You are currently viewing a beta version of our website. If you spot anything unusual, kindly let us know.
Preprint
Review
Recent Advances in Studying the Regulation of Fruit Ripening in Tomato
Altmetrics
Downloads
167
Views
150
Comments
0
A peer-reviewed article of this preprint also exists.
This version is not peer-reviewed
Abstract
Tomato (Solanum lycopersicum L.) is one of the most commercially essential vegetable crops cultivated worldwide. In addition to the nutritional value, tomato is an excellent model for studying climacteric fruits' ripening process. Despite this, the available natural pool of genes that allows expanding phenotypic diversity is limited, and the difficulties of crossing using classical selection methods when stacking traits increase proportionally with each additional feature. Genetic engineering makes it possible to introduce changes in genes of interest without changing the allelic combinations characteristic of successful commercial varieties, to enhance the expression of existing genes, to create and transfer artificial and heterologous genes, and much more. However, this requires understanding the fundamental principles of the molecular interaction. Although the leading candidate genes for these components have been identified, a complete picture of their relationship has yet to be formed. The review summarizes the latest (2017-2023) achievements related to studying the ripening processes of tomato fruits. In this work, an attempt is made to systematize the results of various research and display the interaction pattern of genes regulating the process of tomato fruit ripening.
Keywords:
Subject: Biology and Life Sciences - Plant Sciences
1. Introduction
The fruits of angiosperms are included in the staple diet of humans and livestock. The transition of plants from the vegetative growth phase to the reproductive stage is the main switch in their life cycle. Fruit ripening is initiated and regulated by the combined action of various genetic factors in response to endo- and exogenous stimuli. The functions of ripening regulators are often conserved among different angiosperms.
In addition to the nutritional importance of tomatoes, this crop is a convenient object for studying the mechanisms of fruit ripening regulation. This is explained by its simple diploid genetics, small genome size, short life cycle, ease of transient and stable transformation, pronounced ripening phenotypes, an abundance of bioinformatics data, and well-characterized mutants.
Modern widespread varieties of tomatoes were obtained through domestication and subsequent selection. The selection of seemingly desirable traits carried out without an understanding of the nature of gene relationships has contributed to the reduction of genetic diversity in tomatoes. Consumer preferences and cultivation convenience have also contributed to this. On the other hand, the introgression of elite alleles into a cultivated variety helps create a hybrid genome with the allele of interest but also introduces undesirable genetic backgrounds in the form of linked genes from the allele donor. With the help of backcrossing, breeders can level out the manifestation of unwanted traits, but this is time-consuming and not consistently effective.
Genetic engineering methods provide significant potential for studying the genetic factors regulating fruit ripening. Thus, targeted genome editing technology using the CRISPR/Cas9 system allows researchers to create knockout alleles and make precise changes to the gene sequence. This, in turn, helps to study genes’ functions, relationships, and regulation. Despite the 10-year history of using this powerful technology, the number of publications using it, in which the tomato is the object, is growing steadily yearly (Figure 1). Thus, its research potential still needs to be exhausted.
An analysis of publications where the CRISPR/Cas9 system has been used in tomatoes over the past six years has made it possible to identify topics of most interest to the scientific community (Figure 2). It turned out that more than a quarter of the total number of works is devoted to the study of genes involved in the processes of fruit ripening (27%). Many of these genes encode transcription factors and transcriptional coregulators, microRNAs, or proteins involved in the epigenetic control of gene expression. In many cases, these regulators’ molecular mechanisms of action have yet to be studied, which is the reason for the growing interest in research in this area. Also, a considerable proportion of publications are devoted to the study of the regulation of the processes of flowering and fruit development (18%). The consistently current topic of stress (abiotic and biotic) occupies a third of the total number of publications. The remaining publications cover fields devoted to other physiological processes (15%) and plant architecture and morphology (8%).
Among the most prevalent genetic engineering methods used to study the regulation of ripening processes, the use of the CRISPR/Cas9 system is expected to increase (Figure 3a). At the same time, approaches that have already become classical, such as RNA interference gene silencing, and gene over- and heterologous expression, have not lost their relevance - the number of publications using them has remained consistently high over the past six years (Figure 3, b, c). Interestingly, there are a growing number of studies using gene overexpression to study ripening. All this suggests that, despite the large amount of accumulated data, the regulation of the ripening process still needs to be fully understood.
Ripening, as the final stage of development of tomato fruits, manifests itself in the form of bright pigmentation, increased aroma and taste, and softening of the pulp. These properties make the fruits attractive to animals, which act as seed dispersal vectors. Before the onset of ripening, these physiological changes are suppressed, but once the fruit enters the ripening phase, they occur in a highly synchronized manner with dramatic differences in gene expression patterns. Such radical changes are regulated by environmental factors such as temperature and light and by internal regulators, including hormones, transcription factors, and epigenetic modifications. Due to the commercial and biological importance of tomato fruit ripening, the mechanisms regulating this process are being actively studied.
2. Transcription factors regulating ripening
MADS-box genes are among the most widely represented and diverse transcription factors; consequently, they mediate various biological processes. Among them, regulating fruit ripening is one of the most prominent roles of MADS-box (MCM1, AGAMOUS, DEFICIENS, and SRF) genes. The transcription factor RIN (RIPENING INHIBITOR) has long been considered a major ripening regulator. RIN encodes a SEPALATA class MADS-box transcription factor. MADS-box family transcription factors typically function as multimers, and the MADS-box proteins TAGL1 (TAG-like) and two FRUITFULL (FUL) homologs (TDR4/FUL1 (tapetum degeneration retardation) and MBP7/FUL2 (MADS-box protein)), are coregulators with RIN and ripening regulators with overlapping functions [1]. Silencing of TAGL1 resulted in decreased levels of amino acids in fruit: aspartic acid, L-tyrosine, L-glutamine, L-phenylalanine, L-valine, L-leucine, isoleucine, and 5-caffeoylquinic acid [2]. TAGL1 was also found to regulate the synthesis of the glycoalkaloid α-tomatine negatively. As discussed in [3], TDR4/FUL1 and MBP7/FUL2 do not regulate ethylene biosynthesis but influence fruit ripening in an ethylene-independent manner. RIN often binds to demethylated sites in the promoter regions of ripening-related genes. RIN is induced early in ripening and stimulates ethylene-dependent and ethylene-independent pathways that promote ripening. Mediators in this process are response factors to ethylene (ERF, ethylene-responsive factor) and auxin (ARF, auxin response factor). ERF and ARF control their respective hormonal signaling pathways, regulating gene expression and hormonal signaling.
Several recent studies have clarified the function of RIN. Thus, [4] found that although RIN function is required for full ripening, RIN is not required for the initial ripening induction. The authors suggest that RIN acts redundantly (i.e., there are RIN homologs) or RIN-independent ripening induction occurs due to other transcription factors. In the second case, the authors concluded that a RIN-independent activator can induce the transcription of ripening-related genes even in RIN-deficient plants. Still, a mutant (defective) RIN protein can inhibit its activity. The chimeric transcription factor RIN-MC exhibits a negative role in ripening, promoting the mutant rin phenotype [5]. Other authors think that low ethylene concentrations initiate the ripening of mature green fruits, activate RIN expression, and lead to other changes, including a transition to a burst of autocatalytic ethylene synthesis [6]. Combined with the ethylene biosynthesis gene ACS2 (1-aminocyclopropane-1-carboxylate synthase), RIN has been shown to regulate the heat shock genes HSP17.7 [7] negatively. Therefore, RIN, ethylene, and other factors are necessary to complete the complete fruit ripening program. RIN is not only an activator of ripening but also a repressor of excessive softening [8]. It was found that when controlling the fruit ripening process, RIN binds to 6 lncRNAs [9].
RIN is reported to directly activate the expression of a novel gene, E6-2, involved in tomato fruit ripening [10]. Silencing of E6-2 leads to a delay in fruit ripening suppression of CNR (colorless non-ripening), PG (polygalacturonase), and ERF4 (ethylene-responsive factor) was observed, a decrease in the accumulation of carotenoids and lycopene (due to the suppression of PSY1, PDS and ZDS (phytoene synthase, phytoene desaturase, and zeta-carotene desaturase, respectively)), ethylene (decreased expression of the biosynthetic genes ACS2, ACO1 (1-aminocyclopropane-1-carboxylic acid oxidase), ACO3 and ethylene-sensitive E4, E8), increasing the content of pectin, cellulose, starch and soluble sugar (suppression of cell wall metabolism genes TBG4 (tomato beta-galactosidase), PL (pectate lyase), EXP1 (expansin), and XTH5 (xyloglucan endotransglucosylase/hydrolase)). The broad phenotypic pattern of RIN silencing is an attractive marker for testing molecular editing tools [11,12,13].
Genes with the NAC domain (NAM, ATAF1/2, and CUC2 (apical meristem, ARABIDOPSIS TRANSCRIPTION ACTIVATOR FACTOR, and cup-shaped cotyledon, respectively)) are considered to be another transcription factors that regulate tomato fruit ripening. It has been shown [14] that inhibition of NOR-like1 reduces ethylene production, delayed softening and loss of chlorophyll, and reduced lycopene accumulation. Activation of ethylene synthesis genes by NOR (non-ripening) and NOR-like genes is discussed further in [15]. Knockout of NAC-NOR suppressed fruit ripening (inhibition of ethylene synthesis, reduction of carotenoid accumulation, and fruit softening), and the opposite effect was observed with its overexpression [16]. Replacement of thymine with adenine in the ALC gene (alcobaca, NOR mutation) using homologous recombination contributed to an increase in the shelf life of tomato fruits [17]. Expression of peach NAC1 in tomatoes has been shown to enhance ripening in a delayed ripening (NOR) mutant and restore the synthesis of volatile esters [18]. Overexpression of NAC6 resulted in increased levels of endogenous abscisic acid, which affected the transcription of ripening genes [19]. Transfer of the kumquat NAC22 gene to tomato increased the expression of most carotenoid biosynthesis genes, accelerated the transformation of plastids into chromoplasts, and promoted color changes [20]. NAM1 (no apical meristem), another factor with an NAC domain responsible for the regulation of ethylene biosynthesis, also controls tomato ripening, as confirmed by delayed ripening in CRISPR/Cas9 mutants and accelerated ripening for lines overexpressing NAM1 [21]. Repression of NAM gene domains is carried out by miR164a [22,23,24]. In addition, the HWS (HAWAIIAN SKIRT) gene, encoding an F-box protein, regulates the number of floral organs by modulating the transcription levels of the miR164, CUC1 and CUC2 (cup-shaped cotyledon) genes. HWS is also involved in petals’ cell proliferation and mitotic growth [25].
It was previously shown that a representative of genes with the NAC domain NAP2 (Arabidopsis NAC domain-containing protein) activates the aging gene SAG113 (senescence-associated gene), protein phosphatase), chlorophyll degradation genes SGR1 (stay-green), PAO (polyamine oxidase), and NAP2, and also directly controls the expression of genes essential for the biosynthesis of abscisic acid NCED1 (9-cis-epoxycarotenoid dioxygenase), ABCG40 (Arabidopsis thaliana ATP-binding cassette), and CYP707A2 [26]. These interactions suggest the influence of NAP2 on leaf senescence and yield in tomato. Another new gene, named by the authors HEBE in honor of the Greek goddess of youth, has similar functions [27].
Numerous studies have shown that the CNR gene is the most important regulator of tomato fruit ripening. However, recent research [28] has called this assumption into question. CNR knockout lines exhibited only a ripening-arrested phenotype, while NOR knockout (non-ripening) lines exhibited a partial non-ripening phenotype similar to RIN mutants. Both knockouts differed from the strong, non-ripening phenotypes of their natural mutants. It became apparent that the expression of characteristic ripening genes, such as ACS2, ACO1, PSY, PG, and EXP, is not entirely suppressed in CRISPR/Cas9 lines compared to natural mutants. As the authors concluded, differences in the expression of the genes in question are explained by different degrees of methylation, and they also concluded that the regulatory network of transcription factor genes is redundant. Regulation of NOR may also be associated with something else: sulfoxidation of the NOR transcription factor with the help of MSRs (methionine sulfoxide reductase) proteins modulates the ripening process by reducing the DNA-binding ability of NOR [29].
It is known that the fruits of the tomato epimutant cnr fail to ripen and remain colorless. The SPL (SPOROCYTELESS) gene family consists of a group of genes encoding SBP (SQUAMOSA promoter binding proteins)-box transcription factors, and their protein products bind to the promoter of the floral meristem identity gene SQUAMOSA. Evidence shows that SPL-CNR interacts with SnRK1 (SNF1-related protein kinase) [30]. Suppression of SnRK1 by virus-induced gene silencing (VIGS) inhibits fruit ripening and leads to decreased expression of a wide range of ripening-related genes. This suggests that SnRK1 transcription and subsequent post-translational SPL-CNR-SnRK1 interaction are biologically crucial for tomato fruit ripening. The authors suggest that the involvement of SnRK1 in fruit ripening may be due to the physical interaction of proteins between the SnRK1 gene product and SPL-CNR and subsequent phosphorylation of SPL-CNR due to the kinase activity of SnRK1 [30].
The role of some MBP transcription factors in the ripening process has recently been studied. Thus, suppression of the MBP8 factor shortened the fruit ripening time, suggesting an increase in the activity of ethylene synthesis genes [31]. Meanwhile, carotenoids accumulated to higher levels, and the expression of PSY1, PDS, and ZDS was enhanced in MBP8 RNAi-silenced fruits. The activity of cell wall genes also changed, manifested in the softening of fruits. Silencing MBP15 in [32] delayed tomato ripening, and gibberellin, carotenoid, and ethylene biosynthesis genes were repressed. MBP15 was found to interact with RIN [32].
GRAS (gibberellic acid insensitive (GAI), repressor of GAI (RGA), and scarecrow (SCR)) proteins are plant-specific transcription factors that play critical roles in plant development and stress response. It turned out that they also take part in regulating fruit ripening. For example, silencing GRAS2 reduces tomato fruit weight, which has been attributed to insufficient levels of gibberellic acid during initial ovary development [33]. Overexpression of GRAS4 accelerated fruit ripening (due to activation of expression in the promoter region of ethylene biosynthesis genes and repression of the negative regulator of ripening MADS1). It increased the total content of carotenoids [34]. GRAS24, in addition to flowering and ripening, is responsible for a variety of other agronomic traits, including plant height, leaf architecture, number of lateral branches, root length, and the observed pleiotropic effects in plants overexpressing GRAS24 are due to impaired modulation of gibberellin and auxin signaling [35].
Transcription factors of the WRKY superfamily exhibit up-regulation during fruit ripening. WRKY32 binds to W-box and similar motifs in the regulatory region of the YFT1 (yellow-fruited tomato) promoter and induces its expression [36]. YFT1 encodes the EIN2 protein, a major ethylene signal transduction component. Suppression of ethylene production resulted in delayed chromoplast development, decreased carotenoid accumulation, and a yellow fruit phenotype. Twelve WRKY genes were also shown to be ethylene-responsive (ER), 8 of which activated the promoters of color change-associated genes (PPH (pheophytinase), PAO (polyamine oxidases), PSY1, and PDS) [37]. In addition, protein interactions were found between WRKY17 and RIN/ERF2b/ERF7, WRKY33 and ERF7, WRKY54 and ERF2b, WRKY16 and WRKY1, which only confirms the complexity of the networks of ripening regulators [37].
3. Epigenetic modifications as regulators of ripening
DNA methylation has a critical role in a wide range of cellular functions. For example, a decrease in DNA methylation levels can be observed during fruit ripening, which is explained by DNA demethylases (DML) activation. In DML2 loss-of-function mutants generated by targeted editing, increased DNA methylation was found not only in genes induced during ripening but also in genes repressed during ripening [38]. However, a recent study found that the highly mobile protein positively regulates DML2 expression of 3-hydroxy-3-methylglutaryl-coenzyme A reductase (HMGA) [39]. In [40], expression of the mammalian demethylase TET3c (ten-eleven translocation) in tomatoes resulted in the activation of expression of the previously undescribed gene CEN1.1. The activation intensity of CEN1.1 expression correlated with increased hypomethylation in its promoter, suggesting that CEN1.1 expression is associated with DNA methylation of CHH promoter sites (H=A/C/T). Phenotypically, CEN1.1 has emerged as a repressor of flowering in tomatoes, leading to the development of leaves on inflorescences. Paradoxically, this led to an increase in the number of fruits but to a longer time for their ripening. Thus, this study provides an exciting approach to identifying methylation-associated genes.
In plants, cytosine methylation plays a crucial role in suppressing the movement of transposable elements. Methylation is maintained by DNA methyltransferases MET1, and CMT3 (chromomethylase), as well as additional proteins (for example, DDM1 (decreased DNA methylation)) involved in maintaining heterochromatic structure. The methylase encoded by MET1 is a key DNA methylase responsible for maintaining CG methylation in plants. Loss-of-function mutants of the MET1 gene had pleiotropic developmental phenotypes manifested as small curled leaves, defective flowers, and parthenocarpic fruits [41]. Also, the knockout of MET1 resulted in changes in the expression profiles of RIN target genes, such as ACC2 (acetyl-CoA carboxylase 2). In another study, suppression of MET1 by VIGS in a hypermethylated epimutant CNR promoted vivipary development [42]. The authors explain this by a decrease in the concentration of abscisic acid and NCED transcripts involved in its biosynthesis. The NCED silencing had similar consequences. Methyltransferase DRM7 (domains rearranged methyltransferase) has been shown to influence chloroplast development by modulating starch accumulation and chlorophyll synthesis. It has an epi-effect on leaf senescence, affecting tomatoes’ vegetative growth [43]. The transient expression of arginine methyltransferase PRMT1.5 in tomatoes inhibited the accumulation of carotenoids and anthocyanins [44].
Short interfering RNAs (siRNAs) also mediate DNA methylation through RNA-directed DNA methylation (RdDM). It has been established that heterochromatic mobile elements in plants with DDM1 dysfunction are deprived of mCG and mCHG, which generally keep them inactive [45]. Methylation of CHH sites increased for some heterochromatic transposons and, conversely, decreased for those localized in euchromatin. Knockout of CMT4 chromomethylase caused severe morphological changes in tomato plants, accompanied by defects in leaves, pollen, and seeds [46].
Another mode of epigenetic regulation is post-translational modifications of histones. Histone acetylation is known to be associated with gene activation. In contrast, histone methylation can be associated with either activation or repression depending on the lysine residue and the number of methyl groups added. In [47], RNA-seq profiling showed a significant increase in the expression of methylases MET1 and CMT3 and a minor increase in the demethylase DML2 during the fruit set, which is associated with their role in maintaining post-replication DNA methylation during extensive cell division characteristic of early stages of development fetus However, their abundance was significantly lower than that for histone marks H3K9ac and H3K4me3, determined using chromatin immunoprecipitation sequencing. This implies that changes in the transcriptional profile underlying the fruit set are more closely related to histone modifications than methylation. Histone modification is based on the histone methyltransferase genes SDG27, SDG5, and SDG16 (set domain group). However, the authors could not create homozygous loss-of-function mutants of these genes, suggesting their exceptional biological importance. Mutants heterozygous for these genes exhibited parthenocarpic fruits. The function of other histone lysine methyltransferases SDG33 and SDG34 was revealed in [48]. They were found to regulate the expression of nitrogen-responsive genes and physiological changes in an organ-specific manner.
It has been demonstrated that histone demethylation leads to activating tomato fruit ripening genes [49]. Here, the JMJ6 (Jumonji C-terminal domain-containing demethylase) gene was found to encode a histone lysine demethylase that specifically demethylates H3K27. Its overexpression accelerates the ripening of tomato fruits, which is associated with increased expression of the RIN, ACS4, ACO1, PL, and TBG4 genes. As study [50] showed, JMJ4 mediates abscisic acid-induced leaf senescence in tomatoes.
