [CrossRef]
- Ozawa, T.; Hanaki, A. Spin-trapping of sulfite radical anion, SO3-., by a water-soluble, nitroso-aromatic spin-trap. Biochem. Biophys. Res. Commun. 1987, 142, 410–416. [Google Scholar] [CrossRef]
- Inouye, B.; Morita, K.; Ishida, T.; Ogata, M. Cooperative effect of sulfite and vanadium compounds on lipid peroxidation. Toxicol. Appl. Pharmacol. 1980, 53, 101–107. [Google Scholar] [CrossRef]
- Yang, S.F. Destruction of tryptophan during the aerobic oxidation of sulfite ions. Environmental Research 1973, 6, 395–402. [Google Scholar] [CrossRef] [PubMed]
- Rother, D.; Orawski, G.; Bardischewsky, F.; Friedrich, C.G. SoxRS-mediated regulation of chemotrophic sulfur oxidation in Paracoccus pantotrophus. Microbiology 2005, 151, 1707–1716. [Google Scholar] [CrossRef] [PubMed]
- Mandal, S.; Chatterjee, S.; Dam, B.; Roy, P.; Das Gupta, S.K. The dimeric repressor SoxR binds cooperatively to the promoter(s) regulating expression of the sulfur oxidation (sox) operon of Pseudaminobacter salicylatoxidans KCT001. Microbiology 2007, 153, 80–91. [Google Scholar] [CrossRef] [PubMed]
- Lahiri, C.; Mandal, S.; Ghosh, W.; Dam, B.; Roy, P. A novel gene cluster soxSRT is essential for the chemolithotrophic oxidation of thiosulfate and tetrathionate by Pseudaminobacter salicylatoxidans KCT001. Curr. Microbiol. 2006, 52, 267–273. [Google Scholar] [CrossRef]
- Ma, Z.; Jacobsen, F.E.; Giedroc, D.P. Coordination chemistry of bacterial metal transport and sensing. Chem Rev 2009, 109, 4644–4681. [Google Scholar] [CrossRef]
- Cook, W.J.; Kar, S.R.; Taylor, K.B.; Hall, L.M. Crystal structure of the cyanobacterial metallothionein repressor SmtB: a model for metalloregulatory proteins. J. Mol. Biol. 1998, 275, 337–346. [Google Scholar] [CrossRef]
- Capdevila, Daiana A. ; Edmonds, Katherine A.; Giedroc, David P. Metallochaperones and metalloregulation in bacteria. Essays Biochem. 2017, 61, 177–200. [Google Scholar] [CrossRef]
- Guimaraes, B.G.; Barbosa, R.L.; Soprano, A.S.; Campos, B.M.; de Souza, T.A.; Tonoli, C.C.; Leme, A.F.; Murakami, M.T.; Benedetti, C.E. Plant pathogenic bacteria utilize biofilm growth-associated repressor (BigR), a novel winged-helix redox switch, to control hydrogen sulfide detoxification under hypoxia. J. Biol. Chem. 2011, 286, 26148–26157. [Google Scholar] [CrossRef]
- Shimizu, T.; Shen, J.; Fang, M.; Zhang, Y.; Hori, K.; Trinidad, J.C.; Bauer, C.E.; Giedroc, D.P.; Masuda, S. Sulfide-responsive transcriptional repressor SqrR functions as a master regulator of sulfide-dependent photosynthesis. Proc. Natl. Acad. Sci. USA 2017, 114, 2355–2360. [Google Scholar] [CrossRef]
- Bertani, G. Lysogeny at mid-twentieth century: P1, P2, and other experimental systems. J. Bacteriol. 2004, 186, 595–600. [Google Scholar] [CrossRef]
- Ausubel, F.A.; Brent, R.; Kingston, R.E.; Moore, D.D.; Seidman, J.G.; Smith, J.A.; Struhl, K. Current protocols in molecular biology. John Wiley & Sons, New York, 1997.
