1. Introduction
With advances in industrial technology, many convenient chemical substances have been developed for daily use. Moreover, as these chemical substances have become common in our daily lives, their risks have increased. Therefore, it is important to identify and analyze the environmental factors that affect human health and disease. The concept proposed for this purpose is the exposome, and its importance has been emphasized [
1,
2].
Exposome is defined as the totality of various external factors that an individual is exposed to throughout their lifetime, and includes biological responses related to various factors such as environmental and lifestyle factors [
3]. Exposome is a concept that contrasts with genome [
4], and consists of three domains [
5]: general external domain, such as social status, surrounding environment, and climate; specific external domain, such as chemical pollutants and pathogens; and biological internal domain, such as metabolism, oxidative stress, and aging. Methods for studying exposomes include cohort studies, environment-wide association studies, and omics [
4,
6,
7].
Bisphenol A (BPA), Bisphenol F (BPF), and S (BPS) are chemical substances known as bisphenols. Bisphenols were used to prepare the polycarbonate and epoxy resins. Polycarbonate is transparent and hard and is used as a consumer product for water bottles, sports equipment, etc. BPA is a widely used organic synthetic compound with two hydroxyl phenol groups and the chemical formula (CH
3)
2C(C
6H
4OH)
2. However, BPA can enter the body through various routes, such as inhalation and skin contact, and shows various toxic effects, including atopic diseases [
8]. Therefore, BPF and BPS were used as alternatives to BPA [
9]. BPF is an organic synthetic compound with two hydroxyl phenol groups and the chemical formula (CH
3)
2C(C
6H
3(OH)
2)
2. BPS is an organic synthetic compound with two sulfone phenol groups and chemical formula (CH
3)
2C(C
6H
4SO
2)
2. However, research on the human exposure levels and health effects of BPF and BPS remains insufficient.
Environmental pollutants such as bisphenols are harmful chemicals that affect human health and can enter the body through air, soil, and food contamination. Exposure to these environmental pollutants can cause abnormalities in the immune system and inflammation in organs, such as the skin, respiratory system, and digestive system, which can lead to environmental diseases. Environmental diseases include asthma, atopy, allergy, chronic obstructive pulmonary disease (COPD), chronic sinusitis, etc., and they are diseases that many people suffer from worldwide. AD is a common chronic inflammatory skin disease affecting more than 20% of children and 10% of adults worldwide [
10]. AD occurs owing to defects in skin barrier function, immune regulation disorders, and complex interactions between environmental and infectious factors [
11]. AD is characterized by recurrent itching and localized eczema, and many patients have allergies, such as allergic asthma, allergic rhinitis, food allergy, etc. [
12].
DNA methylation is a phenomenon in which a methyl group is attached to the cytosine base of a DNA molecule. It mainly occurs in the region where cytosine and guanine are connected, and is called a CpG dinucleotide. DNA methylation is involved in important biological processes such as gene expression regulation, chromosome stability, X-chromosome inactivation, and gene imprinting, and affects various life phenomena such as cell differentiation, development, and aging [
13]. Unlike genetic mutations, DNA methylation does not change the DNA sequence itself, but can be reversibly altered by environmental factors inside and outside the cell [
14]. The reversibility of DNA methylation results in cell adaptability [
15]. DNA methylation has a specific pattern that only appears in certain tissues or cells, and can be used to identify the identity and state of cells [
16]. DNA methylation is associated with various diseases. Especially in patients with AD, methylation of skin barrier-related genes such as filaggrin and immune response-related genes such as cytokines decreases or increases, causing damage and inflammation of the skin barrier [
17,
18]. Therefore, DNA methylation is receiving considerable attention as a biomarker for diagnosis, prognosis, treatment response, and other diseases.
In this study, we aimed to observe the correlation between BPS and BPF, which are used as alternatives to BPA, and AD from the perspective of exposome. To do this, we used the GREEN (Growing children’s health and Evaluation of Environment) cohort, which was recruited for the study of environmental diseases caused by environmental pollutants, to assess the exposure levels of bisphenols in adult women and to analyze the correlation between methylation changes and atopic diseases.