By knocking out the HTA1 genes of histone H2A and subsequent production of double homozygous mutants, changes were identified in the expression patterns of many biological ripening processes, including cell redox homeostasis, mRNA splicing, cell cycle regulation, translation, etc. [51]. Moreover, for three genes of carotenoid biosynthesis, PSY1, PDS, and VDE, expression was high regardless of the fruit ripening stage [51]. Histone deacetylation has been associated with transcriptional repression. Histone deacetylases carry out this process. There is evidence that they can act as both positive and negative ripening regulators. Indeed, RNAi silencing of the HDT3 (histone deacetylase) gene led to the suppression of genes for ethylene synthesis (ACS2, ACS4, ACO1, ACO3), carotenoids (PSY1), cell wall metabolism (HEX (acetylhexosaminidase), MAN (mannosidase), TBG4, XTH5, and XYL (xylanase)), as well as general genes, associated with ripening (RIN, E4, E8, PG, Pti4, LOXB (lipoxygenase)) [52]. In contrast, in [53,54], silencing of the HDT1 gene led to opposite results for these same transcripts. In this regard, the molecular mechanisms of regulation of these genes remain to be studied.
Histone acetyltransferase GCN5 acetylates histone H3 lysine (H3K14ac) and affects the levels of H3K9ac and H3K27ac. Its suppression leads to the loss of shoot apical dominance and a decrease in the size of the plant apical meristem [55]. It has also been established that GCN5 can increase WUSCHEL transcript levels. The expression of WUSCHEL can also be regulated by chromatin remodeling factors, such as the histone deacetylase HDA19 [56]. Here, the deacylation mechanism was found to involve the inhibitor gene IMA (inhibitor of meristem activity) acting as an adapter protein to form a chromatin remodeling complex together with the zinc finger protein C2H2 KNU (KNUCKLES) and the transcriptional corepressor TOPLESS.
4. Hormonal control of ripening
Auxin regulation is involved in all plant processes, including cell elongation and division, the formation of the architecture of roots, leaves, inflorescences, the development of embryos and fruits, and responses to stress. The leading site of auxin biosynthesis is young leaves and their primordia. The amino acid tryptophan is the primary precursor of auxins. Knockout of any of the auxin synthesis genes is associated with lethal phenotypes, so attention is paid to genes providing auxin-mediated inactivation (GH3, GRETCHEN HAGEN), transport (PIN, ABCB (PIN-FORMED, ATP binding cassette subfamily B, respectively)), and signal transduction (ARF (auxin response factor), Aux/IAA).
By conjugating auxins to amino acids for storage or degradation, members of the GH3 family, encoding acyl acid amidosynthetases, are critical for maintaining auxin homeostasis. In tomatoes, GH3.15 has been shown to regulate lateral root development and response to gravitropism by modulating auxin homeostasis [57], GH3.8 controls plant height [58], GH3.4 negatively regulates mycorrhization [59,60], GH3.2 affects fruit ripening in the early stages [61].
PINs are one of the facilitators of intercellular auxin transport. VIGS PIN1 accelerates flower abscission by increasing the accumulation of auxin in the ovule and reducing the auxin content in the abscission zone [62], and its negative regulator is the transcription factor MBP9 [63].
ARFs are plant-specific transcription factors that directly bind to auxin response elements in the promoters of auxin-responsive genes. ARF5 has been shown to regulate fruit set and development [64], ARF10 is involved in the accumulation of chlorophyll and sugar during fruit ripening [65], ARF19 is involved in leaf development [66], several ARFs (ARF6A, ARF8A, ARF8B, and ARF24) interact with the transcriptional repressor IAA9 [67] and regulate leaf shape [68], ARF10A is essential for the growth of leaf blades and formation of floral organs [69].
Phytohormones gibberellins regulate various physiological processes of plants. They promote plant growth participate in stem elongation, expansion of leaf blades, pollen development, flowering, and seed germination. Gibberellins are synthesized in plastids and then modified in the endoplasmic reticulum and cytosol.
DELLA (GRAS gene encodes protein containing D-E-L-L-A amino acid sequences) proteins are nuclear-localized negative growth regulators. Gibberellins promote DELLA degradation by assembling the E3 ubiquitin ligase complex, followed by protein degradation. DELLA is encoded by the PROCERA gene, and its loss of function in the homozygous state results in dwarfism [70] and parthenocarpy [71]. The degradation of proteins, including DELLA, is controlled by a complex regulatory network involving connections between several signaling pathways [72]. DELLA proteolysis is mediated by the gibberellin-activated receptor GID. Knockout of their coding genes also results in a dwarf phenotype [73]. There is evidence of cross-signaling between the gibberellin and abscisic pathways [74], and the DELLA protein is an activator of abscisic acid transporters (AIT), regulating transpiration through stomatal closure [75]. Tomato PROCERA activity is assumed to be necessary to transition tomatoes to flowering. DELLA protein directly or indirectly promotes the expression of SFT (SINGLE FLOWER TRUSS) in leaves, as well as SBP and AP1/MC, together with microRNAs in the shoot apex [76].
Recently, factors mediating gibberellin-dependent regulation have also received attention. Thus, silencing of the GRAS15 transcription factor led to pleiotropic phenotypes, including reduced plant height, small leaf size with pointed edges, as well as an increased number of nodes, lateral shoots, and petiole length, which is explained by the suppression of gibberellin synthesis genes [77]. The helix-loop-helix transcription factor gene PRE2 is induced by gibberellin. Its silencing has been shown to cause reductions in fruit size, seed size, pericarp thickness, and placental size [78]. These changes are associated with decreased expression of xyloglucan endotransglucosylases XTH2 and XTH5. PREs regulate many processes - their overexpression in tomatoes leads to multiple morphological changes, including changes in leaf angle, internode length, leaf curl, and pigment composition [79,80].
Cytokinins promote the development of shoots, provide stress resistance, and delay aging. These isopentenyladenine derivatives are formed mainly in the roots and transported to the aerial parts. The main rate-limiting enzyme for cytokinin synthesis is isopentenyltransferase (IPT). It has been shown that overexpression of the IPT gene leads to significant phenotypic changes and slower leaf senescence only under the control of a root-specific promoter [81].
Cytokinin catabolism is carried out by cytokinin oxidases (CKX). Overexpression of CKX2 in tomato fruit decreased cytokinin levels [82]. It is also shown here that endogenous cytokinins regulate the division of pericarp cells, which subsequently determines the size of the fetus. Ethylene is the simplest unsaturated hydrocarbon with the formula C2H2. In plants, it acts as a global regulator of developmental processes. The ethylene biosynthetic pathway includes three steps: S-adenosylmethionine synthetase (SAMS) modifies methionine to form S-adenosylmethionine (SAM), SAM is converted to 1-aminocyclopropane-1-carboxylic acid (ACC) by synthase (ACS), and in the last step ACC converts ACC oxidase (ACO) with the formation of ethylene.
In studies on tomatoes, ethylene is considered a participant in signaling cascades, including during the ripening process. It has been established that its synthesis during ripening is presumably regulated by FER receptor kinases (FERONIA). FERL6 and FERL1 were found to interact physically with the SAMS promoter [83]. Expression of FER genes in tomatoes showed negative regulation of ethylene accumulation at the initial stages of fruit development and, as a consequence, delayed fruit ripening.
The promoter of the transcription factor EIN3 (ethylene insensitive) has been shown to contain several motifs associated with hormones influencing fruit development and ripening [84]. Overexpression of EIN3 in tomatoes resulted in the activation of the expression of ethylene biosynthesis genes ACO1, ACS1, and SAMS1, which promoted early fruit ripening. Accordingly, EIN3 silencing showed the opposite effects. An EIN3-like gene causes premature onset of ovule senescence [85].
Ethylene is bound by a family of ETR (ethylene receptor) proteins located in the membrane of the endoplasmic reticulum. ETRs have functional redundancy. ETR3-mediated signaling inhibits pollen tube growth without sufficient ethylene [86]. ETR3 promotes the activation of cell wall remodeling genes and Ca2+ transporters — overexpression of ETR7 results in earlier flowering, short plants, and small fruits [87]. Targeted base substitution in ETR1/2 causes a delay in ripening and ensures prolonged storage of fruits [88,89].
Ethylene response factors (ERFs) are signaling components involved in ethylene-dependent developmental processes. They can perform both positive and negative regulation of target genes. Their number is large, as is the specificity of the reactions of tomato genes to ethylene: regulation of fruit ripening processes [90,91,92,93], control of aging [94], participation in the activation of protective reactions [95,96,97], growth [98,99], accumulation of chlorophyll and formation of chloroplasts [100], and regulation of other signaling pathways [101].
Brassinosteroids, which include various polyhydroxylated steroidal phytohormones, influence many critical agronomic traits related to growth, photosynthesis, morphology, and yield. The synthesis of brassinosteroids occurs along three pathways, in which campesterol is the initial substrate. Crosstalk between brassinosteroids and redox signals suggests a direct involvement of the former in the plant response to stress [102,103]. However, recent studies also reveal a connection between the brassinosteroid and ethylene pathways. Recently, it was demonstrated that overexpression of one of the genes for the brassinosteroid synthesis enzymes DWARF (DWF) in tomatoes promotes fruit softening, lycopene synthesis, and ethylene production, while gene knockout inhibits them [104]. It was concluded that APETALA2a (AP2a) promotes ethylene signaling to regulate brassinosteroid signaling. Also, tomatoes with overexpression and silencing of the cytochrome P450 monooxygenase CYP90B3 gene showed a correlation of the content of bioactive brassinosteroids with the processes of tomato fruit ripening, including softening, the content of soluble sugars and aromatic volatiles [105].
Research into brassinosteroid-dependent pathways is ongoing. The specific receptor for brassinosteroids is BRI1 (BRASSINOSTEROID INSENSITIVE1). Upon binding of brassinosteroids to its extracellular domain, dimerization of BRI1 and the coreceptor BAK1 (BRI 1-associated receptor kinase 1) occurs. The signal is then transmitted through a phosphorylation cascade involving BSK1 (BR-signaling kinase 1), CDG1 (constitutive differential growth), BSU1 (BRI1 SUPPRESSOR), and BIN2 (BRASSINOSTEROID INSENSITIVE2). Subsequently, BIN2 is inactivated, and two transcription factors, BZR1 (brassinazole resistant) and BES1 (BRI1-extra microsporocytes-suppressor 1), are dephosphorylated by protein phosphatase PP2A. BZR1, BES1, as well as other nuclear factors (for example, BIM1) are regulators of brassinosteroid-dependent genes.
Overexpression of BRI1 [106] in tomatoes improved carotenoid accumulation by increasing the expression of DXS (1-deoxy-D-xylulose 5-phosphate synthase), GGPS (geranylgeranyl pyrophosphate synthase), and PSY1. In addition, brassinosteroids induced the expression of genes involved in its ethylene biosynthesis (ACO1 and ACS2). Similar results were achieved by modifying threonine-1050 BRI1, resulting in plants with high levels of BRI1 autophosphorylation [107]. A recent study on BRI1 showed that the receptor also positively regulates a tomato’s tolerance to cold stress [108]. Brassinosteroids are capable of inducing early flowering. This is supported by the interaction of the suppressor of BIN2 signaling with the early flowering locus FRIGIDA [109]. There is evidence that canonical signaling pathways initiated by BRI1 are involved in xylem differentiation and wood formation in tomatoes through activation of the BZR1/2 transcription factors [110]. The BZR1 homolog has been shown to interact with BIM1 to act as a negative regulator of pericarp cell expansion [111]. According to available information, BZR1 is also a trans-activator of the promoter of the SUN gene (encodes Sad1/Unc-84 (SUN)-domain proteins), responsible for elongation of tomato fruits, and BZR1-knockout tomato phenotypes show redundancy of its homologs [112]. In addition, brassinosteroids promote bud growth in tomatoes through direct transcriptional regulation of BRANCHED1 (BRC1) via the signaling component BZR1 [113].
In the study [114], the authors focused on BES1, a key transcription factor in the brassinosteroid signaling pathway. BES1 was found to bind to the promoter of the fruit-softening inhibitor PMEU1 (pectin methylesterase). Knockdown or knockout of BES1 in tomatoes resulted in increased shelf life without negatively affecting the appearance and nutritional composition of the fruit.
Altered regulation of brassinosteroids may influence cell elongation and division, leading to altered fetal morphology. For example, a premature stop codon at the GLOBE locus containing a brassinosteroid hydroxylase sequence resulted in a spherical phenotype of tomato fruit, which had a flattened shape in the wild type [115]. Since GLOBE and FW3.2 (KLUH) were found to be members of the same cytochrome P450 family, the authors hypothesized that both may act similarly in regulating fruit size and shape.
During oxidative stress following pesticide application, plants use glutathione to clear excess reactive oxygen species. Brassinosteroids induce pesticide metabolism by activating GRX (glutaredoxin) gene expression through transcription factors [116,117].
Abscisic acid is considered an antagonist of brassinosteroids during fruit ripening. The abscisic acid signaling pathway consists of the family of receptor proteins PYR (PYRABACTIN RESISTANCE), PYL (PYR1-like), RCAR (regulatory components of ABA receptors), protein phosphatases PP2C, and SnRK2 kinases. There is evidence of positive regulation of abscisic acid biosynthesis by brassinosteroid signaling. BZR1 (brassinazole resistant) was found to mediate brassinosteroid signaling by promoting abscisic acid biosynthesis through direct transcriptional regulation of NCED1 [118]. Here, we showed that BIN2 negatively regulated BZR1 protein accumulation and cold tolerance by suppressing abscisic acid biosynthesis.
Suppression of PP2C3 expression in tomatoes accelerated the onset of fruit ripening and affected their glossiness by changing the external structure of the epidermis [119]. In transgenic plants, an increase in the expression of SnRK2, PYL receptors, various cutin synthesis and transfer genes, and CYP (cytochrome P) genes was observed. The role of PP2C as a negative regulator in abscisic acid signaling was further supported in [120], where alteration of PP2C5 expression affected fruit quality traits, including pericarp thickness, shape, seed number, and soluble solid content. In addition, PP2C1 silencing increased accumulation of endogenous abscisic acid and accelerated ethylene release in transgenic tomatoes compared to wild-type fruit [121]. PP2C1-RNAi lines had abnormal flowers, and pedicel abscission was impaired.
Abscisic acid homeostasis is regulated by its conjugation with glucose using uridine diphosphate glucosyltransferases (UGT). It was shown that RNAi silencing of the UGT75C1 gene significantly increases the level of expression of the CYP707A2 hydrolase gene while not affecting the expression of the key gene for abscisic acid biosynthesis NCED1 [122]. Suppression of UGT75C1 expression significantly accelerated fruit ripening by increasing abscisic acid levels and promoting early ethylene release.
The PYL9 protein has been identified as a positive regulator of abscisic acid signaling [123]. Depending on abscisic acid concentration, PYL9 can inhibit the protein phosphatase PP2C. In tomatoes overexpressing PYL9, fruit ripening was significantly accelerated due to the early release of ethylene.
The abscisic acid-induced oxidase gene DAO2 (dioxygenase for auxin oxidation) inhibited hypocotyl elongation in tomatoes, exhibiting an antagonistic role to auxins [124].
SnRK phosphorylation is mediated by the protein kinase MAPK11, thereby regulating abscisic acid biosynthesis and signaling [125].
In addition, a new transcriptional repressor of abscisic acid biosynthesis, EAD1 (ERF-associated amphiphilic repression (EAR) motif-containing ABA down-regulated), was recently discovered [126]. Although the authors have not studied the molecular mechanism of repression, its implementation is possible either through the recruitment of histone deacetylases with subsequent formation of a complex with co-suppressors or through direct or indirect binding to transcription factors.
Hydrogen sulfide counteracts the effects of ethylene during ripening. The assimilation of sulfates in chloroplasts can produce endogenous hydrogen sulfide, and the main enzymes in this process are sulfite reductases. Cytosolic hydrogen sulfide can also be generated from cysteine by cysteine desulfhydrase 1 (DES1/LCD1). Loss-of-function mutations in LCD1 in tomatoes [127] increase the expression of genes for ethylene synthesis (ACO1, ACO3, and ACS2), carotenoids (PSY1, PDS, and ZDS), and cell wall metabolism (CEL2, EXP, XTH5, PG, and TBG4). Knockout of the tomato D-cysteine desulfhydrase (DCD) gene results in increased expression of ripening-related genes, including NYC1, PAO, SGR1, PDS, PSY1, ACO1, ACS2, E4, CEL2, and EXP [128].
An attempt to understand hydrogen sulfide-mediated regulation of ripening is made in [129]. The authors suggest that the ubiquitin-protein ligase BRG3 undergoes persulfidation at two cysteine residues, leading to a decrease in ubiquitinating activity and its interaction with the repressor transcription factor WRKY71. This leads to increased binding of WRKY71 to the promoter of cyanoalanine synthase CAS1, which inhibits its transcription and, thus, prolongs fruit ripening.
There is also confirmation that hydrogen sulfide is a regulator of aging, which is noticeable in changes in the expression of chlorophyll degradation genes (NYC1, PAO, PPH, SGR1) and the aging-associated gene SAG [130].
5. Abiotic ripening factors
Fruit ripening is also regulated by signaling systems activated in response to abiotic stimuli. It has been reported that changes in light sensitivity and light-sensitive signaling in tomatoes can significantly change fruit development and quality characteristics. In this context, phytochromes act as molecular switches in response to light. Following light activation, phytochromes deactivate photomorphogenic response repressor proteins (e.g., COP1, CUL4, DDB1, DET1, and PIF).
Using RNAi silencing of the phytochrome genes PHYA, PHYB1, and PHYB2, it was shown that PHYA positively affects the differentiation and division of tomato plastids through changes in the expression of both light-dependent genes and cytokinin-dependent genes [131]. Regulators of carotenoid biosynthesis (GGPS, PSY1, and PDS) were also affected, resulting in decreased carotenoid biosynthesis during fruit ripening.
As for the repressor proteins mentioned above, their effect on ripening is also being studied. Thus, according to [132], overexpression of COP1 (CONSTITUTIVE PHOTOMORPHOGENIC) from Solanum melongena in tomatoes caused a delay in fruit ripening by 3-6 weeks. These transgenic plants showed decreased ethylene production due to suppressing the expression of the central genes of its biosynthesis ACO1, ACO3, and ACS2. The carotenoid biosynthesis genes PSY1, PDS, and ZDS were also downregulated. In [133], using the DDB1, DET1, and CYC-B genes as an example, the multiplex Target-AID (activation-induced cytidine deaminase) technique was developed. As a result, the authors obtained two lines of triple mutants, in which each gene had two-point substitutions, which showed a higher accumulation of carotenoids and lycopene compared to the wild type. In tomatoes, PIF-dependent light signaling has been reported to regulate fruit development and influence nutritional value and ripening time. Transient overexpression of PIF3 in tomato fruit resulted in decreased GGDR mRNA levels, which was inversely related to PIF3 transcript levels [134]. These data indicate that PIF3 mediates PHY-dependent regulation of tocopherol biosynthesis through transcriptional inhibition of geranylgeranyl diphosphate reductase GGDR expression in tomato fruit. Evidence shows that PIF4 can regulate hypocotyl elongation, plant growth, flowering, and leaf senescence in response to light and temperature [135]. The authors support this statement by obtaining tomatoes with RNAi-mediated knockdown of PIF4, which showed increased carotenoid content, accelerated fruit ripening time, and delayed leaf senescence. A small number of flowers and a decrease in vegetative mass were observed in such plants. Knockout of PIF3 using CRISPR/Cas9 led to the arrest of phase I of pollen mitosis, which was reflected in its non-viability [136]. Glutamate synthase (GLT1) and cell wall invertase (CWIN9), involved in auxin and sugar homeostasis, respectively, have also been shown to colocalize with PIF3 in anthers and are directly regulated by PIF3. Knockout lines of GLT1 and CWIN9 (cell wall invertase) showed a similar phenotype. VIGS-mediated silencing of the light-signaling transcription factors HY5 and PIF3 led to changes in glycoalkaloid levels in tomato leaves compared to wild type, suggesting their involvement in the regulation of target genes of glycoalkaloid metabolism [137].