- Horton, R.M. PCR mediated recombination and mutagenesis: SOEing together tailor-made genes. Mol. Biotechnol. 1995, 3, 93–99. [Google Scholar] [CrossRef]
- Dahl, C. Insertional gene inactivation in a phototrophic sulphur bacterium: APS-reductase-deficient mutants of Chromatium vinosum. Microbiology 1996, 142, 3363–3372. [Google Scholar] [CrossRef]
- Chomczynski, P.; Sacchi, N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 1987, 162, 156–159. [Google Scholar] [CrossRef]
- Martineau, C.; Mauffrey, F.; Villemur, R. Comparative analysis of denitrifying activities of Hyphomicrobium nitrativorans, Hyphomicrobium denitrificans, and Hyphomicrobium zavarzinii. Appl. Environ. Microbiol. 2015, 81, 5003–5014. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-D D CT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, S.; Satake, H.; Hisano, T.; Terazawa, T. Potentiometric argentimetric method for the successive titration of sulphide and dissolved sulphur in polysulphide solutions. Talanta 1972, 19, 1650–1654. [Google Scholar] [CrossRef]
- Parks, D.H.; Chuvochina, M.; Waite, D.W.; Rinke, C.; Skarshewski, A.; Chaumeil, P.-A.; Hugenholtz, P. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat. Biotechnol. 2018, 36, 996–1004. [Google Scholar] [CrossRef] [PubMed]
- Rinke, C.; Chuvochina, M.; Mussig, A.J.; Chaumeil, P.A.; Davin, A.A.; Waite, D.W.; Whitman, W.B.; Parks, D.H.; Hugenholtz, P. A standardized archaeal taxonomy for the Genome Taxonomy Database. Nat. Microbiol. 2021, 6, 946–959. [Google Scholar] [CrossRef] [PubMed]
- Hyatt, D.; Chen, G.L.; Locascio, P.F.; Land, M.L.; Larimer, F.W.; Hauser, L.J. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 2010, 11, 119. [Google Scholar] [CrossRef]
- Tanabe, T.S.; Dahl, C. HMS-S-S: a tool for the identification of sulphur metabolism-related genes and analysis of operon structures in genome and metagenome assemblies. Mol. Ecol. Resour. 2022, 22, 2758–2774. [Google Scholar] [CrossRef]
- Dahl, C. Sulfur metabolism in phototrophic bacteria. In Modern topics in the phototrophic prokaryotes: Metabolism, bioenergetics and omics; Hallenbeck, P. C. (ed.), Springer International Publishing: Cham, 2017; pp. 27–66. [Google Scholar]
- Frigaard, N.U.; Dahl, C. Sulfur metabolism in phototrophic sulfur bacteria. Adv. Microb. Physiol. 2009, 54, 103–200. [Google Scholar]
- Gupta, R.S. The phylum Aquificae. In The prokaryotes - other lineages of Bacteria and the Archaea; Rosenberg, E., DeLong, E. F., Lory, S., Stackebrandt, E., Thompson, F., Eds.; Springer: Berlin Heidelberg, 2014. [Google Scholar]
- Kodama, Y.; Watanabe, K. Sulfuricurvum kujiense gen. nov., sp. nov., a facultatively anaerobic, chemolithoautotrophic, sulfur-oxidizing bacterium isolated from an underground crude-oil storage cavity. Int. J. Syst. Evol. Microbiol. 2004, 54, 2297–2300. [Google Scholar] [CrossRef]
- Xie, S.; Wang, S.; Li, D.; Shao, Z.; Lai, Q.; Wang, Y.; Wei, M.; Han, X.; Jiang, L. Sulfurovum indicum sp. nov., a novel hydrogen- and sulfur-oxidizing chemolithoautotroph isolated from a deep-sea hydrothermal plume in the Northwestern Indian Ocean. Int. J. Syst. Evol. Microbiol. 2019, 71. [Google Scholar] [CrossRef] [PubMed]