2. Results
2.1. Information of participants for the analysis of Differentially Methylated Regions (DMR)
We divided the participants into atopic and non-atopic groups and compared their information by group (
Table 1). Of 117 participants, 19 had atopy. Most participants were in their 30s, with an average age of approximately 33.9 years, and most were non-smokers. The average exposure levels of BPA, BPF, and BPS for all participants were 1.56 ug/g cr., 0.78 ug/g cr., and 0.30 ug/g cr., respectively, which were slightly higher than the average exposure levels of BPA, BPF, and BPS for adults according to the results of the National Environmental Health Survey in Korea, which were 1.08 ug/g cr., 0.19 ug/g cr., and 0.19 ug/g cr.
We separated the case and control groups for the DMR analysis and checked the exposure levels of each substance by group (
Table 2). The BPA group consisted of three atopic patients with high BPA exposure and an average age of 38.7 years. The control group consisted of 72 non-atopic participants with low exposure and an average age of 33.9 years. The mean BPA exposure level was 2.5 ug/g cr. in the case group and 0.6 ug/g cr. in the control group, which was lower than that in the case group. In the BPF exposure group, the case group consisted of six atopic patients with an average age of 32.5 years and an average exposure level of 1.1 ug/g cr., whereas the control group consisted of 75 non-atopic participants with an average age of 33.9 years and an average exposure level of 0.1 ug/g cr. Finally, in the BPS exposure group, the case group consisted of five atopic patients with an average age of 33.8 years and an average exposure level of 1.2 u/g cr., whereas the control group consisted of 74 non-atopic participants with an average age of 33.5 years and an average exposure level of 0.1 ug/g cr.