While the most abundant antioxidant in tomato fruit is the lipophilic carotenoid lycopene, levels of water-soluble flavonoids (including anthocyanins) are suboptimal. Plants accumulate anthocyanins in response to various stress events such as low temperature, drought, UV radiation, intense light, and nutrient deficiency, acting as an antioxidant and photoprotective agent. The bZip transcription factor HY5 is believed to be a significant regulator of anthocyanin accumulation in plants in response to light [138,139]. However, research [140] has cast doubt on the accuracy of this statement. By creating HY5 knockout mutants, the authors demonstrated a reduced anthocyanin content, which suggests the presence of additional pathways for their synthesis independent of HY5. Indeed, eight candidate anthocyanin transcription factors have been identified.
A recent study has uncovered the function of the little-studied PHY-F. It turned out that PHY-F is a low-flux radiation sensor [141]. It forms dimers with PHYA and/or PHYB, with which it makes additive contributions to various processes of photomorphogenesis.
In addition to the phytochromes of red and far-red light receptors, there are also cryptochromes of blue light receptors – CRY1 and CRY2. Tomato lines overexpressing CRY1a showed significant accumulation of anthocyanins through the regulation of genes encoding key enzymes of anthocyanin biosynthesis (e.g., AN2, DFR (dihydroflavonol 4-reductase)) [142]. The same study showed that blue light consistently induced overexpressing tagged HY5 protein accumulation in tomatoes. In addition, it was shown that under the influence of blue radiation, repression of COP1 (CONSTITUTIVE PHOTOMORPHOGENIC) transcription was observed, which was confirmed by the creation of lines with RNAi-COP1. Ultimately, the silencing of HY5 and two anthocyanin biosynthesis genes (CHS1 (chalcone synthase) and DFR) in CRY1a lines was accompanied by a decrease in anthocyanin accumulation. Moreover, CRY1a was found to be critical for regulating starch accumulation in chloroplasts by inducing starch degradation through the transcription factor HY5 [143]. Induction of transcription of genes associated with starch degradation under the influence of blue radiation in CRY1a- or HY5-overexpressing plants was also confirmed.
It is known that in tomato, the R2R3-MYB group of factors regulating anthocyanin biosynthesis is represented by AN (ANANTHA) genes. Currently, their biological function is being actively clarified. For example, by generating loss-of-function mutants of AN2, the authors identified it as a positive regulator of anthocyanin biosynthesis in tomato vegetative tissues [144]. In addition to reduced anthocyanin content, the mutants had a dwarf phenotype. Overexpression of AN2 resulted in changes in multiple fruit qualities [145]. Thus, increased production of ethylene and increased content of anthocyanins, phenols, and flavonoids were observed. The content of aromatic volatiles such as aldehydes, phenylpropanoid derivatives, and terpene volatiles was also increased in these fruits. Thus, AN2 was shown here to regulate the transcription of genes in several metabolic pathways. Additionally, it was found that loss-of-function mutations in the AN2 ortholog in wild tomato impair anthocyanin synthesis [146]. Overexpression of ANT1 in tomatoes enriched the anthocyanins in leaves, contributing to more intense light absorption in the blue and red spectrum [147]. However, introducing knockout mutations into the AN2-like gene rather than ANT1 (ANTHOCYANIN) essentially eliminates the accumulation of anthocyanins [149,150]. It was found that AN2-like activated the expression of DFR; however, when AN1 was knocked out, anthocyanin pigmentation in the fruits was also eliminated. The AN2-like antagonist is the R3-MYB protein MYBATV. Meanwhile, a similar conclusion regarding MYBATV was made earlier [151]. It can be summarized that HY5 activates AN2-like, promotes the expression of AN1 and MYBATV, and MYBATV protein competes with AN2-like for binding to AN1 and thereby negatively regulates anthocyanin biosynthesis. Moreover, in [152], overexpression of AN2-like was found to increase jasmonic acid accumulation, activate the defense signaling pathway against Botrytis cinerea, and also increase fruit shelf life by inhibiting the expression of genes associated with the modification cell wall.
The previously mentioned dihydroflavonol 4-reductase (DFR) is involved in the reduction of dihydroflavonols to leukoanthocyanidins during the synthesis of the pigments pelargonidin, cyanidin, and delphinidin. The DFR gene in the tomato genome is represented by a single copy, which prompted its use in developing a natural genome editing marker based on homologous recombination with restoration of the DFR function [153]. DFR expression is also regulated by BBX20, which binds to its promoter region to activate expression [154].
6. System of regulation of tomato fruit ripening process
As a result of our literature review, we have presented a putative model of ripening factor regulatory pathways (Figure 4). We recognize five components of the maturation regulation system: transcription factors, hormones, epigenetics, external stimuli, and ncRNAs.
A significant contribution to regulation is provided by the ethylene-dependent pathway involving important polycistronic regulators like ethylene-sensitive genes, ethylene response genes, and the MADS-RIN complex. The fact that there are many additional factors with which RIN directly interacts suggests the existence of various ad hoc regulatory complexes consisting of several units of transcription factors. Quite often, functional redundancy is observed for them. There are 3 possible relationships of transcription factors: redundancy, additivity, and dependency. Redundancy is manifested in the functional identity of transcription factors. Additivity is associated with the provision of function through the joint contribution of each element. Direct dependence involves activation or repression of the role of one factor only after interaction with another. Moreover, autocatalytic regulation of the participants of regulatory cascades is possible.
As practice shows, turning off the function of any of the factors does not interrupt the entire cascade of ripening gene regulation but only leads to a delay phenotype. This is explained by the mechanism of gene compensation, i.e., the existence of homologs of the nonfunctional factor and/or other regulatory pathways. Indeed, we show that there are other regulators of the ethylene pathway, such as factors of the brassinosteroid and abscisic pathways. Although SA/JA pathways are absent in the proposed scheme, we do not exclude the presence of SA/JA regulators as additional maturation factors. Nevertheless, the number of studies aimed at clarifying their role in these processes does not allow us to conclude that they contribute significantly to the regulation of ripening-related genes.
Activation or deactivation of genes involved in ripening regulation can be mediated epigenetically, as discussed previously with specific examples. RIN-mediated regulation also requires interaction with promoters of lncRNAs, which are regulators of other genes, including those associated with maturation. This provides the so-called ethylene-independent regulatory pathway of maturation genes. Because epigenetic regulation and lncRNA regulation are potentially applicable to each element of regulatory cascades, their display on the scheme is redundant.
7. Future Prospects and Challenges
Significant progress has recently been made in understanding signal transduction systems and processes. The discovery of the function of genes and their regulatory systems in tomato ripening process can allow breeding tomatoes with an increased number of fruits and improved nutritional properties.
Although today there are attempts to generalize the cross-relationship of cascades of hormonal signaling pathways [155,156,157], they are only sometimes consistent with each other. They are united by the general concept of the participation of gibberellin- and auxin-sensitive genes at the stages of flowering, fertilization, and early fruit development, and ethylene-sensitive genes at the direct ripening of tomato fruits. Genes of mediator factors ensure the regulation of these processes. Of course, the proposed concepts still need to be completed. Thus, it was shown that ABA and BS are involved in all stages of tomato reproductive processes. It was also demonstrated with RIN and CNR that established hypotheses are not always perfect. The expanding understanding of the influence of epigenetic changes, microRNAs, and external factors on signaling regulation is also worth mentioning.
All this reminds us of a puzzle: to start putting it together, you need to find the edge of the image; otherwise, it can take infinite time to compare individual elements. But signaling pathways do not have “edges”; they must be created manually. One of the modern approaches to solving problems in this area is the use of a wide range of genetic engineering and bioengineering methods, including, among other things, the collection and processing of bioinformatics data, new sequencing technologies, targeted genome editing using CRISPR technology [158,159,160,161,162,163,164,165], use of base and prime editing for precision gene correction [166,167,168], creation of unique and simple markers for detection of transgenes [169], application transcriptomics, metabolomics and other omics technologies.
However, even here, some difficulties arise. For example, orthologous genes often have different regulatory mechanisms and are unreliable predictors of expression in related species. In addition, not all studies are field-validated, which reduces the significance of the data obtained using transgenic plants. Ideal and, at the same time, simplified laboratory conditions cannot determine all the possible subtleties of gene expression and their regulation. Moreover, changes in the expression of regulatory genes do not always entail significant changes in the transcriptome due to the possible presence in the genome of paralogs of the genes being studied. For genes higher in the regulatory cascades, pleiotropic changes occur. It is worth mentioning that the reproduction processes are discrete and appear in various tissues with excellent synchronization, determined by the life cycle stage, so their identification and description are difficult.
Undoubtedly, the existing conclusions about the regulation of plant life cycle processes obtained on model objects such as the tomato, although extensive, still need to be completed. In the future, genetic approaches will continue to make essential contributions to identifying new candidate genes involved in tomato reproductive signaling cascades. This may open up a broader cluster of regulatory signaling networks involving currently unknown factors and stimuli.
As for the perspectives of the practical application of the study of the considered ripening-related genes, these are improving fruit quality due to increased nutrient content, accelerated ripening, prolonged shelf life, and much more.
8. Conclusions
This review examined recent advances in studying tomato ripening factors using various genetic approaches. Modern research into the factors of tomato fruit ripening continues to expand our understanding of the molecular basis and physiological mechanisms of these processes, which has significant applications for improving breeding methods and growing new varieties with enhanced phenotypic traits that meet the requirements of the modern agricultural industry and consumer demand.
The abundance of current scientific reports cited in this review article reflects the relative ease of studying tomato and potential biotechnological improvements and the breadth of approaches and methods. Despite significant advances, a large number of biochemical pathways, the involvement of hundreds of genes in the fruit ripening process, and fine regulation involving transcription factors and microRNAs, at present, it is premature to talk about a complete understanding of the described processes. The complex new data presented in the review and proposed model of gene regulation will allow us to understand the principle of tomato fruit ripening better and complement the overall knowledge picture.
Author Contributions
Conceptualization, D.B. and V.T.; writing—original draft preparation, D.B.; writing—review and editing, V.T. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Russian Science Foundation, grant No. 22-14-00118.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Wang, R.; Tavano, E.C. da R.; Lammers, M.; Martinelli, A.P.; Angenent, G.C.; de Maagd, R.A. Re-Evaluation of Transcription Factor Function in Tomato Fruit Development and Ripening with CRISPR/Cas9-Mutagenesis. Sci. Rep. 2019, 9. [CrossRef]
- Zhao, X.; Yuan, X.; Chen, S.; Meng, L.; Fu, D. Role of the Tomato TAGL1 Gene in Regulating Fruit Metabolites Elucidated Using RNA Sequence and Metabolomics Analyses. PLoS ONE 2018, 13, e0199083. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Yuan, X.; Chen, S.; Fu, D.-Q.; Jiang, C.-Z. Metabolomic and Transcriptomic Analyses Reveal That a MADS-Box Transcription Factor TDR4 Regulates Tomato Fruit Quality. Front. Plant Sci. 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Ito, Y.; Nishizawa-Yokoi, A.; Endo, M.; Mikami, M.; Shima, Y.; Nakamura, N.; Kotake-Nara, E.; Kawasaki, S.; Toki, S. Re-Evaluation of the Rin Mutation and the Role of RIN in the Induction of Tomato Ripening. Nature Plants 2017, 3, 866–874. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Xu, H.; Ju, Z.; Cao, D.; Zhu, H.; Fu, D.; Grierson, D.; Qin, G.; Luo, Y.; Zhu, B. The RIN-MC Fusion of MADS-Box Transcription Factors Has Transcriptional Activity and Modulates Expression of Many Ripening Genes. Plant Physiol. 2017, 176, 891–909. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Zhu, B.; Pirrello, J.; Xu, C.; Zhang, B.; Bouzayen, M.; Chen, K.; Grierson, D. Roles of RIN and Ethylene in Tomato Fruit Ripening and Ripening-associated Traits. New Phytol. 2019, 226, 460–475. [Google Scholar] [CrossRef] [PubMed]