- Labrenz, M.; Grote, J.; Mammitzsch, K.; Boschker, H.T.S.; Laue, M.; Jost, G.; Glaubitz, S.; Jurgens, K. Sulfurimonas gotlandica sp. nov., a chemoautotrophic and psychrotolerant epsilonproteobacterium isolated from a pelagic redoxcline, and an emended description of the genus Sulfurimonas. Int. J. Syst. Evol. Microbiol. 2013, 63, 4141–4148. [Google Scholar] [CrossRef]
- Skirnisdottir, S.; Hreggvidsson, G.O.; Holst, O.; Kristjansson, J.K. Isolation and characterization of a mixotrophic sulfur-oxidizing Thermus scotoductus. Extremophiles 2001, 5, 45–51. [Google Scholar] [CrossRef] [PubMed]
- Bjornsdottir, S.H.; Petursdottir, S.K.; Hreggvidsson, G.O.; Skirnisdottir, S.; Hjorleifsdottir, S.; Arnfinnsson, J.; Kristjansson, J.K. Thermus islandicus sp. nov., a mixotrophic sulfur-oxidizing bacterium isolated from the Torfajokull geothermal area. Int. J. Syst. Evol. Microbiol. 2009, 59, 2962–2966. [Google Scholar] [CrossRef] [PubMed]
- Imhoff, J.F. Phylogenetic taxonomy of the family Chlorobiaceae on the basis of 16S rRNA and fmo (Fenna-Matthews-Olson protein) gene sequences. Int. J. Syst. Evol. Microbiol. 2003, 53, 941–951. [Google Scholar] [CrossRef]
- Carareto Alves, L.M.; de Souza, J.A.M.; Varani, A.d.M.; Lemos, E.G.d.M. The Family Rhizobiaceae. In The prokaryotes; Rosenberg, E., DeLong, E. F., Lory, S., Stackebrandt, E., Eds.; Thompson, F. (eds.), Springer: Berlin Heidelberg, 2014; pp. 419–437. [Google Scholar]
- Pujalte, M.J.; Lucena, T.; Ruvira, M.A.; Arahal, D.R.; Macián, M.C. The Family Rhodobacteraceae. In The prokaryotes; Rosenberg, E., DeLong, E. F., Lory, S., Stackebrandt, E., Eds.; Thompson, F. (eds.), Springer: Berlin, Heidelberg, 2014; pp. 439–512. [Google Scholar]
- Spietz, R.L.; Lundeen, R.A.; Zhao, X.; Nicastro, D.; Ingalls, A.E.; Morris, R.M. Heterotrophic carbon metabolism and energy acquisition in Candidatus Thioglobus singularis strain PS1, a member of the SUP05 clade of marine Gammaproteobacteria. Environ. Microbiol. 2019, 21, 2391–2401. [Google Scholar] [CrossRef]
- Imhoff, J.F. Family I. Chromatiaceae Bavendamm 1924, 125 AL emend. Imhoff 1984b, 339. In Bergey’s manual of systematic bacteriology; Brenner, D. J., Krieg, N. R., Staley, J. T., Eds.; Garrity, G. M. (eds.), Springer: New York, 2005. [Google Scholar]
- Imhoff, J.F. Family II. Ectothiorhodospiraceae Imhoff 1984b, 339 VP In Bergey’s manual of systematic bacteriology; Brenner, D. J., Krieg, N. R., Staley, J. T., Garrity, G. M., Eds.; Springer: New York, 2005; Vol. 2,pp. 41-57. [Google Scholar]
- Boden, R.; Scott, K.M.; Williams, J.; Russel, S.; Antonen, K.; Rae, A.W.; Hutt, L.P. An evaluation of Thiomicrospira, Hydrogenovibrio and Thioalkalimicrobium: reclassification of four species of Thiomicrospira to each Thiomicrorhabdus gen. nov. and Hydrogenovibrio, and reclassification of all four species of Thioalkalimicrobium to Thiomicrospira. Int. J. Syst. Evol. Microbiol. 2017, 67, 1140–1151. [Google Scholar]
- Boden, R.; Scott, K.M. Evaluation of the genus Thiothrix Winogradsky 1888 (Approved Lists 1980) emend. Aruga et al. 2002: reclassification of Thiothrix disciformis to Thiolinea disciformis gen. nov., comb. nov., and of Thiothrix flexilis to Thiofilum flexile gen. nov., comb nov., with emended description of Thiothrix. Int. J. Syst. Evol. Microbiol. 2018, 68, 2226–2239. [Google Scholar]
- Shimizu, T.; Masuda, S. Characterization of redox-active cysteine residues of persulfide-responsive transcriptional repressor SqrR. Commun. Integr. Biol. 2017, 10, e1329786. [Google Scholar] [CrossRef]