2.2. DMR analysis according to bisphenol exposure
DMR analysis was performed by comparing the case and control groups grouped by each substance and setting the window size to 100 bp. We analyzed with a cutoff of |FC| > 2, p-value > 0.05, and summarized the DMR analysis results in a table (
Table 3). The BPA exposure group included 85,305 DMRs, of which 90 were hypermethylated and 85,215 were hypomethylated. The BPF and BPS exposure groups had 97,560 and 68,982 DMRs, respectively, of which 178 and 3,450 were hypermethylated, and 97,382 and 65,532 were hypomethylated, respectively.
2.3. Bisphenol and AD
We matched the regions summarized using DMR analysis with the genes, summarized the methylation levels of the genes, and then compared them with the AD-related genes organized by the database (
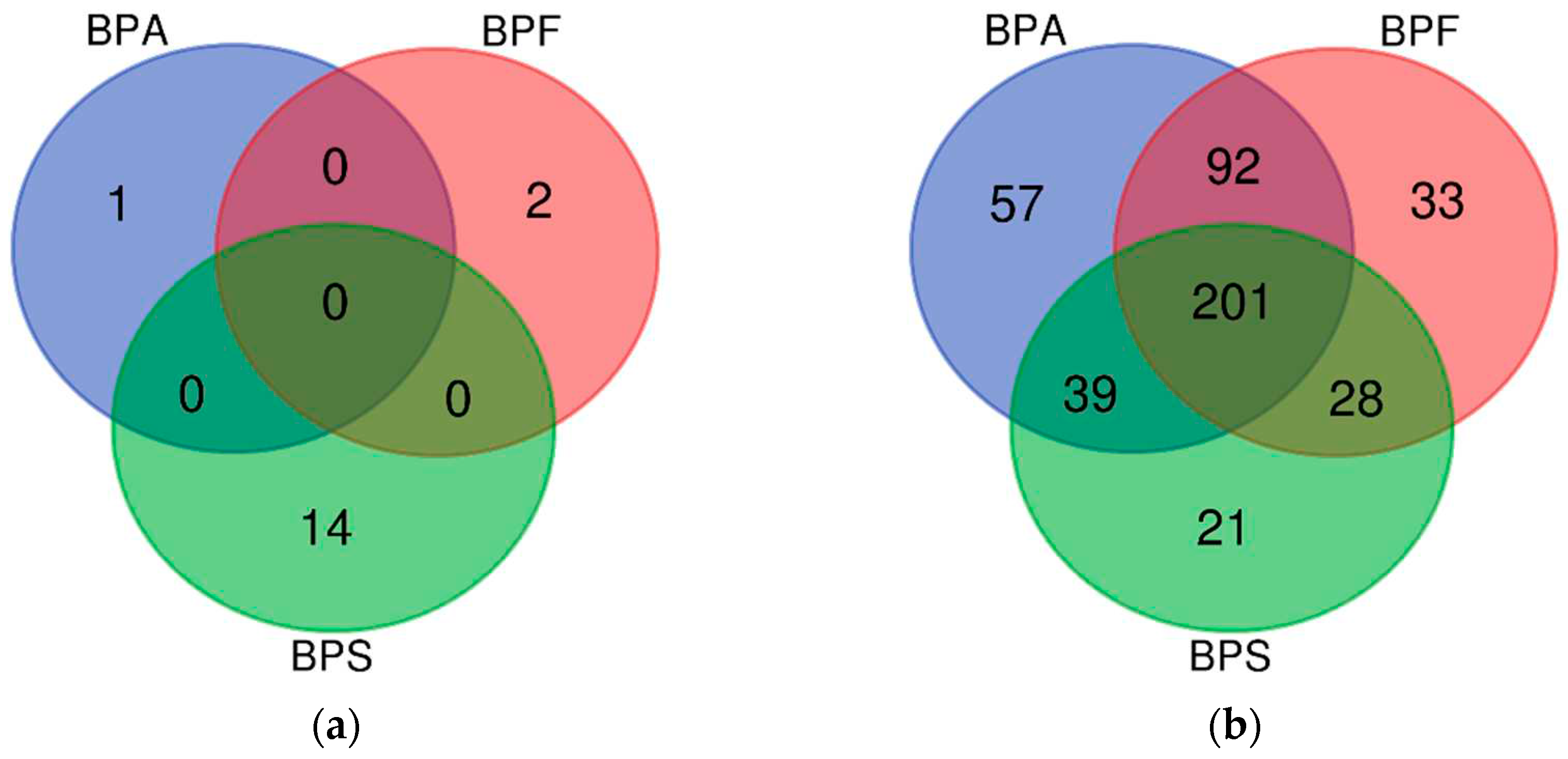
Table 4). The number of AD-related genes was 390, 356, and 303 in the BPA, BPF, and BPS exposure groups, respectively. The number of hypermethylated genes was one, two, and 14 in the BPA, BPF, and BPS exposure groups. The number of hypomethylated genes was 389, 354, and 289 in the BPA, BPF, and BPS exposure groups.
The number of genes derived from DMR analysis is shown in a Venn diagram (
Figure 1). Among the hypermethylated genes, no common genes were expressed after BPA, BPF, and BPS exposure (
Figure 1a). Most of the hypomethylated genes were included in all BPA, BPF, and BPS exposure groups. Among them, IL4 (BPA logFC −1.76, BPF logFC −1.75, BPS logFC −1.62), one of the cytokines known to be associated with atopy [
19], was included, and IL13RA1 (BPA logFC −2.67, BPF logFC −1.60, BPS logFC −2.08), which is known to be expressed in keratinocytes [
20], was also commonly hypo methylated in all three exposure groups (
Figure 1b).