- Upadhyay, R.K.; Tucker, M.L.; Mattoo, A.K. Ethylene and RIPENING INHIBITOR Modulate Expression of SlHSP17.7A, B Class I Small Heat Shock Protein Genes During Tomato Fruit Ripening. Front. Plant Sci. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Ito, Y.; Sekiyama, Y.; Nakayama, H.; Nishizawa-Yokoi, A.; Endo, M.; Shima, Y.; Nakamura, N.; Kotake-Nara, E.; Kawasaki, S.; Hirose, S.; et al. Allelic Mutations in the Ripening-Inhibitor Locus Generate Extensive Variation in Tomato Ripening. Plant Physiol. 2020, 183, 80–95. [Google Scholar] [CrossRef]
- Yu, T.; Tzeng, D.T.W.; Li, R.; Chen, J.; Zhong, S.; Fu, D.; Zhu, B.; Luo, Y.; Zhu, H. Genome-Wide Identification of Long Non-Coding RNA Targets of the Tomato MADS Box Transcription Factor RIN and Function Analysis. Annals of Botany 2018, 123, 469–482. [Google Scholar] [CrossRef]
- Kang, J.; Gong, J.; Zhang, L.; Gao, Z.; Xie, Q.; Hu, Z.; Chen, G. A Novel E6-like Gene, E6-2, Affects Fruit Ripening in Tomato. Plant Science 2021, 313, 111066. [Google Scholar] [CrossRef]
- Osakabe, K.; Wada, N.; Miyaji, T.; Murakami, E.; Marui, K.; Ueta, R.; Hashimoto, R.; Abe-Hara, C.; Kong, B.; Yano, K.; et al. Genome Editing in Plants Using CRISPR Type I-D Nuclease. Commun. Biol. 2020, 3. [Google Scholar] [CrossRef] [PubMed]
- Niu, Q.; Wu, S.; Li, Y.; Yang, X.; Liu, P.; Xu, Y.; Lang, Z. Expanding the Scope of CRISPR/Cas9-mediated Genome Editing in Plants Using an xCas9 and Cas9-NG Hybrid. JIPB 2020, 62, 398–402. [Google Scholar] [CrossRef] [PubMed]
- Niu, Q.; Wu, S.; Xie, H.; Wu, Q.; Liu, P.; Xu, Y.; Lang, Z. Efficient A·T to G·C Base Conversions in Dicots Using Adenine Base Editors Expressed under the Tomato EF1α Promoter. Plant Biotechnol. J. 2022, 21, 5–7. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Wei, W.; Zhao, X.; Tan, X.; Fan, Z.; Zhang, Y.; Jing, Y.; Meng, L.; Zhu, B.; Zhu, H.; et al. A NAC Transcription Factor, NOR-Like1, Is a New Positive Regulator of Tomato Fruit Ripening. Hortic. Res. 2018, 5. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Chen, H.; Nonaka, S.; Sato-Izawa, K.; Kusano, M.; Ezura, H. Ethylene Biosynthesis Controlled by NON-RIPENING: A Regulatory Conflict between Wounding and Ripening. Plant Physiol. Biochem. 2018, 132, 720–726. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Wei, W.; Fan, Z.; Zhao, X.; Zhang, Y.; Jing, Y.; Zhu, B.; Zhu, H.; Shan, W.; Chen, J.; et al. Re-Evaluation of the nor Mutation and the Role of the NAC-NOR Transcription Factor in Tomato Fruit Ripening. J. Exp. Bot. 2020, 71, 3560–3574. [Google Scholar] [CrossRef]
- Yu, Q.; Wang, B.; Li, N.; Tang, Y.; Yang, S.; Yang, T.; Xu, J.; Guo, C.; Yan, P.; Wang, Q.; et al. CRISPR/Cas9-Induced Targeted Mutagenesis and Gene Replacement to Generate Long-Shelf Life Tomato Lines. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef] [PubMed]
- Cao, X.; Wei, C.; Duan, W.; Gao, Y.; Kuang, J.; Liu, M.; Chen, K.; Klee, H.; Zhang, B. Transcriptional and Epigenetic Analysis Reveals That NAC Transcription Factors Regulate Fruit Flavor Ester Biosynthesis. Plant J. 2021, 106, 785–800. [Google Scholar] [CrossRef]
- Jian, W.; Zheng, Y.; Yu, T.; Cao, H.; Chen, Y.; Cui, Q.; Xu, C.; Li, Z. SlNAC6, A NAC Transcription Factor, Is Involved in Drought Stress Response and Reproductive Process in Tomato. J. Plant Physiol. 2021, 264, 153483. [Google Scholar] [CrossRef]
- Gong, J.; Zeng, Y.; Meng, Q.; Guan, Y.; Li, C.; Yang, H.; Zhang, Y.; Ampomah-Dwamena, C.; Liu, P.; Chen, C.; et al. Red Light-Induced Kumquat Fruit Coloration Is Attributable to Increased Carotenoid Metabolism Regulated by FcrNAC22. J. Exp. Bot. 2021, 72, 6274–6290. [Google Scholar] [CrossRef]
- Gao, Y.; Fan, Z.; Zhang, Q.; Li, H.; Liu, G.; Jing, Y.; Zhang, Y.; Zhu, B.; Zhu, H.; Chen, J.; et al. A Tomato NAC Transcription Factor, SlNAM1, Positively Regulates Ethylene Biosynthesis and the Onset of Tomato Fruit Ripening. Plant J. 2021, 108, 1317–1331. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.K.; Vishwakarma, A.; Kenea, H.D.; Galsurker, O.; Cohen, H.; Aharoni, A.; Arazi, T. CRISPR/Cas9 Mutants of Tomato MICRORNA164 Genes Uncover Their Functional Specialization in Development. Plant Physiol. 2021, 187, 1636–1652. [Google Scholar] [CrossRef] [PubMed]
- Lin, D.; Zhu, X.; Qi, B.; Gao, Z.; Tian, P.; Li, Z.; Lin, Z.; Zhang, Y.; Huang, T. SlMIR164A Regulates Fruit Ripening and Quality by Controlling SlNAM2 and SlNAM3 in Tomato. Plant Biotechnol. J. 2022, 20, 1456–1469. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Tang, M.; Huang, Z.; Song, J.; Xu, J.; Ahammed, G.J.; Yu, J.; Zhou, Y. The miR164a-NAM3 Module Confers Cold Tolerance by Inducing Ethylene Production in Tomato. Plant J. 2022, 111, 440–456. [Google Scholar] [CrossRef] [PubMed]
- Yuan, S.; Kawasaki, S.; Abdellatif, I.M.Y.; Nishida, K.; Kondo, A.; Ariizumi, T.; Ezura, H.; Miura, K. Efficient Base Editing in Tomato Using a Highly Expressed Transient System. Plant Cell Rep. 2021, 40, 667–676. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Zhang, Y.; Tureckova, V.; Xue, G.-P.; Fernie, A.R.; Mueller-Roeber, B.; Balazadeh, S. The NAC Transcription Factor SlNAP2 Regulates Leaf Senescence and Fruit Yield in Tomato. Plant Physiol. 2018, 177, 1286–1302. [Google Scholar] [CrossRef]
- Forlani, S.; Cozzi, C.; Rosa, S.; Tadini, L.; Masiero, S.; Mizzotti, C. HEBE, a Novel Positive Regulator of Senescence in Solanum lycopersicum. Sci Rep 2020, 10. [Google Scholar] [CrossRef]
- Gao, Y.; Zhu, N.; Zhu, X.; Wu, M.; Jiang, C.-Z.; Grierson, D.; Luo, Y.; Shen, W.; Zhong, S.; Fu, D.-Q.; et al. Diversity and Redundancy of the Ripening Regulatory Networks Revealed by the fruitENCODE and the New CRISPR/Cas9 CNR and NOR Mutants. Hortic. Res. 2019, 6. [Google Scholar] [CrossRef] [PubMed]
- Jiang, G.; Zeng, J.; Li, Z.; Song, Y.; Yan, H.; He, J.; Jiang, Y.; Duan, X. Redox Regulation of the NOR Transcription Factor Is Involved in the Regulation of Fruit Ripening in Tomato. Plant Physiol. 2020, 183, 671–685. [Google Scholar] [CrossRef]
- Lai, T.; Wang, X.; Ye, B.; Jin, M.; Chen, W.; Wang, Y.; Zhou, Y.; Blanks, A.M.; Gu, M.; Zhang, P.; et al. Molecular and Functional Characterization of the SBP-Box Transcription Factor SPL-CNR in Tomato Fruit Ripening and Cell Death. J. Exp. Bot. 2020, 71, 2995–3011. [Google Scholar] [CrossRef]
- Yin, W.; Hu, Z.; Cui, B.; Guo, X.; Hu, J.; Zhu, Z.; Chen, G. Suppression of the MADS-Box Gene SlMBP8 Accelerates Fruit Ripening of Tomato (Solanum lycopersicum). Plant Physiol. Biochem. 2017, 118, 235–244. [Google Scholar] [CrossRef] [PubMed]
- Yin, W.; Yu, X.; Chen, G.; Tang, B.; Wang, Y.; Liao, C.; Zhang, Y.; Hu, Z. Suppression of SlMBP15 Inhibits Plant Vegetative Growth and Delays Fruit Ripening in Tomato. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Wang, X.; Li, C.; Li, H.; Zhang, J.; Ye, Z. Silencing GRAS2 Reduces Fruit Weight in Tomato. JIPB 2018, 60, 498–513. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Shi, Y.; Su, D.; Lu, W.; Li, Z. SlGRAS4 Accelerates Fruit Ripening by Regulating Ethylene Biosynthesis Genes and SlMADS1 in Tomato. Hortic. Res. 2021, 8. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Peng, S.; Xian, Z.; Lin, D.; Hu, G.; Yang, L.; Ren, M.; Li, Z. Overexpression of a Tomato miR171 Target Gene SlGRAS24 Impacts Multiple Agronomical Traits via Regulating Gibberellin and Auxin Homeostasis. Plant Biotechnol. J. 2016, 15, 472–488. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Li, Y.; Fan, S.; Wen, T.; Wang, M.; Zhang, L.; Zhao, L. The Transcription Factor WRKY32 Affects Tomato Fruit Colour by Regulating YELLOW FRUITED-TOMATO 1, a Core Component of Ethylene Signal Transduction. J. Exp. Bot. 2021, 72, 4269–4282. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Zhang, X.; Wang, L.; Tian, Y.; Jia, N.; Chen, S.; Shi, N.; Huang, X.; Zhou, C.; Yu, Y.; et al. Regulation of Ethylene-Responsive SlWRKYs Involved in Color Change during Tomato Fruit Ripening. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Lang, Z.; Wang, Y.; Tang, K.; Tang, D.; Datsenka, T.; Cheng, J.; Zhang, Y.; Handa, A.K.; Zhu, J.-K. Critical Roles of DNA Demethylation in the Activation of Ripening-Induced Genes and Inhibition of Ripening-Repressed Genes in Tomato Fruit. Proc. Natl. Acad. Sci. USA 2017, 114. [Google Scholar] [CrossRef]
- Li, Z.; Pi, Y.; Fan, J.; Yang, X.; Zhai, C.; Chen, H.; Wang, F.; Ding, J.; Gu, T.; Li, Y.; et al. High Mobility Group A3 Enhances Transcription of the DNA Demethylase Gene SlDML2 to Promote Tomato Fruit Ripening. Plant Physiol. 2022, 189, 315–328. [Google Scholar] [CrossRef]
- Hollwey, E.; Out, S.; Watson, M.R.; Heidmann, I.; Meyer, P. TET3-Mediated Demethylation in Tomato Activates Expression of a CETS Gene That Stimulates Vegetative Growth. Plant Direct 2017, 1. [Google Scholar] [CrossRef]
- Yang, Y.; Tang, K.; Datsenka, T.U.; Liu, W.; Lv, S.; Lang, Z.; Wang, X.; Gao, J.; Wang, W.; Nie, W.; et al. Critical Function of DNA Methyltransferase 1 in Tomato Development and Regulation of the DNA Methylome and Transcriptome. JIPB 2019, 61, 1224–1242. [Google Scholar] [CrossRef] [PubMed]
- Yao, M.; Chen, W.; Kong, J.; Zhang, X.; Shi, N.; Zhong, S.; Ma, P.; Gallusci, P.; Jackson, S.; Liu, Y.; et al. METHYLTRANSFERASE1 and Ripening Modulate Vivipary during Tomato Fruit Development. Plant Physiol. 2020, 183, 1883–1897. [Google Scholar] [CrossRef] [PubMed]
- Wen, Y.X.; Wang, J.Y.; Zhu, H.H.; Han, G.H.; Huang, R.N.; Huang, L.; Hong, Y.G.; Zheng, S.J.; Yang, J.L.; Chen, W.W. Potential Role of Domains Rearranged Methyltransferase7 in Starch and Chlorophyll Metabolism to Regulate Leaf Senescence in Tomato. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef] [PubMed]
- Jia, H.; Jia, H.; Lu, S.; Zhang, Z.; Su, Z.; Sadeghnezhad, E.; Li, T.; Xiao, X.; Wang, M.; Pervaiz, T.; et al. DNA and Histone Methylation Regulates Different Types of Fruit Ripening by Transcriptome and Proteome Analyses. J. Agric. Food Chem. 2022, 70, 3541–3556. [Google Scholar] [CrossRef] [PubMed]
- Corem, S.; Doron-Faigenboim, A.; Jouffroy, O.; Maumus, F.; Arazi, T.; Bouché, N. Redistribution of CHH Methylation and Small Interfering RNAs across the Genome of Tomato Ddm1 Mutants. Plant Cell 2018, 30, 1628–1644. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Zhao, J.; Chen, Z.; Qiao, J.; Zhang, Y.; Shen, H.; Hu, Z. CRISPR/Cas9-Targeted Mutagenesis ofSlCMT4Causes Changes in Plant Architecture and Reproductive Organs in Tomato. Hortic. Res. 2022, 9. [Google Scholar] [CrossRef] [PubMed]
- Hu, G.; Huang, B.; Wang, K.; Frasse, P.; Maza, E.; Djari, A.; Benhamed, M.; Gallusci, P.; Li, Z.; Zouine, M.; et al. Histone Posttranslational Modifications Rather than DNA Methylation Underlie Gene Reprogramming in Pollination-dependent and Pollination-independent Fruit Set in Tomato. New Phytol. 2020, 229, 902–919. [Google Scholar] [CrossRef] [PubMed]
- Bvindi, C.; Tang, L.; Lee, S.; Patrick, R.M.; Yee, Z.R.; Mengiste, T.; Li, Y. Histone Methyltransferases SDG33 and SDG34 Regulate Organ-Specific Nitrogen Responses in Tomato. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef]
- Li, Z.; Jiang, G.; Liu, X.; Ding, X.; Zhang, D.; Wang, X.; Zhou, Y.; Yan, H.; Li, T.; Wu, K.; et al. Histone Demethylase SlJMJ6 Promotes Fruit Ripening by Removing H3K27 Methylation of Ripening-related Genes in Tomato. New Phytol. 2020, 227, 1138–1156. [Google Scholar] [CrossRef]
- Ding, X.; Zhang, D.; Gu, D.; Li, Z.; Liang, H.; Zhu, H.; Jiang, Y.; Duan, X. The Histone H3K27 Demethylase SlJMJ4 Promotes Dark- and ABA-Induced Leaf Senescence in Tomato. Hortic. Res. 2022, 9. [Google Scholar] [CrossRef]
- Yang, X.; Zhang, X.; Yang, Y.; Zhang, H.; Zhu, W.; Nie, W.-F. The Histone Variant Sl_H2A.Z Regulates Carotenoid Biosynthesis and Gene Expression during Tomato Fruit Ripening. Hortic. Res. 2021, 8. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.-E.; Hu, Z.; Li, F.; Zhang, L.; Yu, X.; Tang, B.; Chen, G. Silencing of Histone Deacetylase SlHDT3 Delays Fruit Ripening and Suppresses Carotenoid Accumulation in Tomato. Plant Sci. 2017, 265, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.-E.; Hu, Z.; Zhu, M.; Li, F.; Zhu, Z.; Lu, Y.; Chen, G. The Tomato Histone Deacetylase SlHDA1 Contributes to the Repression of Fruit Ripening and Carotenoid Accumulation. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.-E. Histone Deacetylase Gene SlHDT1 Regulates Tomato Fruit Ripening by Affecting Carotenoid Accumulation and Ethylene Biosynthesis. Plant Sci. 2022, 318, 111235. [Google Scholar] [CrossRef]
- Hawar, A.; Xiong, S.; Yang, Z.; Sun, B. Histone Acetyltransferase SlGCN5 Regulates Shoot Meristem and Flower Development in Solanum lycopersicum. Front. Plant Sci. 2022, 12. [Google Scholar] [CrossRef] [PubMed]
- Bollier, N.; Sicard, A.; Leblond, J.; Latrasse, D.; Gonzalez, N.; Gévaudant, F.; Benhamed, M.; Raynaud, C.; Lenhard, M.; Chevalier, C.; et al. At-MINI ZINC FINGER2 and Sl-INHIBITOR OF MERISTEM ACTIVITY, a Conserved Missing Link in the Regulation of Floral Meristem Termination in Arabidopsis and Tomato. Plant Cell 2018, 30, 83–100. [Google Scholar] [CrossRef] [PubMed]
- Ai, G.; Huang, R.; Zhang, D.; Li, M.; Li, G.; Li, W.; Ahiakpa, J.K.; Wang, Y.; Hong, Z.; Zhang, J. SlGH3.15, a Member of the GH3 Gene Family, Regulates Lateral Root Development and Gravitropism Response by Modulating Auxin Homeostasis in Tomato. Plant Sci. 2023, 330, 111638. [Google Scholar] [CrossRef]
- Sun, M.; Li, H.; Li, Y.; Xiang, H.; Liu, Y.; He, Y.; Qi, M.; Li, T. Tomato YABBY2b Controls Plant Height through Regulating Indole-3-Acetic Acid-Amido Synthetase (GH3.8) Expression. Plant Scie. 2020, 297, 110530. [Google Scholar] [CrossRef]
- Chen, X.; Liao, D.; Yang, X.; Ji, M.; Wang, S.; Gu, M.; Chen, A.; Xu, G. Three Cis-Regulatory Motifs, AuxRE, MYCRS1 and MYCRS2, Are Required for Modulating the Auxin- and Mycorrhiza-Responsive Expression of a Tomato GH3 Gene. Plant Cell Physiol. 2017, 58, 770–778. [Google Scholar] [CrossRef]
- Chen, X.; Chen, J.; Liao, D.; Ye, H.; Li, C.; Luo, Z.; Yan, A.; Zhao, Q.; Xie, K.; Li, Y.; et al. Auxin-mediated Regulation of Arbuscular Mycorrhizal Symbiosis: A Role of SlGH3.4 in Tomato. Plant Cell Environ. 2021, 45, 955–968. [Google Scholar] [CrossRef]
- Sravankumar, T.; Akash; Naik, N.; Kumar, R. A Ripening-Induced SlGH3-2 Gene Regulates Fruit Ripening via Adjusting Auxin-Ethylene Levels in Tomato (Solanum lycopersicum L.). Plant Mol. Biol. 2018, 98, 455–469. [CrossRef] [PubMed]