- Mukherjee, D.; Datta, A.B.; Chakrabarti, P. Crystal structure of HlyU, the hemolysin gene transcription activator, from Vibrio cholerae N16961 and functional implications. Biochim. Biophys. Acta 2014, 1844, 2346–2354. [Google Scholar] [CrossRef] [PubMed]
- Gueune, H.; Durand, M.J.; Thouand, G.; DuBow, M.S. The ygaVP genes of Escherichia coli form a tributyltin-inducible operon. Appl. Environ. Microbiol. 2008, 74, 1954–1958. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Deutsch, C. Pegylation: a method for assessing topological accessibilities in Kv1.3. Biochemistry 2001, 40, 13288–13301. [Google Scholar] [CrossRef] [PubMed]
- Capdevila, D.A.; Walsh, B.J.C.; Zhang, Y.; Dietrich, C.; Gonzalez-Gutierrez, G.; Giedroc, D.P. Structural basis for persulfide-sensing specificity in a transcriptional regulator. Nat. Chem. Biol. 2021, 17, 65–70. [Google Scholar] [CrossRef]
- Barbosa, R.L.; Benedetti, C.E. BigR, a transcriptional repressor from plant-associated bacteria, regulates an operon implicated in biofilm growth. J. Bacteriol. 2007, 189, 6185–6194. [Google Scholar] [CrossRef]
- Grossoehme, N.; Kehl-Fie, T.E.; Ma, Z.; Adams, K.W.; Cowart, D.M.; Scott, R.A.; Skaar, E.P.; Giedroc, D.P. Control of copper resistance and inorganic sulfur metabolism by paralogous regulators in Staphylococcus aureus. J. Biol. Chem. 2011, 286, 13522–13531. [Google Scholar] [CrossRef]
- Balasubramanian, R.; Hori, K.; Shimizu, T.; Kasamatsu, S.; Okamura, K.; Tanaka, K.; Ihara, H.; Masuda, S. The sulfide-responsive SqrR/BigR homologous regulator YgaV of Escherichia coli controls expression of anaerobic respiratory genes and antibiotic tolerance. Antioxidants (Basel) 2022, 11. [Google Scholar] [CrossRef]
- Tang, C.; Li, J.; Shen, Y.; Liu, M.; Liu, H.; Liu, H.; Xun, L.; Xia, Y. A sulfide-sensor and a sulfane sulfur-sensor collectively regulate sulfur-oxidation for feather degradation by Bacillus licheniformis. Commun. Biol. 2023, 6, 167. [Google Scholar] [CrossRef]
- Tanaka, Y.; Yoshikaie, K.; Takeuchi, A.; Ichikawa, M.; Mori, T.; Uchino, S.; Sugano, Y.; Hakoshima, T.; Takagi, H.; Nonaka, G.; Tsukazaki, T. Crystal structure of a YeeE/YedE family protein engaged in thiosulfate uptake. Science Advances 2020, 6, eaba7637. [Google Scholar] [CrossRef]
- Tanabe, T.S.; Leimkühler, S.; Dahl, C. The functional diversity of the prokaryotic sulfur carrier protein TusA. Adv. Microb. Physiol. 2019, 75, 233–277. [Google Scholar] [PubMed]


| Treatment | SoxR Mass [Da] (addition: [Da]) |
SoxR C50S Mass [Da] (addition: [Da]) |
SoxR C116S Mass [Da] (addition: [Da]) |
SoxR C50S C116S Mass [Da] (addition: [Da]) |
|---|---|---|---|---|
| Native | 15212.8 | 15197.3 | 15198.2 | 15182.3 |
| DTT reduced | 15212.5 | 15199.3 | 15198.8 | nd |
| CuCl2 oxidized | 15210.5 | 15196.7 | 15196.9 | 15182.0 |
| Iodoacetamide | 15328.0 (2 CAM: 2 × 57.07) |
15255.2 (1 CAM: 57.07) |
15255.22 (1 CAM: 57.07) |
nd |
| Polysulfide | 15212.5 15244.7 (1 S: 32) 15276.4 (2 S: 64) 15308.0 (3 S: 96) |
15198.9 15230.0 (1 S: 32) |
15198.0 15230.3 (1 S: 32) 15261.1 (2 S: 64) |
15182.0 |
| Polysulfide + Iodoacetamide | 15212.9 15244.3 (1 S: 32) 15275.2 (2 S: 64) 15306.0 (3 S: 96) |
15198.0 15286.0 (1 S + 1 CAM: 90) |
15197.6 15228.5 (1 S: 32) 15285.4 (1 S + 1 CAM: 90) |
15180.5 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