We verified the association between 201 AD-related genes derived from the three bisphenol exposure results and atopy. Among the common pathways, we confirmed that the JAK-STAT and PI3K-AKT signaling pathways are both influenced by IL4 and are related to AD [
21,
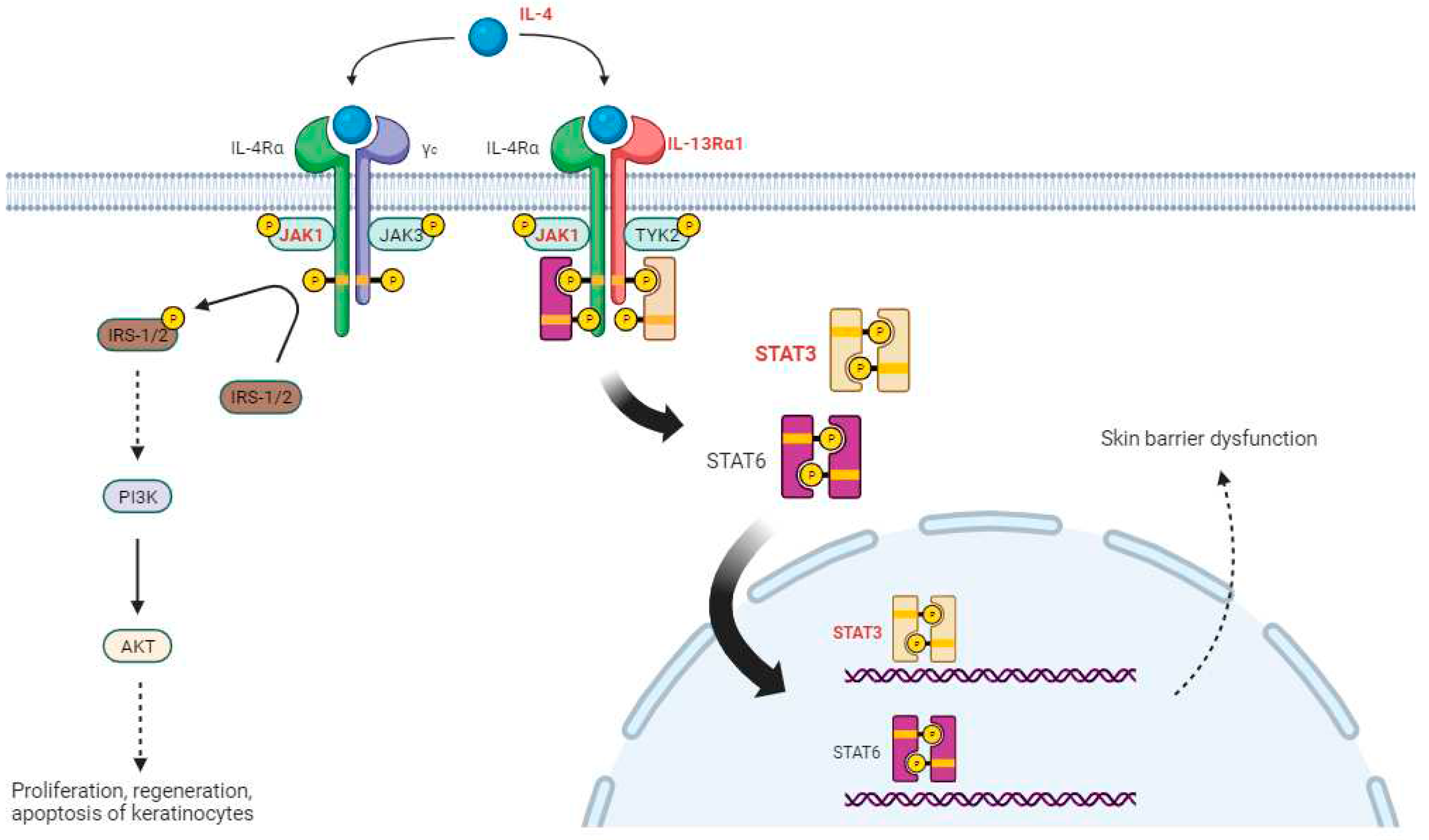
22]. From the DMR analysis results, we identified various genes that were commonly hypomethylated in the BPA, BPF, and BPS exposure groups, including IL4, JAK1 (BPA logFC −2.77, BPF logFC −2.13, BPS logFC −1.90), IL13RA1, and STAT3 (BPA logFC −2.41, BPF logFC −1.91, BPS logFC −1.50), which belong to the PI3K-AKT and JAK-STAT signaling pathways (marked in red in
Figure 2). BCL2 (BPA logFC −2.59, BPF logFC −1.09, and BPS logFC −1.65), which is involved in the PI3K-AKT signaling pathway, was also found to be common in the three bisphenol exposure groups and was overexpressed in patients with AD [
23].
In addition, various genes involved in the PI3K-AKT and JAK-STAT signaling pathways showed changes in methylation (
Table 5). IL33, another cytokine gene associated with AD [
24] besides IL4, was hypomethylated in both BPA and BPF exposure groups, as well as in TYK2, which phosphorylates and activates STAT3 in the JAK-STAT pathway. JAK3, which is phosphorylated along with JAK1 in type 1 IL4-R, was also hypomethylated in both BPA and BPS exposure groups.