- Shi, Z.; Jiang, Y.; Han, X.; Liu, X.; Cao, R.; Qi, M.; Xu, T.; Li, T. SlPIN1 Regulates Auxin Efflux to Affect Flower Abscission Process. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef] [PubMed]
- Li, A.; Chen, G.; Yu, X.; Zhu, Z.; Zhang, L.; Zhou, S.; Hu, Z. The Tomato MADS-Box Gene SlMBP9 Negatively Regulates Lateral Root Formation and Apical Dominance by Reducing Auxin Biosynthesis and Transport. Plant Cell Rep. 2019, 38, 951–963. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Zhang, Y.; Feng, Q.; Qin, L.; Pan, C.; Lamin-Samu, A.T.; Lu, G. Tomato AUXIN RESPONSE FACTOR 5 Regulates Fruit Set and Development via the Mediation of Auxin and Gibberellin Signaling. Sci. Rep. 2018, 8. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; Mei, L.; Wu, M.; Wei, W.; Shan, W.; Gong, Z.; Zhang, Q.; Yang, F.; Yan, F.; Zhang, Q.; et al. SlARF10, an Auxin Response Factor, Is Involved in Chlorophyll and Sugar Accumulation during Tomato Fruit Development. J. Exp. Bot. 2018. [Google Scholar] [CrossRef]
- Israeli, A.; Capua, Y.; Shwartz, I.; Tal, L.; Meir, Z.; Levy, M.; Bar, M.; Efroni, I.; Ori, N. Multiple Auxin-Response Regulators Enable Stability and Variability in Leaf Development. Curr. Biol. 2019, 29, 1746–1759.e5. [Google Scholar] [CrossRef] [PubMed]
- Abe-Hara, C.; Yamada, K.; Wada, N.; Ueta, R.; Hashimoto, R.; Osakabe, K.; Osakabe, Y. Effects of the Sliaa9 Mutation on Shoot Elongation Growth of Tomato Cultivars. Front. Plant Sci. 2021, 12. [Google Scholar] [CrossRef]
- Wu, L.; Tian, Z.; Zhang, J. Functional Dissection of Auxin Response Factors in Regulating Tomato Leaf Shape Development. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Damodharan, S.; Corem, S.; Gupta, S.K.; Arazi, T. Tuning of SlARF10A Dosage by sly-miR160a Is Critical for Auxin-mediated Compound Leaf and Flower Development. Plant J. 2018, 96, 855–868. [Google Scholar] [CrossRef]
- Tomlinson, L.; Yang, Y.; Emenecker, R.; Smoker, M.; Taylor, J.; Perkins, S.; Smith, J.; MacLean, D.; Olszewski, N.E.; Jones, J.D.G. Using CRISPR/Cas9 Genome Editing in Tomato to Create a Gibberellin-responsive Dominant Dwarf DELLA Allele. Plant Biotechnol. J. 2018, 17, 132–140. [Google Scholar] [CrossRef]
- Shinozaki, Y.; Ezura, K.; Hu, J.; Okabe, Y.; Bénard, C.; Prodhomme, D.; Gibon, Y.; Sun, T.; Ezura, H.; Ariizumi, T. Identification and Functional Study of a Mild Allele of SlDELLA Gene Conferring the Potential for Improved Yield in Tomato. Sci. Rep. 2018, 8. [Google Scholar] [CrossRef] [PubMed]
- Cheng, W.; Yin, S.; Tu, Y.; Mei, H.; Wang, Y.; Yang, Y. SlCAND1, Encoding Cullin-Associated Nedd8-Dissociated Protein 1, Regulates Plant Height, Flowering Time, Seed Germination, and Root Architecture in Tomato. Plant Mol. Biol. 2020, 102, 537–551. [Google Scholar] [CrossRef] [PubMed]
- Illouz-Eliaz, N.; Ramon, U.; Shohat, H.; Blum, S.; Livne, S.; Mendelson, D.; Weiss, D. Multiple Gibberellin Receptors Contribute to Phenotypic Stability under Changing Environments. Plant Cell 2019, 31, 1506–1519. [Google Scholar] [CrossRef] [PubMed]
- Nir, I.; Shohat, H.; Panizel, I.; Olszewski, N.; Aharoni, A.; Weiss, D. The Tomato DELLA Protein PROCERA Acts in Guard Cells to Promote Stomatal Closure. Plant Cell 2017, 29, 3186–3197. [Google Scholar] [CrossRef] [PubMed]
- Shohat, H.; Illouz-Eliaz, N.; Kanno, Y.; Seo, M.; Weiss, D. The Tomato DELLA Protein PROCERA Promotes Abscisic Acid Responses in Guard Cells by Upregulating an Abscisic Acid Transporter. Plant Physiol. 2020, 184, 518–528. [Google Scholar] [CrossRef] [PubMed]
- Silva, G.F.F.; Silva, E.M.; Correa, J.P.O.; Vicente, M.H.; Jiang, N.; Notini, M.M.; Junior, A.C.; De Jesus, F.A.; Castilho, P.; Carrera, E.; et al. Tomato Floral Induction and Flower Development Are Orchestrated by the Interplay between Gibberellin and Two Unrelated microRNA-controlled Modules. New Phytol. 2018, 221, 1328–1344. [Google Scholar] [CrossRef]
- Naeem, M.; Waseem, M.; Zhu, Z.; Zhang, L. Downregulation of SlGRAS15 Manipulates Plant Architecture in Tomato (Solanum lycopersicum). Dev. Genes. Evol. 2019, 230, 1–12. [Google Scholar] [CrossRef]
- Zhu, Z.; Liang, H.; Chen, G.; Li, F.; Wang, Y.; Liao, C.; Hu, Z. The bHLH Transcription Factor SlPRE2 Regulates Tomato Fruit Development and Modulates Plant Response to Gibberellin. Plant Cell Rep. 2019, 38, 1053–1064. [Google Scholar] [CrossRef]
- Zhu, Z.; Chen, G.; Guo, X.; Yin, W.; Yu, X.; Hu, J.; Hu, Z. Overexpression of SlPRE2, an Atypical bHLH Transcription Factor, Affects Plant Morphology and Fruit Pigment Accumulation in Tomato. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Li, J.; Gong, J.; Zhang, L.; Shen, H.; Chen, G.; Xie, Q.; Hu, Z. Overexpression of SlPRE5, an Atypical bHLH Transcription Factor, Affects Plant Morphology and Chlorophyll Accumulation in Tomato. Journal of Plant Physiol. 2022, 273, 153698. [Google Scholar] [CrossRef]
- Glanz-Idan, N.; Lach, M.; Tarkowski, P.; Vrobel, O.; Wolf, S. Delayed Leaf Senescence by Upregulation of Cytokinin Biosynthesis Specifically in Tomato Roots. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef]
- Gan, L.; Song, M.; Wang, X.; Yang, N.; Li, H.; Liu, X.; Li, Y. Cytokinins Are Involved in Regulation of Tomato Pericarp Thickness and Fruit Size. Hortic. Res. 2022, 9. [Google Scholar] [CrossRef] [PubMed]
- Jia, M.; Du, P.; Ding, N.; Zhang, Q.; Xing, S.; Wei, L.; Zhao, Y.; Mao, W.; Li, J.; Li, B.; et al. Two FERONIA-Like Receptor Kinases Regulate Apple Fruit Ripening by Modulating Ethylene Production. Front. Plant Sci. 2017, 8. [Google Scholar] [CrossRef]
- Yanping, Z.; Yuqing, H.; Chen, W.; Qian, M.; Songtao, J.; Xudong, Z.; Ting, Z.; Kekun, Z.; Haifeng, J.; Tariq, P.; et al. Characterization and Identification of PpEIN3 during the Modulation of Fruit Ripening Process by Ectopic Expressions in Tomato. Plant Genome 2019, 12. [Google Scholar] [CrossRef]
- Wang, H.; Zhang, H.; Liang, F.; Cong, L.; Song, L.; Li, X.; Zhai, R.; Yang, C.; Wang, Z.; Ma, F.; et al. PbEIL1 Acts Upstream ofPbCysp1to Regulate Ovule Senescence in Seedless Pear. Hortic. Res. 2021, 8. [Google Scholar] [CrossRef]
- Althiab-Almasaud, R.; Chen, Y.; Maza, E.; Djari, A.; Frasse, P.; Mollet, J.; Mazars, C.; Jamet, E.; Chervin, C. Ethylene Signaling Modulates Tomato Pollen Tube Growth through Modifications of Cell Wall Remodeling and Calcium Gradient. Plant J. 2021, 107, 893–908. [Google Scholar] [CrossRef]
- Chen, Y.; Hu, G.; Rodriguez, C.; Liu, M.; Binder, B.M.; Chervin, C. Roles of SlETR7, a Newly Discovered Ethylene Receptor, in Tomato Plant and Fruit Development. Hortic. Res. 2020, 7. [Google Scholar] [CrossRef]
- Shimatani, Z.; Kashojiya, S.; Takayama, M.; Terada, R.; Arazoe, T.; Ishii, H.; Teramura, H.; Yamamoto, T.; Komatsu, H.; Miura, K.; et al. Targeted Base Editing in Rice and Tomato Using a CRISPR-Cas9 Cytidine Deaminase Fusion. Nat. Biotechnol. 2017, 35, 441–443. [Google Scholar] [CrossRef]
- Kashojiya, S.; Lu, Y.; Takayama, M.; Komatsu, H.; Minh, L.H.T.; Nishida, K.; Shirasawa, K.; Miura, K.; Nonaka, S.; Masuda, J.; et al. Modification of Tomato Breeding Traits and Plant Hormone Signaling by Target-AID, the Genome-Editing System Inducing Efficient Nucleotide Substitution. Hortic. Res. 2022, 9. [Google Scholar] [CrossRef]
- Guo, H.; Mao, M.; Deng, Y.; Sun, L.; Chen, R.; Cao, P.; Lai, J.; Zhang, Y.; Wang, C.; Li, C.; et al. Multi-Omics Analysis Reveals That SlERF.D6 Synergistically Regulates SGAs and Fruit Development. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef]
- Gambhir, P.; Singh, V.; Parida, A.; Raghuvanshi, U.; Kumar, R.; Sharma, A.K. Ethylene Response Factor ERF.D7 Activates Auxin Response Factor 2 Paralogs to Regulate Tomato Fruit Ripening. Plant Physiol. 2022, 190, 2775–2796. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Gao, M.; Zhao, Y.; Chen, Y.; Wu, L.; Yin, H.; Yang, J.; Xiong, S.; Wang, S.; Wang, J.; et al. LcERF19, an AP2/ERF Transcription Factor from Litsea Cubeba, Positively Regulates Geranial and Neral Biosynthesis. Hortic. Res. 2022, 9. [Google Scholar] [CrossRef]
- Li, G.; Wang, J.; Zhang, C.; Ai, G.; Zhang, D.; Wei, J.; Cai, L.; Li, C.; Zhu, W.; Larkin, R.M.; et al. L2, a Chloroplast Metalloproteinase, Regulates Fruit Ripening by Participating in Ethylene Autocatalysis under the Control of Ethylene Response Factors. J. Exp. Bot. 2021, 72, 7035–7048. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Feng, P.; Tang, B.; Hu, Z.; Xie, Q.; Zhou, S.; Chen, G. The AP2/ERF Transcription Factor SlERF.F5 Functions in Leaf Senescence in Tomato. Plant Cell Rep. 2022, 41, 1181–1195. [Google Scholar] [CrossRef]
- Hu, C.; Wu, S.; Li, J.; Dong, H.; Zhu, C.; Sun, T.; Hu, Z.; Foyer, C.H.; Yu, J. Herbivore-induced Ca2+ Signals Trigger a Jasmonate Burst by Activating ERF16-mediated Expression in Tomato. New Phytol. 2022, 236, 1796–1808. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Hu, K.; Wei, S.; Yao, G.; Zhang, H. ETHYLENE RESPONSE FACTORS 4.1/4.2 with an EAR Motif Repress Anthocyanin Biosynthesis in Red-Skinned Pears. Plant Physiol. 2023, 192, 1892–1912. [Google Scholar] [CrossRef] [PubMed]
- Liu, A.; Cheng, C. Pathogen-induced ERF68 Regulates Hypersensitive Cell Death in Tomato. Molecular Plant Pathology 2016, 18, 1062–1074. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Yuan, L.; Guo, W.; Wu, W. Transcription Factor TERF1 Promotes Seed Germination under Osmotic Conditions by Activating Gibberellin Acid Signaling. Plant Sci. 2022, 322, 111350. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Yang, H.; Tang, B.; Li, F.; Xie, Q.; Chen, G.; Hu, Z. The AP2/ERF Transcription Factor SlERF.J2 Functions in Hypocotyl Elongation and Plant Height in Tomato. Plant Cell Rep. 2022. [Google Scholar] [CrossRef]
- Chen, Y.; Cai, X.; Tang, B.; Xie, Q.; Chen, G.; Chen, X.; Hu, Z. SlERF.J2 Reduces Chlorophyll Accumulation and Inhibits Chloroplast Biogenesis and Development in Tomato Leaves. Plant Sci. 2023, 328, 111578. [Google Scholar] [CrossRef]
- Gupta, A.; Upadhyay, R.K.; Prabhakar, R.; Tiwari, N.; Garg, R.; Sane, V.A.; Sane, A.P. SlDREB3, a Negative Regulator of ABA Responses, Controls Seed Germination, Fruit Size and the Onset of Ripening in Tomato. Plant Sci. 2022, 319, 111249. [Google Scholar] [CrossRef] [PubMed]
- Fang, P.; Wang, Y.; Wang, M.; Wang, F.; Chi, C.; Zhou, Y.; Zhou, J.; Shi, K.; Xia, X.; Foyer, C.H.; et al. Crosstalk between Brassinosteroid and Redox Signaling Contributes to the Activation of CBF Expression during Cold Responses in Tomato. Antioxidants 2021, 10, 509. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Cao, J.-J.; Wang, K.-X.; Xia, X.-J.; Shi, K.; Zhou, Y.-H.; Yu, J.-Q.; Zhou, J. BZR1 Mediates Brassinosteroid-Induced Autophagy and Nitrogen Starvation in Tomato. Plant Physiol. 2018, 179, 671–685. [Google Scholar] [CrossRef] [PubMed]
- Sang, K.; Li, J.; Qian, X.; Yu, J.; Zhou, Y.; Xia, X. The APETALA2a/DWARF/BRASSINAZOLE-RESISTANT 1 Module Contributes to Carotenoid Synthesis in Tomato Fruits. Plant J. 2022, 112, 1238–1251. [Google Scholar] [CrossRef]
- Hu, S.; Liu, L.; Li, S.; Shao, Z.; Meng, F.; Liu, H.; Duan, W.; Liang, D.; Zhu, C.; Xu, T.; et al. Regulation of Fruit Ripening by the Brassinosteroid Biosynthetic Gene SlCYP90B3 via an Ethylene-Dependent Pathway in Tomato. Hortic. Res. 2020, 7. [Google Scholar] [CrossRef] [PubMed]
- Nie, S.; Huang, S.; Wang, S.; Cheng, D.; Liu, J.; Lv, S.; Li, Q.; Wang, X. Enhancing Brassinosteroid Signaling via Overexpression of Tomato (Solanum lycopersicum) SlBRI1 Improves Major Agronomic Traits. Front. Plant Sci. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Liu, J.; Zhao, T.; Du, C.; Nie, S.; Zhang, Y.; Lv, S.; Huang, S.; Wang, X. Modification of Threonine-1050 of SlBRI1 Regulates BR Signalling and Increases Fruit Yield of Tomato. BMC Plant Biol. 2019, 19. [Google Scholar] [CrossRef]
- Wang, D.; Yang, Z.; Wu, M.; Wang, W.; Wang, Y.; Nie, S. Enhanced Brassinosteroid Signaling via the Overexpression of SlBRI1 Positively Regulates the Chilling Stress Tolerance of Tomato. Plant Sci. 2022, 320, 111281. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.; Luo, B.; Hu, M.; Fu, S.; Liu, J.; Jiang, M.; Zhao, Y.; Huang, S.; Wang, S.; Wang, X. Brassinosteroid Signaling Downstream Suppressor BIN2 Interacts with SLFRIGIDA-LIKE to Induce Early Flowering in Tomato. IJMS 2022, 23, 11264. [Google Scholar] [CrossRef]
- Lee, J.; Han, S.; Lee, H.-Y.; Jeong, B.; Heo, T.-Y.; Hyun, T.K.; Kim, K.; Je, B.I.; Lee, H.; Shim, D.; et al. Brassinosteroids Facilitate Xylem Differentiation and Wood Formation in Tomato. Planta 2019, 249, 1391–1403. [Google Scholar] [CrossRef]
- Mori, K.; Lemaire-Chamley, M.; Jorly, J.; Carrari, F.; Conte, M.; Asamizu, E.; Mizoguchi, T.; Ezura, H.; Rothan, C. The Conserved Brassinosteroid-Related Transcription Factor BIM1a Negatively Regulates Fruit Growth in Tomato. J. Exp. Bot. 2020, 72, 1181–1197. [Google Scholar] [CrossRef] [PubMed]
- Yu, T.; Ai, G.; Xie, Q.; Wang, W.; Song, J.; Wang, J.; Tao, J.; Zhang, X.; Hong, Z.; Lu, Y.; et al. Regulation of Tomato Fruit Elongation by Transcription Factor BZR1.7 through Promotion of SUN Gene Expression. Hortic. Res. 2022, 9. [Google Scholar] [CrossRef] [PubMed]
- Xia, X.; Dong, H.; Yin, Y.; Song, X.; Gu, X.; Sang, K.; Zhou, J.; Shi, K.; Zhou, Y.; Foyer, C.H.; et al. Brassinosteroid Signaling Integrates Multiple Pathways to Release Apical Dominance in Tomato. Proc. Natl. Acad. Sci. U.S.A. 2021, 118. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Liu, L.; Liang, D.; Zhang, M.; Jia, C.; Qi, M.; Liu, Y.; Shao, Z.; Meng, F.; Hu, S.; et al. SlBES1 Promotes Tomato Fruit Softening through Transcriptional Inhibition of PMEU1. iScience 2021, 24, 102926. [Google Scholar] [CrossRef] [PubMed]
- Sierra-Orozco, E.; Shekasteband, R.; Illa-Berenguer, E.; Snouffer, A.; van der Knaap, E.; Lee, T.G.; Hutton, S.F. Identification and Characterization of GLOBE, a Major Gene Controlling Fruit Shape and Impacting Fruit Size and Marketability in Tomato. Hortic Res 2021, 8. [Google Scholar] [CrossRef] [PubMed]
- Hou, J.; Sun, Q.; Li, J.; Ahammed, G.J.; Yu, J.; Fang, H.; Xia, X. Glutaredoxin S25 and Its Interacting TGACG Motif-Binding Factor TGA2 Mediate Brassinosteroid-Induced Chlorothalonil Metabolism in Tomato Plants. Environ. Pollut. 2019, 255, 113256. [Google Scholar] [CrossRef] [PubMed]
- Hou, J.; Zhang, Q.; Zhou, Y.; Ahammed, G.J.; Zhou, Y.; Yu, J.; Fang, H.; Xia, X. Glutaredoxin GRXS16 Mediates Brassinosteroid-Induced Apoplastic H2O2 Production to Promote Pesticide Metabolism in Tomato. Environ. Pollut. 2018, 240, 227–234. [Google Scholar] [CrossRef] [PubMed]
- An, S.; Liu, Y.; Sang, K.; Wang, T.; Yu, J.; Zhou, Y.; Xia, X. Brassinosteroid Signaling Positively Regulates Abscisic Acid Biosynthesis in Response to Chilling Stress in Tomato. JIPB 2023, 65, 10–24. [Google Scholar] [CrossRef]
- Liang, B.; Sun, Y.; Wang, J.; Zheng, Y.; Zhang, W.; Xu, Y.; Li, Q.; Leng, P. Tomato Protein Phosphatase 2C Influences the Onset of Fruit Ripening and Fruit Glossiness. J. Exp. Bot. 2020, 72, 2403–2418. [Google Scholar] [CrossRef]
- Wang, J.; Xu, Y.; Zhang, W.; Zheng, Y.; Yuan, B.; Li, Q.; Leng, P. Tomato SlPP2C5 Is Involved in the Regulation of Fruit Development and Ripening. Plant Cell Physiol. 2021, 62, 1760–1769. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, Q.; Jiang, L.; Kai, W.; Liang, B.; Wang, J.; Du, Y.; Zhai, X.; Wang, J.; Zhang, Y.; et al. Suppressing Type 2C Protein Phosphatases Alters Fruit Ripening and the Stress Response in Tomato. Plant Cell Physiol. 2017, 59, 142–154. [Google Scholar] [CrossRef]
- Sun, Y.; Ji, K.; Liang, B.; Du, Y.; Jiang, L.; Wang, J.; Kai, W.; Zhang, Y.; Zhai, X.; Chen, P.; et al. Suppressing ABA Uridine Diphosphate Glucosyltransferase (SlUGT75C1) Alters Fruit Ripening and the Stress Response in Tomato. Plant J. 2017, 91, 574–589. [Google Scholar] [CrossRef] [PubMed]
- Kai, W.; Wang, J.; Liang, B.; Fu, Y.; Zheng, Y.; Zhang, W.; Li, Q.; Leng, P. PYL9 Is Involved in the Regulation of ABA Signaling during Tomato Fruit Ripening. J. Exp. Bot. 2019, 70, 6305–6319. [Google Scholar] [CrossRef]
- Lei, L.; Zhang, J.; Pu, D.; Liu, B.; Meng, X.; Shang, Q.; Duan, Y.; Zhang, F.; Zhang, M.; Dong, C. ABA-responsive AREB1/ABI3-1/ABI5 Cascade Regulates IAA Oxidase Gene SlDAO2 to Inhibit Hypocotyl Elongation in Tomato. Plant Cell Environ. 2022, 46, 498–517. [Google Scholar] [CrossRef]
- Song, J.; Shang, L.; Wang, X.; Xing, Y.; Xu, W.; Zhang, Y.; Wang, T.; Li, H.; Zhang, J.; Ye, Z. MAPK11 Regulates Seed Germination and ABA Signaling in Tomato by Phosphorylating SnRKs. J. Exp. Bot. 2020, 72, 1677–1690. [Google Scholar] [CrossRef]
- Wang, W.; Wang, X.; Wang, Y.; Zhou, G.; Wang, C.; Hussain, S.; Adnan; Lin, R.; Wang, T.; Wang, S. SlEAD1, an EAR Motif-Containing ABA down-Regulated Novel Transcription Repressor Regulates ABA Response in Tomato. GM Crops Food 2020, 11, 275–289. [CrossRef]
- Hu, K.-D.; Zhang, X.-Y.; Yao, G.-F.; Rong, Y.-L.; Ding, C.; Tang, J.; Yang, F.; Huang, Z.-Q.; Xu, Z.-M.; Chen, X.-Y.; et al. A Nuclear-Localized Cysteine Desulfhydrase Plays a Role in Fruit Ripening in Tomato. Hortic. Res. 2020, 7. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.-Q.; Hu, K.-D.; Yao, G.-F.; Wang, S.-Y.; Peng, X.-J.; Zhang, H. A D-Cysteine Desulfhydrase, SlDCD2, Participates in Tomato Fruit Ripening by Modulating ROS Homoeostasis and Ethylene Biosynthesis. Hortic. Res. 2023, 10. [Google Scholar] [CrossRef] [PubMed]
- Sun, C.; Yao, G.; Li, L.; Li, T.; Zhao, Y.; Hu, K.; Zhang, C.; Zhang, H. E3 Ligase BRG3 Persulfidation Delays Tomato Ripening by Reducing Ubiquitination of the Repressor WRKY71. Plant Physiol. 2023, 192, 616–632. [Google Scholar] [CrossRef]
- Hu, K.; Peng, X.; Yao, G.; Zhou, Z.; Yang, F.; Li, W.; Zhao, Y.; Li, Y.; Han, Z.; Chen, X.; et al. Roles of a Cysteine Desulfhydrase LCD1 in Regulating Leaf Senescence in Tomato. IJMS 2021, 22, 13078. [Google Scholar] [CrossRef]
- Ernesto Bianchetti, R.; Silvestre Lira, B.; Santos Monteiro, S.; Demarco, D.; Purgatto, E.; Rothan, C.; Rossi, M.; Freschi, L. Fruit-Localized Phytochromes Regulate Plastid Biogenesis, Starch Synthesis, and Carotenoid Metabolism in Tomato. J. Exp. Bot. 2018, 69, 3573–3586. [Google Scholar] [CrossRef]
- Naeem, M.; Muqarab, R.; Waseem, M. The Solanum Melongena COP1 Delays Fruit Ripening and Influences Ethylene Signaling in Tomato. Journal of Plant Physiol. 2019, 240, 152997. [Google Scholar] [CrossRef] [PubMed]
- Hunziker, J.; Nishida, K.; Kondo, A.; Kishimoto, S.; Ariizumi, T.; Ezura, H. Multiple Gene Substitution by Target-AID Base-Editing Technology in Tomato. Sci. Rep. 2020, 10. [Google Scholar] [CrossRef] [PubMed]
- Gramegna, G.; Rosado, D.; Sánchez Carranza, A.P.; Cruz, A.B.; Simon-Moya, M.; Llorente, B.; Rodríguez-Concepcíon, M.; Freschi, L.; Rossi, M. PHYTOCHROME-INTERACTING FACTOR 3 Mediates Light-dependent Induction of Tocopherol Biosynthesis during Tomato Fruit Ripening. Plant Cell Environ. 2018, 42, 1328–1339. [Google Scholar] [CrossRef] [PubMed]
- Rosado, D.; Trench, B.; Bianchetti, R.; Zuccarelli, R.; Rodrigues Alves, F.R.; Purgatto, E.; Segal Floh, E.I.; Silveira Nogueira, F.T.; Freschi, L.; Rossi, M. Downregulation of PHYTOCHROME-INTERACTING FACTOR 4 Influences Plant Development and Fruit Production. Plant Physiol. 2019, 181, 1360–1370. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Liu, Y.; Ali, M.; Ye, L.; Pan, C.; Li, M.; Zhao, X.; Yu, F.; Zhao, X.; Lu, G. Phytochrome Interacting Factor 3 Regulates Pollen Mitotic Division through Auxin Signalling and Sugar Metabolism Pathways in Tomato. New Phytol. 2021, 234, 560–577. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Meng, L.; Gao, Y.; Grierson, D.; Fu, D. Manipulation of Light Signal Transduction Factors as a Means of Modifying Steroidal Glycoalkaloids Accumulation in Tomato Leaves. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Wang, X.; Zhang, Y.; Yan, J.; Ahammed, G.J.; Bu, X.; Sun, X.; Liu, Y.; Xu, T.; Qi, H.; et al. SlFHY3 and SlHY5 Act Compliantly to Enhance Cold Tolerance through the Integration of Myo-inositol and Light Signaling in Tomato. New Phytol. 2022, 233, 2127–2143. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Zhang, C.; Dong, H.; Liu, X.; Guo, H.; Tong, B.; Fang, F.; Zhao, Y.; Yu, Y.; Liu, Y.; et al. Activation and Negative Feedback Regulation ofSlHY5Transcription by the SlBBX20/21–SlHY5 Transcription Factor Module in UV-B Signaling. Plant Cell 2022, 34, 2038–2055. [Google Scholar] [CrossRef]
- Qiu, Z.; Wang, H.; Li, D.; Yu, B.; Hui, Q.; Yan, S.; Huang, Z.; Cui, X.; Cao, B. Identification of Candidate HY5-Dependent and -Independent Regulators of Anthocyanin Biosynthesis in Tomato. Plant Cell Physiol. 2018, 60, 643–656. [Google Scholar] [CrossRef]
- Balderrama, D.; Barnwell, S.; Carlson, K.D.; Salido, E.; Guevara, R.; Nguyen, C.; Madlung, A. Phytochrome F Mediates Red Light Responsiveness Additively with Phytochromes B1 and B2 in Tomato. Plant Physiol. 2023, 191, 2353–2366. [Google Scholar] [CrossRef]
- Liu, C.; Chi, C.; Jin, L.; Zhu, J.; Yu, J.; Zhou, Y. The bZip Transcription Factor HY5 Mediates CRY1a-induced Anthocyanin Biosynthesis in Tomato. Plant Cell Environ. 2018, 41, 1762–1775. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Hu, C.; Liu, C.; Wang, J.; Zhou, Y.; Yu, J. ELONGATED HYPOCOTYL 5 Mediates Blue Light-Induced Starch Degradation in Tomato. J. Exp. Bot. 2020, 72, 2627–2641. [Google Scholar] [CrossRef]
- Zhi, J.; Liu, X.; Li, D.; Huang, Y.; Yan, S.; Cao, B.; Qiu, Z. CRISPR/Cas9-Mediated SlAN2 Mutants Reveal Various Regulatory Models of Anthocyanin Biosynthesis in Tomato Plant. Plant Cell Rep. 2020, 39, 799–809. [Google Scholar] [CrossRef] [PubMed]
- Jian, W.; Cao, H.; Yuan, S.; Liu, Y.; Lu, J.; Lu, W.; Li, N.; Wang, J.; Zou, J.; Tang, N.; et al. SlMYB75, an MYB-Type Transcription Factor, Promotes Anthocyanin Accumulation and Enhances Volatile Aroma Production in Tomato Fruits. Hortic. Res. 2019, 6. [Google Scholar] [CrossRef] [PubMed]
- Heo, J.; Bang, W.Y.; Jeong, J.C.; Park, S.-C.; Lee, J.M.; Choi, S.; Lee, B.; Lee, Y.K.; Kim, K.; Park, S.J. The Comparisons of Expression Pattern Reveal Molecular Regulation of Fruit Metabolites in S. nigrum and S. lycopersicum. Sci. Rep. 2022, 12. [Google Scholar] [CrossRef]
- Cerqueira, J.V.A.; Zhu, F.; Mendes, K.; Nunes-Nesi, A.; Martins, S.C.V.; Benedito, V.; Fernie, A.R.; Zsogon, A. Promoter Replacement of ANT1 Induces Anthocyanin Accumulation and Triggers the Shade Avoidance Response through Developmental, Physiological and Metabolic Reprogramming in Tomato. Hortic. Res. 2022, 10. [Google Scholar] [CrossRef] [PubMed]
- Deng, L.; Wang, H.; Sun, C.; Li, Q.; Jiang, H.; Du, M.; Li, C.-B.; Li, C. Efficient Generation of Pink-Fruited Tomatoes Using CRISPR/Cas9 System. J. Genet. Genomics 2018, 45, 51–54. [Google Scholar] [CrossRef]
- Sun, C.; Deng, L.; Du, M.; Zhao, J.; Chen, Q.; Huang, T.; Jiang, H.; Li, C.-B.; Li, C. A Transcriptional Network Promotes Anthocyanin Biosynthesis in Tomato Flesh. Mol. Plant 2020, 13, 42–58. [Google Scholar] [CrossRef]
- Yan, S.; Chen, N.; Huang, Z.; Li, D.; Zhi, J.; Yu, B.; Liu, X.; Cao, B.; Qiu, Z. Anthocyanin Fruit Encodes an R2R3-MYB Transcription Factor, SlAN2-like, Activating the Transcription of SlMYBATV to Fine-tune Anthocyanin Content in Tomato Fruit. New Phytol. 2019, 225, 2048–2063. [Google Scholar] [CrossRef]
- Colanero, S.; Perata, P.; Gonzali, S. The Atroviolacea Gene Encodes an R3-MYB Protein Repressing Anthocyanin Synthesis in Tomato Plants. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Zhang, Z.; Xu, Z.; Wang, L.; Chen, C.; Ren, Z. Overexpression of SlMYB75 Enhances Resistance to Botrytis Cinerea and Prolongs Fruit Storage Life in Tomato. Plant Cell Rep. 2020, 40, 43–58. [Google Scholar] [CrossRef] [PubMed]
- Danilo, B.; Perrot, L.; Botton, E.; Nogué, F.; Mazier, M. The DFR Locus: A Smart Landing Pad for Targeted Transgene Insertion in Tomato. PLoS ONE 2018, 13, e0208395. [Google Scholar] [CrossRef] [PubMed]
- Luo, D.; Xiong, C.; Lin, A.; Zhang, C.; Sun, W.; Zhang, J.; Yang, C.; Lu, Y.; Li, H.; Ye, Z.; et al. SlBBX20 Interacts with the COP9 Signalosome Subunit SlCSN5-2 to Regulate Anthocyanin Biosynthesis by Activating SlDFR Expression in Tomato. Hortic. Res. 2021, 8. [Google Scholar] [CrossRef] [PubMed]
- Quinet, M.; Angosto, T.; Yuste-Lisbona, F.J.; Blanchard-Gros, R.; Bigot, S.; Martinez, J.-P.; Lutts, S. Tomato Fruit Development and Metabolism. Front. Plant Sci. 2019, 10. [Google Scholar] [CrossRef]
- Li, S.; Chen, K.; Grierson, D. Molecular and Hormonal Mechanisms Regulating Fleshy Fruit Ripening. Cells 2021, 10, 1136. [Google Scholar] [CrossRef] [PubMed]
- Fenn, M.A.; Giovannoni, J.J. Phytohormones in Fruit Development and Maturation. Plant J. 2021, 105, 446–458. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Andersson, M.; Granell, A.; Cardi, T.; Hofvander, P.; Nicolia, A. Establishment of a DNA-Free Genome Editing and Protoplast Regeneration Method in Cultivated Tomato (Solanum lycopersicum). Plant Cell Rep. 2022, 41, 1843–1852. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Wang, X.; Li, Q.; Peng, W.; Zhang, Z.; Chu, P.; Guo, S.; Fan, Y.; Lyu, S. AtGCS Promoter-Driven Clustered Regularly Interspaced Short Palindromic Repeats/Cas9 Highly Efficiently Generates Homozygous/Biallelic Mutations in the Transformed Roots by Agrobacterium rhizogenes–Mediated Transformation. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef]
- Zhang, N.; Roberts, H.M.; Van Eck, J.; Martin, G.B. Generation and Molecular Characterization of CRISPR/Cas9-Induced Mutations in 63 Immunity-Associated Genes in Tomato Reveals Specificity and a Range of Gene Modifications. Front. Plant Sci. 2020, 11. [Google Scholar] [CrossRef]
- Vu, T.V.; Doan, D.T.H.; Tran, M.T.; Sung, Y.W.; Song, Y.J.; Kim, J.-Y. Improvement of the LbCas12a-crRNA System for Efficient Gene Targeting in Tomato. Front. Plant Sci. 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Kirke, J.; Kaplan, N.; Velez, S.; Jin, X.-L.; Vichyavichien, P.; Zhang, X.-H. Tissue-Preferential Activity and Induction of the Pepper Capsaicin Synthase PUN1 Promoter by Wounding, Heat and Metabolic Pathway Precursor in Tobacco and Tomato Plants. Mol. Biotechnol. 2018, 60, 194–202. [Google Scholar] [CrossRef] [PubMed]
- Vu, T.V.; Sivankalyani, V.; Kim, E.; Doan, D.T.H.; Tran, M.T.; Kim, J.; Sung, Y.W.; Park, M.; Kang, Y.J.; Kim, J. Highly Efficient Homology-directed Repair Using CRISPR/Cpf1-geminiviral Replicon in Tomato. Plant Biotechnol. J. 2020, 18, 2133–2143. [Google Scholar] [CrossRef] [PubMed]
- de Maagd, R.A.; Loonen, A.; Chouaref, J.; Pele, A.; Meijer-Dekens, F.; Fransz, P.; Bai, Y. CRISPR/Cas Inactivation of RECQ4 Increases Homeologous Crossovers in an Interspecific Tomato Hybrid. Plant Biotechnol. J. 2019, 18, 805–813. [Google Scholar] [CrossRef] [PubMed]
- Wada, N.; Osakabe, K.; Osakabe, Y. Type I-D CRISPR System-Mediated Genome Editing in Plants. Methods Mol. Biol. 2023, 21–38. [Google Scholar] [CrossRef]
- Veillet, F.; Perrot, L.; Guyon-Debast, A.; Kermarrec, M.-P.; Chauvin, L.; Chauvin, J.-E.; Gallois, J.-L.; Mazier, M.; Nogue, F. Expanding the CRISPR Toolbox in P. Patens Using SpCas9-NG Variant and Application for Gene and Base Editing in Solanaceae Crops. IJMS 2020, 21, 1024. [Google Scholar] [CrossRef] [PubMed]
- Shimatani, Z.; Kashojiya, S.; Takayama, M.; Terada, R.; Arazoe, T.; Ishii, H.; Teramura, H.; Yamamoto, T.; Komatsu, H.; Miura, K.; et al. Targeted Base Editing in Rice and Tomato Using a CRISPR-Cas9 Cytidine Deaminase Fusion. Nat. Biotechnol. 2017, 35, 441–443. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; He, Y.; Sretenovic, S.; Liu, S.; Cheng, Y.; Han, Y.; Liu, G.; Bao, Y.; Fang, Q.; Zheng, X.; et al. CRISPR-BETS: A Base-editing Design Tool for Generating Stop Codons. Plant Biotechnol. J. 2021, 20, 499–510. [Google Scholar] [CrossRef]
- Matsumoto, A.; Schlüter, T.; Melkonian, K.; Takeda, A.; Nakagami, H.; Mine, A. A Versatile Tn7 Transposon-Based Bioluminescence Tagging Tool for Quantitative and Spatial Detection of Bacteria in Plants. Plant Commun. 2022, 3, 100227. [Google Scholar] [CrossRef]
Figure 1.
CRISPR/Cas9-related studies on tomato by year.
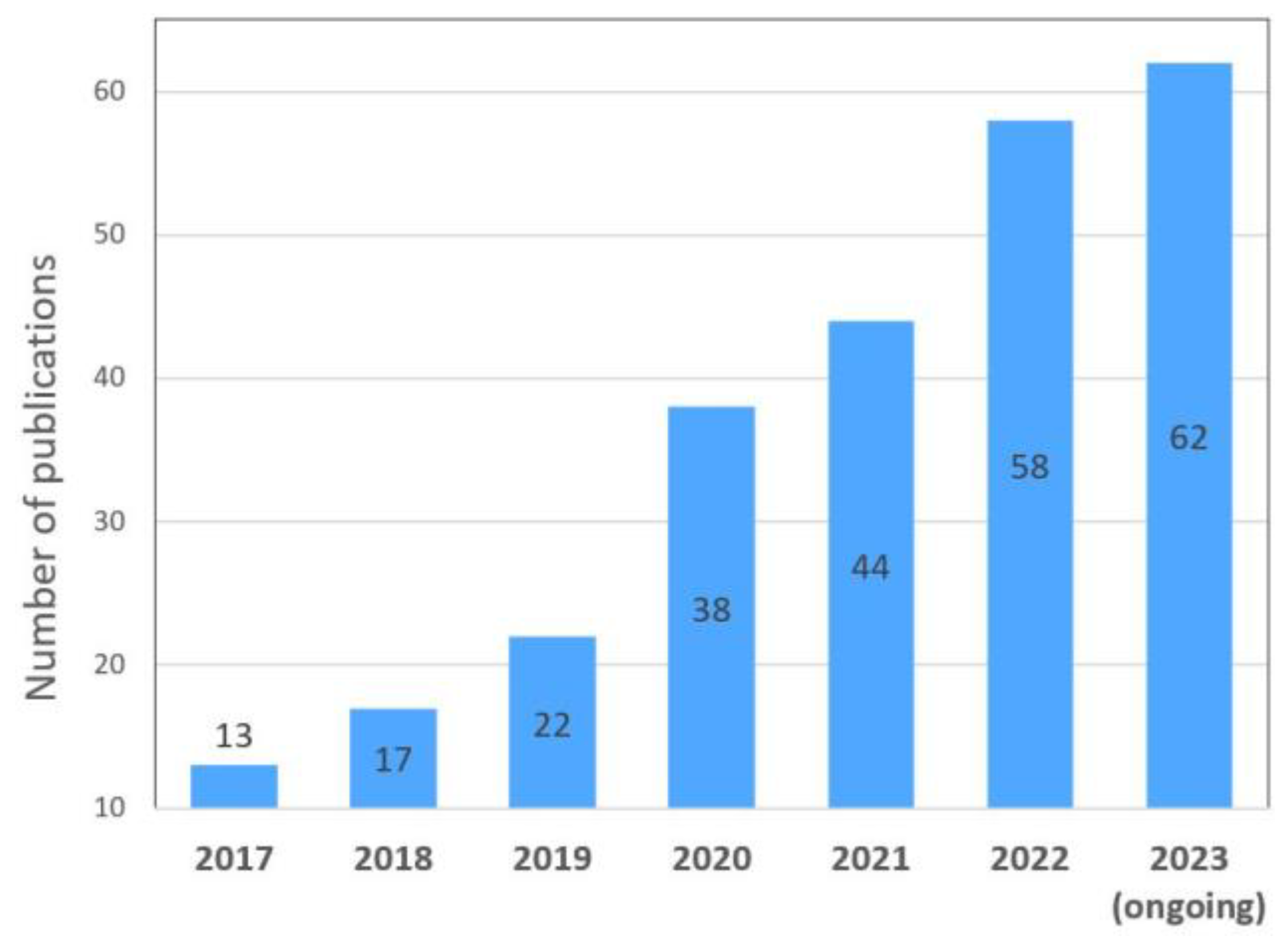
Figure 2.
Topics of studied tomato genes (2017-2023) by CRISPR/Cas9.
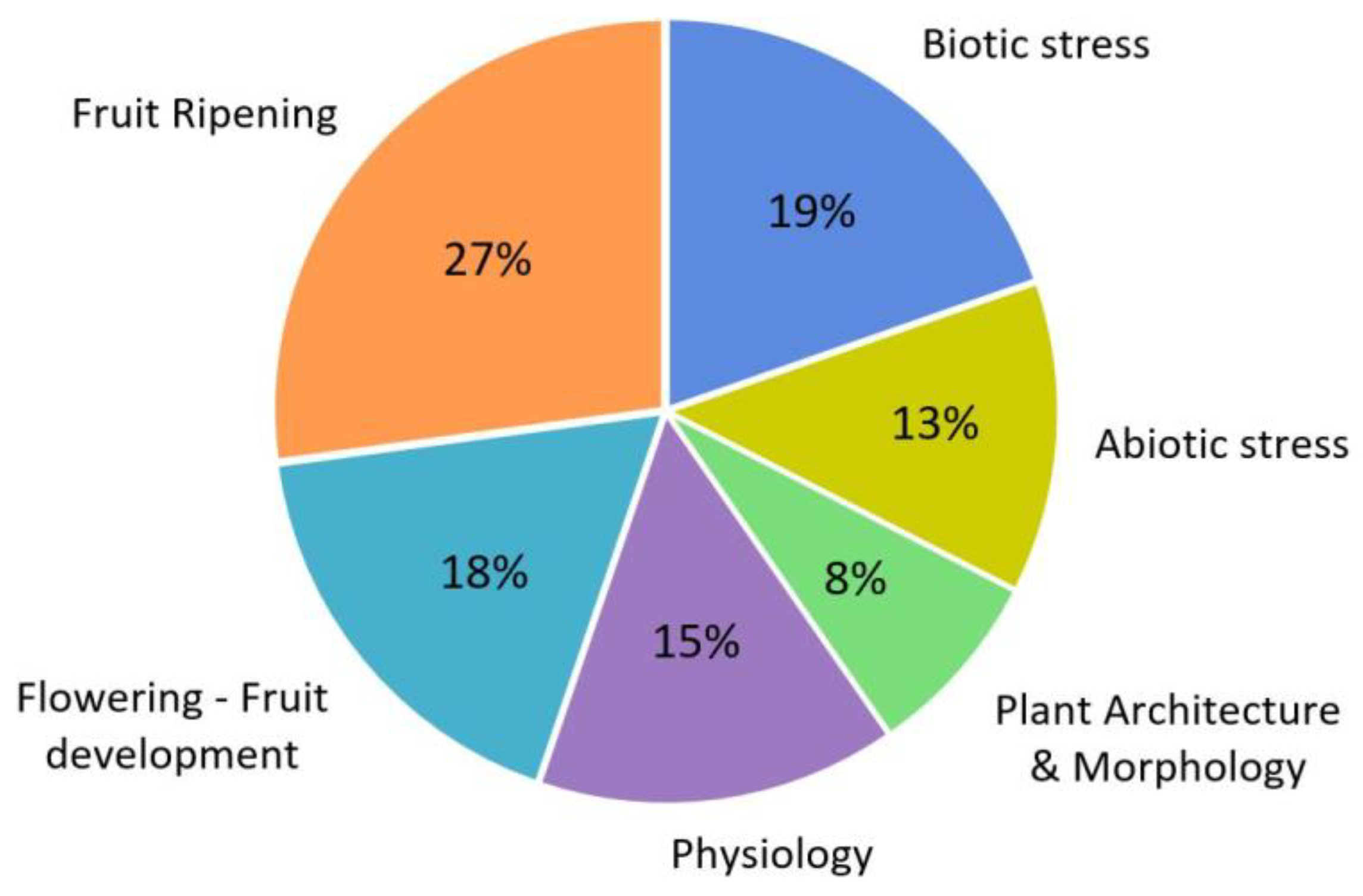
Figure 3.
The number of publications devoted to exploring the processes of flowering and ripening in tomatoes. Using CRISPR/Cas9 technology (a), using over- and heterologous expression approach (b), and using gene silencing technologies (c).
Figure 3.
The number of publications devoted to exploring the processes of flowering and ripening in tomatoes. Using CRISPR/Cas9 technology (a), using over- and heterologous expression approach (b), and using gene silencing technologies (c).
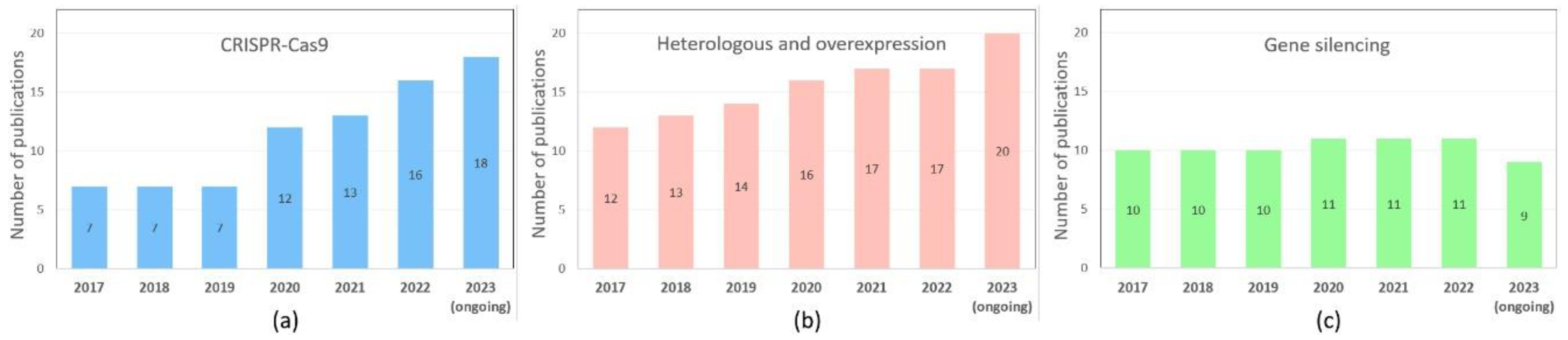
Figure 4.