3. Discussion
AD is an inflammatory skin disease with increasing prevalence worldwide [
25]. The cause of AD is a complex interaction between genetic and environmental factors. One of the suggested environmental factors is BPA exposure. BPA is a chemical substance widely used in plastic products and food containers [
26], which can affect the immune system and skin barrier function in the body [
27]. Several studies have shown that populations exposed to BPA have a higher risk of atopy. As concerns regarding the safety of BPA increased, BPF and BPS were used as alternatives. However, research on BPF and BPS is relatively scarce compared to that on BPA, especially from an epigenetic perspective. Epigenetics studies the various mechanisms that regulate gene expression without changing the gene sequence, which can be altered by environmental factors that affect the onset and progression of diseases.
This study observed changes in DNA methylation in the PBMC of pregnant women exposed to BPA, BPF, and BPS. Of the 117 pregnant women, 19 had AD. The body exposure level of bisphenols was the highest for BPA, and the average exposure level of all participants was slightly higher than the results of the National Environmental Health Survey. This difference may be due to the fact that the National Environmental Health Survey surveyed all adults, while this study only targeted women in their 30s who were in their reproductive age.
For the DMR analysis, we separated the case group of AD patients with high exposure and the control group of non-AD patients with low exposure. We confirmed that IL4, a cytokine known to be associated with atopy; IL13RA1, known to be expressed in keratinocytes; and BCL2, known to be overexpressed in patients with AD, were commonly hypomethylated in all BPA, BPF, and BPS exposure groups. In addition, we confirmed that IL33, TYK2, JAK1, and JAK3 are hypomethylated. This suggests that these genes were relatively overexpressed, and we confirmed that these genes belong to the JAK-STAT and PI3-AKT signaling pathways.
There are two types of IL-4 receptors in the JAK-STAT and PI3K-Akt signaling pathways. When IL-4Rα binds to γc, it becomes IL-4R type 1, and when it binds to IL-13Rα1, it becomes IL-4R type 2 [
28]. Type 1 IL-4R binds to IL-4 and phosphorylates insulin receptor substances (IRS) through JAK1 phosphorylation [
29]. This induces the activation of AKT signaling, including that of PI3K, and affects the growth, survival, and proliferation of keratinocytes [
22]. Type 2 IL-4R binds to IL-4 and phosphorylates JAK1 and TYK2, which, in turn, phosphorylate and dimerize STAT6 and STAT3, respectively. STAT6 and STAT3 move to the nucleus and affect skin barrier dysfunction, which is known to be involved in the production and regulation of Th2 cytokines, major factors in the development of AD. Bisphenols can worsen the symptoms of AD by reducing the methylation of these genes and increasing their expression.
This study shows that both BPA and its substitutes, BPF and BPS, can affect AD through epigenetic mechanisms. However, this study had some limitations. First, the cohort used in this study only targeted pregnant women; therefore, the body exposure level to bisphenols may differ from that of adults in general. Second, this study only used DMR analysis; therefore, other factors involved in gene expression regulation were not considered. Recent studies have used multi-omics instead of single-omics [
30,
31]. Therefore, future studies should elucidate the complex gene expression network by integrating DMR analysis with other omics data [
32]. This integrated analysis reflects the actual situation of human exposure to bisphenols, identifies the biochemical changes that occur in the body, and reveals the causal relationship and risk factors of disease onset.
4. Materials and Methods
4.1. Human-derived samples
This study conducted experiments on pregnant women from the GREEN (Growing Children’s Health and Evaluation of Environment) cohort (IRB #2016-12-111) recruited between 2017 and 2021. The GREEN cohort consisted of 151 pregnant women and 99 infants, and blood and urine samples were collected from pregnant women. PBMCs were collected from the blood, and gDNA was obtained to perform MeDIP-seq; urine samples were analyzed for urinary concentrations of BPA, BPF, and BPS using liquid chromatography mass spectrometry (LC/MS/MS) by Smartive Co., Ltd. (Hanam City, Gyeonggi-do, Republic of Korea) and then corrected for creatinine.