A molecular interaction network model of ripening-related genes in tomato.
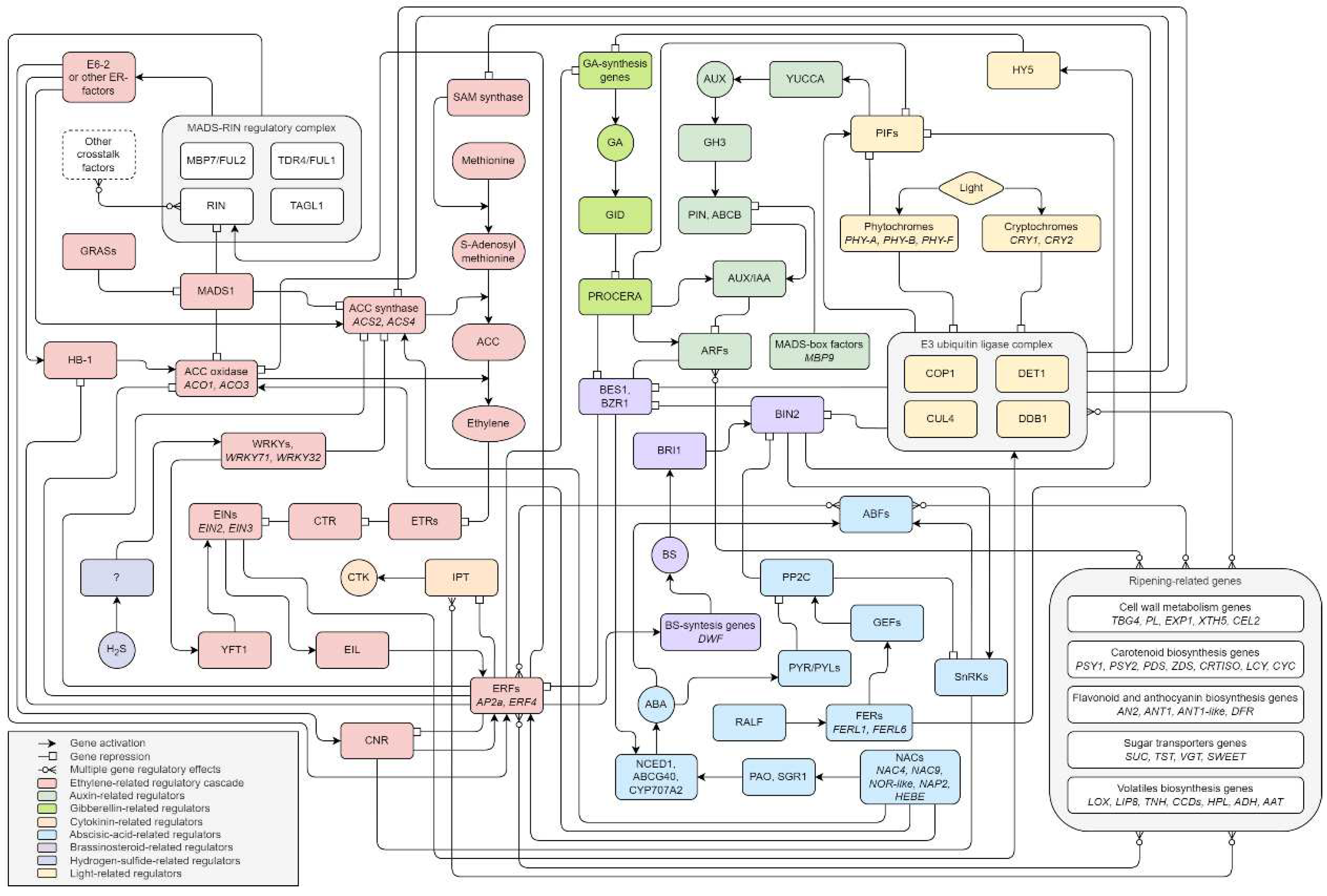
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Copyright: This open access article is published under a Creative Commons CC BY 4.0 license, which permit the free download, distribution, and reuse, provided that the author and preprint are cited in any reuse.
Submitted:
13 December 2023
Posted:
14 December 2023
You are already at the latest version
Alerts
A peer-reviewed article of this preprint also exists.
This version is not peer-reviewed
Submitted:
13 December 2023
Posted:
14 December 2023
You are already at the latest version
Alerts
Abstract
Tomato (Solanum lycopersicum L.) is one of the most commercially essential vegetable crops cultivated worldwide. In addition to the nutritional value, tomato is an excellent model for studying climacteric fruits' ripening process. Despite this, the available natural pool of genes that allows expanding phenotypic diversity is limited, and the difficulties of crossing using classical selection methods when stacking traits increase proportionally with each additional feature. Genetic engineering makes it possible to introduce changes in genes of interest without changing the allelic combinations characteristic of successful commercial varieties, to enhance the expression of existing genes, to create and transfer artificial and heterologous genes, and much more. However, this requires understanding the fundamental principles of the molecular interaction. Although the leading candidate genes for these components have been identified, a complete picture of their relationship has yet to be formed. The review summarizes the latest (2017-2023) achievements related to studying the ripening processes of tomato fruits. In this work, an attempt is made to systematize the results of various research and display the interaction pattern of genes regulating the process of tomato fruit ripening.
Keywords:
Subject: Biology and Life Sciences - Plant Sciences
1. Introduction
The fruits of angiosperms are included in the staple diet of humans and livestock. The transition of plants from the vegetative growth phase to the reproductive stage is the main switch in their life cycle. Fruit ripening is initiated and regulated by the combined action of various genetic factors in response to endo- and exogenous stimuli. The functions of ripening regulators are often conserved among different angiosperms.
In addition to the nutritional importance of tomatoes, this crop is a convenient object for studying the mechanisms of fruit ripening regulation. This is explained by its simple diploid genetics, small genome size, short life cycle, ease of transient and stable transformation, pronounced ripening phenotypes, an abundance of bioinformatics data, and well-characterized mutants.
Modern widespread varieties of tomatoes were obtained through domestication and subsequent selection. The selection of seemingly desirable traits carried out without an understanding of the nature of gene relationships has contributed to the reduction of genetic diversity in tomatoes. Consumer preferences and cultivation convenience have also contributed to this. On the other hand, the introgression of elite alleles into a cultivated variety helps create a hybrid genome with the allele of interest but also introduces undesirable genetic backgrounds in the form of linked genes from the allele donor. With the help of backcrossing, breeders can level out the manifestation of unwanted traits, but this is time-consuming and not consistently effective.
Genetic engineering methods provide significant potential for studying the genetic factors regulating fruit ripening. Thus, targeted genome editing technology using the CRISPR/Cas9 system allows researchers to create knockout alleles and make precise changes to the gene sequence. This, in turn, helps to study genes’ functions, relationships, and regulation. Despite the 10-year history of using this powerful technology, the number of publications using it, in which the tomato is the object, is growing steadily yearly (Figure 1). Thus, its research potential still needs to be exhausted.
An analysis of publications where the CRISPR/Cas9 system has been used in tomatoes over the past six years has made it possible to identify topics of most interest to the scientific community (Figure 2). It turned out that more than a quarter of the total number of works is devoted to the study of genes involved in the processes of fruit ripening (27%). Many of these genes encode transcription factors and transcriptional coregulators, microRNAs, or proteins involved in the epigenetic control of gene expression. In many cases, these regulators’ molecular mechanisms of action have yet to be studied, which is the reason for the growing interest in research in this area. Also, a considerable proportion of publications are devoted to the study of the regulation of the processes of flowering and fruit development (18%). The consistently current topic of stress (abiotic and biotic) occupies a third of the total number of publications. The remaining publications cover fields devoted to other physiological processes (15%) and plant architecture and morphology (8%).
Among the most prevalent genetic engineering methods used to study the regulation of ripening processes, the use of the CRISPR/Cas9 system is expected to increase (Figure 3a). At the same time, approaches that have already become classical, such as RNA interference gene silencing, and gene over- and heterologous expression, have not lost their relevance - the number of publications using them has remained consistently high over the past six years (Figure 3, b, c). Interestingly, there are a growing number of studies using gene overexpression to study ripening. All this suggests that, despite the large amount of accumulated data, the regulation of the ripening process still needs to be fully understood.
Ripening, as the final stage of development of tomato fruits, manifests itself in the form of bright pigmentation, increased aroma and taste, and softening of the pulp. These properties make the fruits attractive to animals, which act as seed dispersal vectors. Before the onset of ripening, these physiological changes are suppressed, but once the fruit enters the ripening phase, they occur in a highly synchronized manner with dramatic differences in gene expression patterns. Such radical changes are regulated by environmental factors such as temperature and light and by internal regulators, including hormones, transcription factors, and epigenetic modifications. Due to the commercial and biological importance of tomato fruit ripening, the mechanisms regulating this process are being actively studied.
2. Transcription factors regulating ripening
MADS-box genes are among the most widely represented and diverse transcription factors; consequently, they mediate various biological processes. Among them, regulating fruit ripening is one of the most prominent roles of MADS-box (MCM1, AGAMOUS, DEFICIENS, and SRF) genes. The transcription factor RIN (RIPENING INHIBITOR) has long been considered a major ripening regulator. RIN encodes a SEPALATA class MADS-box transcription factor. MADS-box family transcription factors typically function as multimers, and the MADS-box proteins TAGL1 (TAG-like) and two FRUITFULL (FUL) homologs (TDR4/FUL1 (tapetum degeneration retardation) and MBP7/FUL2 (MADS-box protein)), are coregulators with RIN and ripening regulators with overlapping functions [1]. Silencing of TAGL1 resulted in decreased levels of amino acids in fruit: aspartic acid, L-tyrosine, L-glutamine, L-phenylalanine, L-valine, L-leucine, isoleucine, and 5-caffeoylquinic acid [2]. TAGL1 was also found to regulate the synthesis of the glycoalkaloid α-tomatine negatively. As discussed in [3], TDR4/FUL1 and MBP7/FUL2 do not regulate ethylene biosynthesis but influence fruit ripening in an ethylene-independent manner. RIN often binds to demethylated sites in the promoter regions of ripening-related genes. RIN is induced early in ripening and stimulates ethylene-dependent and ethylene-independent pathways that promote ripening. Mediators in this process are response factors to ethylene (ERF, ethylene-responsive factor) and auxin (ARF, auxin response factor). ERF and ARF control their respective hormonal signaling pathways, regulating gene expression and hormonal signaling.
Several recent studies have clarified the function of RIN. Thus, [4] found that although RIN function is required for full ripening, RIN is not required for the initial ripening induction. The authors suggest that RIN acts redundantly (i.e., there are RIN homologs) or RIN-independent ripening induction occurs due to other transcription factors. In the second case, the authors concluded that a RIN-independent activator can induce the transcription of ripening-related genes even in RIN-deficient plants. Still, a mutant (defective) RIN protein can inhibit its activity. The chimeric transcription factor RIN-MC exhibits a negative role in ripening, promoting the mutant rin phenotype [5]. Other authors think that low ethylene concentrations initiate the ripening of mature green fruits, activate RIN expression, and lead to other changes, including a transition to a burst of autocatalytic ethylene synthesis [6]. Combined with the ethylene biosynthesis gene ACS2 (1-aminocyclopropane-1-carboxylate synthase), RIN has been shown to regulate the heat shock genes HSP17.7 [7] negatively. Therefore, RIN, ethylene, and other factors are necessary to complete the complete fruit ripening program. RIN is not only an activator of ripening but also a repressor of excessive softening [8]. It was found that when controlling the fruit ripening process, RIN binds to 6 lncRNAs [9].
RIN is reported to directly activate the expression of a novel gene, E6-2, involved in tomato fruit ripening [10]. Silencing of E6-2 leads to a delay in fruit ripening suppression of CNR (colorless non-ripening), PG (polygalacturonase), and ERF4 (ethylene-responsive factor) was observed, a decrease in the accumulation of carotenoids and lycopene (due to the suppression of PSY1, PDS and ZDS (phytoene synthase, phytoene desaturase, and zeta-carotene desaturase, respectively)), ethylene (decreased expression of the biosynthetic genes ACS2, ACO1 (1-aminocyclopropane-1-carboxylic acid oxidase), ACO3 and ethylene-sensitive E4, E8), increasing the content of pectin, cellulose, starch and soluble sugar (suppression of cell wall metabolism genes TBG4 (tomato beta-galactosidase), PL (pectate lyase), EXP1 (expansin), and XTH5 (xyloglucan endotransglucosylase/hydrolase)). The broad phenotypic pattern of RIN silencing is an attractive marker for testing molecular editing tools [11,12,13].
Genes with the NAC domain (NAM, ATAF1/2, and CUC2 (apical meristem, ARABIDOPSIS TRANSCRIPTION ACTIVATOR FACTOR, and cup-shaped cotyledon, respectively)) are considered to be another transcription factors that regulate tomato fruit ripening. It has been shown [14] that inhibition of NOR-like1 reduces ethylene production, delayed softening and loss of chlorophyll, and reduced lycopene accumulation. Activation of ethylene synthesis genes by NOR (non-ripening) and NOR-like genes is discussed further in [15]. Knockout of NAC-NOR suppressed fruit ripening (inhibition of ethylene synthesis, reduction of carotenoid accumulation, and fruit softening), and the opposite effect was observed with its overexpression [16]. Replacement of thymine with adenine in the ALC gene (alcobaca, NOR mutation) using homologous recombination contributed to an increase in the shelf life of tomato fruits [17]. Expression of peach NAC1 in tomatoes has been shown to enhance ripening in a delayed ripening (NOR) mutant and restore the synthesis of volatile esters [18]. Overexpression of NAC6 resulted in increased levels of endogenous abscisic acid, which affected the transcription of ripening genes [19]. Transfer of the kumquat NAC22 gene to tomato increased the expression of most carotenoid biosynthesis genes, accelerated the transformation of plastids into chromoplasts, and promoted color changes [20]. NAM1 (no apical meristem), another factor with an NAC domain responsible for the regulation of ethylene biosynthesis, also controls tomato ripening, as confirmed by delayed ripening in CRISPR/Cas9 mutants and accelerated ripening for lines overexpressing NAM1 [21]. Repression of NAM gene domains is carried out by miR164a [22,23,24]. In addition, the HWS (HAWAIIAN SKIRT) gene, encoding an F-box protein, regulates the number of floral organs by modulating the transcription levels of the miR164, CUC1 and CUC2 (cup-shaped cotyledon) genes. HWS is also involved in petals’ cell proliferation and mitotic growth [25].
It was previously shown that a representative of genes with the NAC domain NAP2 (Arabidopsis NAC domain-containing protein) activates the aging gene SAG113 (senescence-associated gene), protein phosphatase), chlorophyll degradation genes SGR1 (stay-green), PAO (polyamine oxidase), and NAP2, and also directly controls the expression of genes essential for the biosynthesis of abscisic acid NCED1 (9-cis-epoxycarotenoid dioxygenase), ABCG40 (Arabidopsis thaliana ATP-binding cassette), and CYP707A2 [26]. These interactions suggest the influence of NAP2 on leaf senescence and yield in tomato. Another new gene, named by the authors HEBE in honor of the Greek goddess of youth, has similar functions [27].
Numerous studies have shown that the CNR gene is the most important regulator of tomato fruit ripening. However, recent research [28] has called this assumption into question. CNR knockout lines exhibited only a ripening-arrested phenotype, while NOR knockout (non-ripening) lines exhibited a partial non-ripening phenotype similar to RIN mutants. Both knockouts differed from the strong, non-ripening phenotypes of their natural mutants. It became apparent that the expression of characteristic ripening genes, such as ACS2, ACO1, PSY, PG, and EXP, is not entirely suppressed in CRISPR/Cas9 lines compared to natural mutants. As the authors concluded, differences in the expression of the genes in question are explained by different degrees of methylation, and they also concluded that the regulatory network of transcription factor genes is redundant. Regulation of NOR may also be associated with something else: sulfoxidation of the NOR transcription factor with the help of MSRs (methionine sulfoxide reductase) proteins modulates the ripening process by reducing the DNA-binding ability of NOR [29].
It is known that the fruits of the tomato epimutant cnr fail to ripen and remain colorless. The SPL (SPOROCYTELESS) gene family consists of a group of genes encoding SBP (SQUAMOSA promoter binding proteins)-box transcription factors, and their protein products bind to the promoter of the floral meristem identity gene SQUAMOSA. Evidence shows that SPL-CNR interacts with SnRK1 (SNF1-related protein kinase) [30]. Suppression of SnRK1 by virus-induced gene silencing (VIGS) inhibits fruit ripening and leads to decreased expression of a wide range of ripening-related genes. This suggests that SnRK1 transcription and subsequent post-translational SPL-CNR-SnRK1 interaction are biologically crucial for tomato fruit ripening. The authors suggest that the involvement of SnRK1 in fruit ripening may be due to the physical interaction of proteins between the SnRK1 gene product and SPL-CNR and subsequent phosphorylation of SPL-CNR due to the kinase activity of SnRK1 [30].
The role of some MBP transcription factors in the ripening process has recently been studied. Thus, suppression of the MBP8 factor shortened the fruit ripening time, suggesting an increase in the activity of ethylene synthesis genes [31]. Meanwhile, carotenoids accumulated to higher levels, and the expression of PSY1, PDS, and ZDS was enhanced in MBP8 RNAi-silenced fruits. The activity of cell wall genes also changed, manifested in the softening of fruits. Silencing MBP15 in [32] delayed tomato ripening, and gibberellin, carotenoid, and ethylene biosynthesis genes were repressed. MBP15 was found to interact with RIN [32].
GRAS (gibberellic acid insensitive (GAI), repressor of GAI (RGA), and scarecrow (SCR)) proteins are plant-specific transcription factors that play critical roles in plant development and stress response. It turned out that they also take part in regulating fruit ripening. For example, silencing GRAS2 reduces tomato fruit weight, which has been attributed to insufficient levels of gibberellic acid during initial ovary development [33]. Overexpression of GRAS4 accelerated fruit ripening (due to activation of expression in the promoter region of ethylene biosynthesis genes and repression of the negative regulator of ripening MADS1). It increased the total content of carotenoids [34]. GRAS24, in addition to flowering and ripening, is responsible for a variety of other agronomic traits, including plant height, leaf architecture, number of lateral branches, root length, and the observed pleiotropic effects in plants overexpressing GRAS24 are due to impaired modulation of gibberellin and auxin signaling [35].
Transcription factors of the WRKY superfamily exhibit up-regulation during fruit ripening. WRKY32 binds to W-box and similar motifs in the regulatory region of the YFT1 (yellow-fruited tomato) promoter and induces its expression [36]. YFT1 encodes the EIN2 protein, a major ethylene signal transduction component. Suppression of ethylene production resulted in delayed chromoplast development, decreased carotenoid accumulation, and a yellow fruit phenotype. Twelve WRKY genes were also shown to be ethylene-responsive (ER), 8 of which activated the promoters of color change-associated genes (PPH (pheophytinase), PAO (polyamine oxidases), PSY1, and PDS) [37]. In addition, protein interactions were found between WRKY17 and RIN/ERF2b/ERF7, WRKY33 and ERF7, WRKY54 and ERF2b, WRKY16 and WRKY1, which only confirms the complexity of the networks of ripening regulators [37].