4.2. Classification of exposure groups for BPA, BPF, and BPS
To divide into control and case groups according to the difference in exposure levels of BPA, BPF, and BPS, the upper 25% of exposure level was set as the high-exposure group and the lower as the low-exposure group (BPA:1.685 ug/g creatinine, BPF:0.445 ug/g creatinine, BPS:0.178 ug/g creatinine). For each substance, those who had AD and were in the high-exposure group were designated as the case group, and those who did not have AD and were in the low-exposure group were designated as the control group.
4.3. MeDIP-seq (DNA preparation, library construction, sequencing)
Blood was collected using heparin tubes, and PBMCs were separated from the collected blood. gDNA was extracted from the PBMCs using an Invitrogen PureLink Genomic DNA Mini Kit (ThermoFisher, Waltham, MA, USA). The purity of the extracted gDNA was measured using a Nanodrop (ThermoFisher) and quantified using a Qubit (ThermoFisher). DNA fragmentation was performed using a Bioruptor (Diagenode, Seraing, Belgium) and immunoprecipitation was performed using an antibody. An adapter-attached library was constructed using the adaptor index included in the TruSeq RNA UD Indices (Illumina, San Diego, CA, USA). PCR was performed to amplify DNA with an adaptor index attached, and only 300–350 bp size were selected using Pippin Prep (Sage Science, Beverly, MA, USA). The constructed library was sequenced with 2*75bp read length of paired-end using Nextseq500 (Illumina).
4.4. Differentially Methylated Regions (DMR) analysis
We performed a quality check on the raw data using FastQC (Ver. 1.11.9) and trimmed the raw data using Trimmomatic (Ver. 0.40) program. We used TruSeq3-PE.fa, which was provided by Trimmomatic for the adapter sequence during the trimming process. We created an index for the hg38 reference genome using bowtie2 (Ver. 2.4.2) and mapped them. We converted the SAM format derived later into the BAM format using SAMtools (Ver. 1.11.0), and sorted the files based on the mapped position information. Duplicate sequences were removed from the sorted files using Picard (Ver. 2.25.0). The DMR analysis was performed using MEDIPS (ver. 1.42.0), a R (Ver. 4.3.1) package (p-value < 0.05, |FC| > 2). We analyzed only autosomes and the X chromosome because we performed the analysis for adult women, divided the window size into 100 bp and proceeded and integrated the continuous position. We set DMR annotation as ‘GENE’ and marked it as gene containing the position.
4.5. Functional Analysis
We obtained a list of 842 genes related to atopy from the Comparative Toxicogenomics Database (CTD) and DisGeNET DB. We also compared the genes derived from DMR analysis with AD-related genes and analyzed the functions and cellular pathways of the derived genes using the KEGG PATHWAY Database of the Database for Annotation, Visualization and Integrated Discovery (DAVID) tool.
4.6. Ethical Considerations
All patients provided informed written consent for participation in the study, as regulated by the local law. This study was approved by the Institutional Review Board (IRB #2016-12-111) of Samsung Seoul Hospital, and written consent was obtained from the parents who participated in this study.
5. Conclusions
In this study, we used DMR analysis in pregnant women to investigate the correlation between bisphenols (BPA, BPF, and BPS) and AD. The results showed hypomethylation of various genes known to affect AD in BPA, BPF, and BPS and confirmed that these genes were involved in the JAK-STAT and PI3K-AKT signaling pathways. This study confirmed that BPA, BPF, and BPS can negatively affect atopy.
Author Contributions
Methodology, validation, formal analysis, data curation, writing—original draft preparation, visualization, S.H.K.; formal analysis, investigation, S. Y. Y. and J.H.C.; data curation, investigation, J.K. and K.A.; and supervision, editing, project administration, funding, S.Y.H. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korean Government (MSIT) [No. 2022R1A2C1092120]. This study was supported by the Korea Environment Industry & Technology Institute(KEITI) through "the Environmental Health Action Program" funded by the Korea Ministry of Environment(MOE) (2017001360005).
Institutional Review Board Statement
This study was approved by the Institutional Review Board (IRB #2016-12-111) of Samsung Seoul Hospital, and written consent was obtained from the parents who participated in this study.
Informed Consent Statement
Informed consent was obtained from all participants involved in the study.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Wild, C.P. The exposome: from concept to utility. International journal of epidemiology 2012, 41, 24–32. [Google Scholar] [CrossRef]
- Gaylord, A.; Barrett, E.S.; Sathyanarayana, S.; Swan, S.H.; Nguyen, R.H.; Bush, N.R.; Carroll, K.; Day, D.B.; Kannan, K.; Trasande, L. Prenatal bisphenol A and S exposure and atopic disease phenotypes at age 6. Environmental Research 2023, 226, 115630. [Google Scholar] [CrossRef]
- Vermeulen, R.; Schymanski, E.L.; Barabási, A.-L.; Miller, G.W. The exposome and health: Where chemistry meets biology. Science 2020, 367, 392–396. [Google Scholar] [CrossRef]
- Haddad, N.; Andrianou, X.D.; Makris, K.C. A scoping review on the characteristics of human exposome studies. Current Pollution Reports 2019, 5, 378–393. [Google Scholar] [CrossRef]
- Miller, G.W.; Jones, D.P. The nature of nurture: refining the definition of the exposome. Toxicological sciences 2014, 137, 1–2. [Google Scholar] [CrossRef]
- Stefanovic, N.; Irvine, A.D.; Flohr, C. The role of the environment and exposome in atopic dermatitis. Current Treatment Options in Allergy 2021, 8, 222–241. [Google Scholar] [CrossRef]
- Koh, E.J.; Kim, S.H.; Hwang, S.Y. Sample management: a primary critical starting point for successful omics studies. Molecular & Cellular Toxicology 2022, 18, 141–148. [Google Scholar] [CrossRef]
- Stefanovic, N.; Flohr, C.; Irvine, A.D. The exposome in atopic dermatitis. Allergy 2020, 75, 63–74. [Google Scholar] [CrossRef]
- Rochester, J.R.; Bolden, A.L. Bisphenol S and F: a systematic review and comparison of the hormonal activity of bisphenol A substitutes. Environmental health perspectives 2015, 123, 643–650. [Google Scholar] [CrossRef]
- Malajian, D.; Guttman-Yassky, E. New pathogenic and therapeutic paradigms in atopic dermatitis. Cytokine 2015, 73, 311–318. [Google Scholar] [CrossRef]
- Solomon, I.; Ilie, M.A.; Draghici, C.; Voiculescu, V.M.; Căruntu, C.; Boda, D.; Zurac, S. The impact of lifestyle factors on evolution of atopic dermatitis: An alternative approach. Experimental and therapeutic medicine 2019, 17, 1078–1084. [Google Scholar] [CrossRef]
- Ghazal, S.; Ridha, Z.; D'Aguanno, K.; Nassim, D.; Quaiattini, A.; Netchiporouk, E.; Poulin, Y.; Kalia, S.; Marcoux, D.; Piguet, V. Treatment guidelines for atopic dermatitis since the approval of dupilumab: a systematic review and quality appraisal using AGREE-II. Frontiers in Medicine 2022, 253. [Google Scholar] [CrossRef]
- Villicaña, S.; Bell, J.T. Genetic impacts on DNA methylation: research findings and future perspectives. Genome biology 2021, 22, 127. [Google Scholar] [CrossRef]
- Ruiz-Hernandez, A.; Kuo, C.-C.; Rentero-Garrido, P.; Tang, W.-Y.; Redon, J.; Ordovas, J.M.; Navas-Acien, A.; Tellez-Plaza, M. Environmental chemicals and DNA methylation in adults: a systematic review of the epidemiologic evidence. Clinical epigenetics 2015, 7, 1–24. [Google Scholar] [CrossRef]
- Šestáková, Š.; Šálek, C.; Remešová, H. DNA methylation validation methods: a coherent review with practical comparison. Biological procedures online 2019, 21, 1–11. [Google Scholar] [CrossRef]
- Rauluseviciute, I.; Drabløs, F.; Rye, M.B. DNA methylation data by sequencing: experimental approaches and recommendations for tools and pipelines for data analysis. Clinical epigenetics 2019, 11, 1–13. [Google Scholar] [CrossRef]
- Gupta, J.; Margolis, D.J. Filaggrin gene mutations with special reference to atopic dermatitis. Current treatment options in allergy 2020, 7, 403–413. [Google Scholar] [CrossRef]
- Moosbrugger-Martinz, V.; Leprince, C.; Méchin, M.-C.; Simon, M.; Blunder, S.; Gruber, R.; Dubrac, S. Revisiting the roles of filaggrin in atopic dermatitis. International Journal of Molecular Sciences 2022, 23, 5318. [Google Scholar] [CrossRef]
- Chiricozzi, A.; Maurelli, M.; Peris, K.; Girolomoni, G. Targeting IL-4 for the treatment of atopic dermatitis. ImmunoTargets and therapy 2020, 151–156. [Google Scholar] [CrossRef]
- Furue, M. Regulation of skin barrier function via competition between AHR axis versus IL-13/IL-4‒JAK‒STAT6/STAT3 axis: pathogenic and therapeutic implications in atopic dermatitis. Journal of Clinical Medicine 2020, 9, 3741. [Google Scholar] [CrossRef]
- Huang, I.; Chung, W.-H.; Wu, P.-C.; Chen, C.-B. JAK–STAT signaling pathway in the pathogenesis of atopic dermatitis: An updated review. Frontiers in immunology 2022, 13, 1068260. [Google Scholar] [CrossRef] [PubMed]
- Teng, Y.; Fan, Y.; Ma, J.; Lu, W.; Liu, N.; Chen, Y.; Pan, W.; Tao, X. The PI3K/Akt pathway: emerging roles in skin homeostasis and a group of non-malignant skin disorders. Cells 2021, 10, 1219. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, K.; Hashida, R.; Miyagawa, M.; Kagaya, S.; Sugita, Y.; Matsumoto, K.; Katsunuma, T.; Akasawa, A.; Tsujimoto, G.; Saito, H. Analysis of gene expression in peripheral blood eosinophils from patients with atopic dermatitis and in vitro cytokine-stimulated blood eosinophils. Clinical & Experimental Immunology 2003, 131, 436–445. [Google Scholar] [CrossRef]
- Imai, Y. Interleukin-33 in atopic dermatitis. Journal of dermatological science 2019, 96, 2–7. [Google Scholar] [CrossRef] [PubMed]
- Spergel, J.M.; Paller, A.S. Atopic dermatitis and the atopic march. Journal of Allergy and Clinical Immunology 2003, 112, S118–S127. [Google Scholar] [CrossRef]
- Vandenberg, L.N.; Hauser, R.; Marcus, M.; Olea, N.; Welshons, W.V. Human exposure to bisphenol A (BPA). Reproductive toxicology 2007, 24, 139–177. [Google Scholar] [CrossRef]
- Rogers, J.A.; Metz, L.; Yong, V.W. Endocrine disrupting chemicals and immune responses: A focus on bisphenol-A and its potential mechanisms. Molecular immunology 2013, 53, 421–430. [Google Scholar] [CrossRef] [PubMed]
- Harb, H.; Chatila, T.A. Mechanisms of dupilumab. Clinical & Experimental Allergy 2020, 50, 5–14. [Google Scholar] [CrossRef]
- Wymann, M.P.; Zvelebil, M.; Laffargue, M. Phosphoinositide 3-kinase signalling–which way to target? Trends in pharmacological sciences 2003, 24, 366–376. [Google Scholar] [CrossRef]
- Nam, S.-E.; Bae, D.-Y.; Ki, J.-S.; Ahn, C.-Y.; Rhee, J.-S. The importance of multi-omics approaches for the health assessment of freshwater ecosystems. Molecular & Cellular Toxicology 2023, 19, 3–11. [Google Scholar] [CrossRef]
- Graw, S.; Chappell, K.; Washam, C.L.; Gies, A.; Bird, J.; Robeson, M.S.; Byrum, S.D. Multi-omics data integration considerations and study design for biological systems and disease. Molecular omics 2021, 17, 170–185. [Google Scholar] [CrossRef] [PubMed]
- Shin, G.-h.; Hong, J.-m.; Park, S.-w. Novel data archival system for multi-omics data of human exposure to harmful substances. Molecular & Cellular Toxicology 2022, 18, 277–283. [Google Scholar] [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).