3. Epigenetic modifications as regulators of ripening
DNA methylation has a critical role in a wide range of cellular functions. For example, a decrease in DNA methylation levels can be observed during fruit ripening, which is explained by DNA demethylases (DML) activation. In DML2 loss-of-function mutants generated by targeted editing, increased DNA methylation was found not only in genes induced during ripening but also in genes repressed during ripening [38]. However, a recent study found that the highly mobile protein positively regulates DML2 expression of 3-hydroxy-3-methylglutaryl-coenzyme A reductase (HMGA) [39]. In [40], expression of the mammalian demethylase TET3c (ten-eleven translocation) in tomatoes resulted in the activation of expression of the previously undescribed gene CEN1.1. The activation intensity of CEN1.1 expression correlated with increased hypomethylation in its promoter, suggesting that CEN1.1 expression is associated with DNA methylation of CHH promoter sites (H=A/C/T). Phenotypically, CEN1.1 has emerged as a repressor of flowering in tomatoes, leading to the development of leaves on inflorescences. Paradoxically, this led to an increase in the number of fruits but to a longer time for their ripening. Thus, this study provides an exciting approach to identifying methylation-associated genes.
In plants, cytosine methylation plays a crucial role in suppressing the movement of transposable elements. Methylation is maintained by DNA methyltransferases MET1, and CMT3 (chromomethylase), as well as additional proteins (for example, DDM1 (decreased DNA methylation)) involved in maintaining heterochromatic structure. The methylase encoded by MET1 is a key DNA methylase responsible for maintaining CG methylation in plants. Loss-of-function mutants of the MET1 gene had pleiotropic developmental phenotypes manifested as small curled leaves, defective flowers, and parthenocarpic fruits [41]. Also, the knockout of MET1 resulted in changes in the expression profiles of RIN target genes, such as ACC2 (acetyl-CoA carboxylase 2). In another study, suppression of MET1 by VIGS in a hypermethylated epimutant CNR promoted vivipary development [42]. The authors explain this by a decrease in the concentration of abscisic acid and NCED transcripts involved in its biosynthesis. The NCED silencing had similar consequences. Methyltransferase DRM7 (domains rearranged methyltransferase) has been shown to influence chloroplast development by modulating starch accumulation and chlorophyll synthesis. It has an epi-effect on leaf senescence, affecting tomatoes’ vegetative growth [43]. The transient expression of arginine methyltransferase PRMT1.5 in tomatoes inhibited the accumulation of carotenoids and anthocyanins [44].
Short interfering RNAs (siRNAs) also mediate DNA methylation through RNA-directed DNA methylation (RdDM). It has been established that heterochromatic mobile elements in plants with DDM1 dysfunction are deprived of mCG and mCHG, which generally keep them inactive [45]. Methylation of CHH sites increased for some heterochromatic transposons and, conversely, decreased for those localized in euchromatin. Knockout of CMT4 chromomethylase caused severe morphological changes in tomato plants, accompanied by defects in leaves, pollen, and seeds [46].
Another mode of epigenetic regulation is post-translational modifications of histones. Histone acetylation is known to be associated with gene activation. In contrast, histone methylation can be associated with either activation or repression depending on the lysine residue and the number of methyl groups added. In [47], RNA-seq profiling showed a significant increase in the expression of methylases MET1 and CMT3 and a minor increase in the demethylase DML2 during the fruit set, which is associated with their role in maintaining post-replication DNA methylation during extensive cell division characteristic of early stages of development fetus However, their abundance was significantly lower than that for histone marks H3K9ac and H3K4me3, determined using chromatin immunoprecipitation sequencing. This implies that changes in the transcriptional profile underlying the fruit set are more closely related to histone modifications than methylation. Histone modification is based on the histone methyltransferase genes SDG27, SDG5, and SDG16 (set domain group). However, the authors could not create homozygous loss-of-function mutants of these genes, suggesting their exceptional biological importance. Mutants heterozygous for these genes exhibited parthenocarpic fruits. The function of other histone lysine methyltransferases SDG33 and SDG34 was revealed in [48]. They were found to regulate the expression of nitrogen-responsive genes and physiological changes in an organ-specific manner.
It has been demonstrated that histone demethylation leads to activating tomato fruit ripening genes [49]. Here, the JMJ6 (Jumonji C-terminal domain-containing demethylase) gene was found to encode a histone lysine demethylase that specifically demethylates H3K27. Its overexpression accelerates the ripening of tomato fruits, which is associated with increased expression of the RIN, ACS4, ACO1, PL, and TBG4 genes. As study [50] showed, JMJ4 mediates abscisic acid-induced leaf senescence in tomatoes.
By knocking out the HTA1 genes of histone H2A and subsequent production of double homozygous mutants, changes were identified in the expression patterns of many biological ripening processes, including cell redox homeostasis, mRNA splicing, cell cycle regulation, translation, etc. [51]. Moreover, for three genes of carotenoid biosynthesis, PSY1, PDS, and VDE, expression was high regardless of the fruit ripening stage [51]. Histone deacetylation has been associated with transcriptional repression. Histone deacetylases carry out this process. There is evidence that they can act as both positive and negative ripening regulators. Indeed, RNAi silencing of the HDT3 (histone deacetylase) gene led to the suppression of genes for ethylene synthesis (ACS2, ACS4, ACO1, ACO3), carotenoids (PSY1), cell wall metabolism (HEX (acetylhexosaminidase), MAN (mannosidase), TBG4, XTH5, and XYL (xylanase)), as well as general genes, associated with ripening (RIN, E4, E8, PG, Pti4, LOXB (lipoxygenase)) [52]. In contrast, in [53,54], silencing of the HDT1 gene led to opposite results for these same transcripts. In this regard, the molecular mechanisms of regulation of these genes remain to be studied.
Histone acetyltransferase GCN5 acetylates histone H3 lysine (H3K14ac) and affects the levels of H3K9ac and H3K27ac. Its suppression leads to the loss of shoot apical dominance and a decrease in the size of the plant apical meristem [55]. It has also been established that GCN5 can increase WUSCHEL transcript levels. The expression of WUSCHEL can also be regulated by chromatin remodeling factors, such as the histone deacetylase HDA19 [56]. Here, the deacylation mechanism was found to involve the inhibitor gene IMA (inhibitor of meristem activity) acting as an adapter protein to form a chromatin remodeling complex together with the zinc finger protein C2H2 KNU (KNUCKLES) and the transcriptional corepressor TOPLESS.
4. Hormonal control of ripening
Auxin regulation is involved in all plant processes, including cell elongation and division, the formation of the architecture of roots, leaves, inflorescences, the development of embryos and fruits, and responses to stress. The leading site of auxin biosynthesis is young leaves and their primordia. The amino acid tryptophan is the primary precursor of auxins. Knockout of any of the auxin synthesis genes is associated with lethal phenotypes, so attention is paid to genes providing auxin-mediated inactivation (GH3, GRETCHEN HAGEN), transport (PIN, ABCB (PIN-FORMED, ATP binding cassette subfamily B, respectively)), and signal transduction (ARF (auxin response factor), Aux/IAA).
By conjugating auxins to amino acids for storage or degradation, members of the GH3 family, encoding acyl acid amidosynthetases, are critical for maintaining auxin homeostasis. In tomatoes, GH3.15 has been shown to regulate lateral root development and response to gravitropism by modulating auxin homeostasis [57], GH3.8 controls plant height [58], GH3.4 negatively regulates mycorrhization [59,60], GH3.2 affects fruit ripening in the early stages [61].
PINs are one of the facilitators of intercellular auxin transport. VIGS PIN1 accelerates flower abscission by increasing the accumulation of auxin in the ovule and reducing the auxin content in the abscission zone [62], and its negative regulator is the transcription factor MBP9 [63].
ARFs are plant-specific transcription factors that directly bind to auxin response elements in the promoters of auxin-responsive genes. ARF5 has been shown to regulate fruit set and development [64], ARF10 is involved in the accumulation of chlorophyll and sugar during fruit ripening [65], ARF19 is involved in leaf development [66], several ARFs (ARF6A, ARF8A, ARF8B, and ARF24) interact with the transcriptional repressor IAA9 [67] and regulate leaf shape [68], ARF10A is essential for the growth of leaf blades and formation of floral organs [69].
Phytohormones gibberellins regulate various physiological processes of plants. They promote plant growth participate in stem elongation, expansion of leaf blades, pollen development, flowering, and seed germination. Gibberellins are synthesized in plastids and then modified in the endoplasmic reticulum and cytosol.
DELLA (GRAS gene encodes protein containing D-E-L-L-A amino acid sequences) proteins are nuclear-localized negative growth regulators. Gibberellins promote DELLA degradation by assembling the E3 ubiquitin ligase complex, followed by protein degradation. DELLA is encoded by the PROCERA gene, and its loss of function in the homozygous state results in dwarfism [70] and parthenocarpy [71]. The degradation of proteins, including DELLA, is controlled by a complex regulatory network involving connections between several signaling pathways [72]. DELLA proteolysis is mediated by the gibberellin-activated receptor GID. Knockout of their coding genes also results in a dwarf phenotype [73]. There is evidence of cross-signaling between the gibberellin and abscisic pathways [74], and the DELLA protein is an activator of abscisic acid transporters (AIT), regulating transpiration through stomatal closure [75]. Tomato PROCERA activity is assumed to be necessary to transition tomatoes to flowering. DELLA protein directly or indirectly promotes the expression of SFT (SINGLE FLOWER TRUSS) in leaves, as well as SBP and AP1/MC, together with microRNAs in the shoot apex [76].
Recently, factors mediating gibberellin-dependent regulation have also received attention. Thus, silencing of the GRAS15 transcription factor led to pleiotropic phenotypes, including reduced plant height, small leaf size with pointed edges, as well as an increased number of nodes, lateral shoots, and petiole length, which is explained by the suppression of gibberellin synthesis genes [77]. The helix-loop-helix transcription factor gene PRE2 is induced by gibberellin. Its silencing has been shown to cause reductions in fruit size, seed size, pericarp thickness, and placental size [78]. These changes are associated with decreased expression of xyloglucan endotransglucosylases XTH2 and XTH5. PREs regulate many processes - their overexpression in tomatoes leads to multiple morphological changes, including changes in leaf angle, internode length, leaf curl, and pigment composition [79,80].
Cytokinins promote the development of shoots, provide stress resistance, and delay aging. These isopentenyladenine derivatives are formed mainly in the roots and transported to the aerial parts. The main rate-limiting enzyme for cytokinin synthesis is isopentenyltransferase (IPT). It has been shown that overexpression of the IPT gene leads to significant phenotypic changes and slower leaf senescence only under the control of a root-specific promoter [81].
Cytokinin catabolism is carried out by cytokinin oxidases (CKX). Overexpression of CKX2 in tomato fruit decreased cytokinin levels [82]. It is also shown here that endogenous cytokinins regulate the division of pericarp cells, which subsequently determines the size of the fetus. Ethylene is the simplest unsaturated hydrocarbon with the formula C2H2. In plants, it acts as a global regulator of developmental processes. The ethylene biosynthetic pathway includes three steps: S-adenosylmethionine synthetase (SAMS) modifies methionine to form S-adenosylmethionine (SAM), SAM is converted to 1-aminocyclopropane-1-carboxylic acid (ACC) by synthase (ACS), and in the last step ACC converts ACC oxidase (ACO) with the formation of ethylene.
In studies on tomatoes, ethylene is considered a participant in signaling cascades, including during the ripening process. It has been established that its synthesis during ripening is presumably regulated by FER receptor kinases (FERONIA). FERL6 and FERL1 were found to interact physically with the SAMS promoter [83]. Expression of FER genes in tomatoes showed negative regulation of ethylene accumulation at the initial stages of fruit development and, as a consequence, delayed fruit ripening.
The promoter of the transcription factor EIN3 (ethylene insensitive) has been shown to contain several motifs associated with hormones influencing fruit development and ripening [84]. Overexpression of EIN3 in tomatoes resulted in the activation of the expression of ethylene biosynthesis genes ACO1, ACS1, and SAMS1, which promoted early fruit ripening. Accordingly, EIN3 silencing showed the opposite effects. An EIN3-like gene causes premature onset of ovule senescence [85].
Ethylene is bound by a family of ETR (ethylene receptor) proteins located in the membrane of the endoplasmic reticulum. ETRs have functional redundancy. ETR3-mediated signaling inhibits pollen tube growth without sufficient ethylene [86]. ETR3 promotes the activation of cell wall remodeling genes and Ca2+ transporters — overexpression of ETR7 results in earlier flowering, short plants, and small fruits [87]. Targeted base substitution in ETR1/2 causes a delay in ripening and ensures prolonged storage of fruits [88,89].
Ethylene response factors (ERFs) are signaling components involved in ethylene-dependent developmental processes. They can perform both positive and negative regulation of target genes. Their number is large, as is the specificity of the reactions of tomato genes to ethylene: regulation of fruit ripening processes [90,91,92,93], control of aging [94], participation in the activation of protective reactions [95,96,97], growth [98,99], accumulation of chlorophyll and formation of chloroplasts [100], and regulation of other signaling pathways [101].
Brassinosteroids, which include various polyhydroxylated steroidal phytohormones, influence many critical agronomic traits related to growth, photosynthesis, morphology, and yield. The synthesis of brassinosteroids occurs along three pathways, in which campesterol is the initial substrate. Crosstalk between brassinosteroids and redox signals suggests a direct involvement of the former in the plant response to stress [102,103]. However, recent studies also reveal a connection between the brassinosteroid and ethylene pathways. Recently, it was demonstrated that overexpression of one of the genes for the brassinosteroid synthesis enzymes DWARF (DWF) in tomatoes promotes fruit softening, lycopene synthesis, and ethylene production, while gene knockout inhibits them [104]. It was concluded that APETALA2a (AP2a) promotes ethylene signaling to regulate brassinosteroid signaling. Also, tomatoes with overexpression and silencing of the cytochrome P450 monooxygenase CYP90B3 gene showed a correlation of the content of bioactive brassinosteroids with the processes of tomato fruit ripening, including softening, the content of soluble sugars and aromatic volatiles [105].
Research into brassinosteroid-dependent pathways is ongoing. The specific receptor for brassinosteroids is BRI1 (BRASSINOSTEROID INSENSITIVE1). Upon binding of brassinosteroids to its extracellular domain, dimerization of BRI1 and the coreceptor BAK1 (BRI 1-associated receptor kinase 1) occurs. The signal is then transmitted through a phosphorylation cascade involving BSK1 (BR-signaling kinase 1), CDG1 (constitutive differential growth), BSU1 (BRI1 SUPPRESSOR), and BIN2 (BRASSINOSTEROID INSENSITIVE2). Subsequently, BIN2 is inactivated, and two transcription factors, BZR1 (brassinazole resistant) and BES1 (BRI1-extra microsporocytes-suppressor 1), are dephosphorylated by protein phosphatase PP2A. BZR1, BES1, as well as other nuclear factors (for example, BIM1) are regulators of brassinosteroid-dependent genes.
Overexpression of BRI1 [106] in tomatoes improved carotenoid accumulation by increasing the expression of DXS (1-deoxy-D-xylulose 5-phosphate synthase), GGPS (geranylgeranyl pyrophosphate synthase), and PSY1. In addition, brassinosteroids induced the expression of genes involved in its ethylene biosynthesis (ACO1 and ACS2). Similar results were achieved by modifying threonine-1050 BRI1, resulting in plants with high levels of BRI1 autophosphorylation [107]. A recent study on BRI1 showed that the receptor also positively regulates a tomato’s tolerance to cold stress [108]. Brassinosteroids are capable of inducing early flowering. This is supported by the interaction of the suppressor of BIN2 signaling with the early flowering locus FRIGIDA [109]. There is evidence that canonical signaling pathways initiated by BRI1 are involved in xylem differentiation and wood formation in tomatoes through activation of the BZR1/2 transcription factors [110]. The BZR1 homolog has been shown to interact with BIM1 to act as a negative regulator of pericarp cell expansion [111]. According to available information, BZR1 is also a trans-activator of the promoter of the SUN gene (encodes Sad1/Unc-84 (SUN)-domain proteins), responsible for elongation of tomato fruits, and BZR1-knockout tomato phenotypes show redundancy of its homologs [112]. In addition, brassinosteroids promote bud growth in tomatoes through direct transcriptional regulation of BRANCHED1 (BRC1) via the signaling component BZR1 [113].
In the study [114], the authors focused on BES1, a key transcription factor in the brassinosteroid signaling pathway. BES1 was found to bind to the promoter of the fruit-softening inhibitor PMEU1 (pectin methylesterase). Knockdown or knockout of BES1 in tomatoes resulted in increased shelf life without negatively affecting the appearance and nutritional composition of the fruit.
Altered regulation of brassinosteroids may influence cell elongation and division, leading to altered fetal morphology. For example, a premature stop codon at the GLOBE locus containing a brassinosteroid hydroxylase sequence resulted in a spherical phenotype of tomato fruit, which had a flattened shape in the wild type [115]. Since GLOBE and FW3.2 (KLUH) were found to be members of the same cytochrome P450 family, the authors hypothesized that both may act similarly in regulating fruit size and shape.
During oxidative stress following pesticide application, plants use glutathione to clear excess reactive oxygen species. Brassinosteroids induce pesticide metabolism by activating GRX (glutaredoxin) gene expression through transcription factors [116,117].
Abscisic acid is considered an antagonist of brassinosteroids during fruit ripening. The abscisic acid signaling pathway consists of the family of receptor proteins PYR (PYRABACTIN RESISTANCE), PYL (PYR1-like), RCAR (regulatory components of ABA receptors), protein phosphatases PP2C, and SnRK2 kinases. There is evidence of positive regulation of abscisic acid biosynthesis by brassinosteroid signaling. BZR1 (brassinazole resistant) was found to mediate brassinosteroid signaling by promoting abscisic acid biosynthesis through direct transcriptional regulation of NCED1 [118]. Here, we showed that BIN2 negatively regulated BZR1 protein accumulation and cold tolerance by suppressing abscisic acid biosynthesis.
Suppression of PP2C3 expression in tomatoes accelerated the onset of fruit ripening and affected their glossiness by changing the external structure of the epidermis [119]. In transgenic plants, an increase in the expression of SnRK2, PYL receptors, various cutin synthesis and transfer genes, and CYP (cytochrome P) genes was observed. The role of PP2C as a negative regulator in abscisic acid signaling was further supported in [120], where alteration of PP2C5 expression affected fruit quality traits, including pericarp thickness, shape, seed number, and soluble solid content. In addition, PP2C1 silencing increased accumulation of endogenous abscisic acid and accelerated ethylene release in transgenic tomatoes compared to wild-type fruit [121]. PP2C1-RNAi lines had abnormal flowers, and pedicel abscission was impaired.
Abscisic acid homeostasis is regulated by its conjugation with glucose using uridine diphosphate glucosyltransferases (UGT). It was shown that RNAi silencing of the UGT75C1 gene significantly increases the level of expression of the CYP707A2 hydrolase gene while not affecting the expression of the key gene for abscisic acid biosynthesis NCED1 [122]. Suppression of UGT75C1 expression significantly accelerated fruit ripening by increasing abscisic acid levels and promoting early ethylene release.
The PYL9 protein has been identified as a positive regulator of abscisic acid signaling [123]. Depending on abscisic acid concentration, PYL9 can inhibit the protein phosphatase PP2C. In tomatoes overexpressing PYL9, fruit ripening was significantly accelerated due to the early release of ethylene.
The abscisic acid-induced oxidase gene DAO2 (dioxygenase for auxin oxidation) inhibited hypocotyl elongation in tomatoes, exhibiting an antagonistic role to auxins [124].
SnRK phosphorylation is mediated by the protein kinase MAPK11, thereby regulating abscisic acid biosynthesis and signaling [125].
In addition, a new transcriptional repressor of abscisic acid biosynthesis, EAD1 (ERF-associated amphiphilic repression (EAR) motif-containing ABA down-regulated), was recently discovered [126]. Although the authors have not studied the molecular mechanism of repression, its implementation is possible either through the recruitment of histone deacetylases with subsequent formation of a complex with co-suppressors or through direct or indirect binding to transcription factors.
Hydrogen sulfide counteracts the effects of ethylene during ripening. The assimilation of sulfates in chloroplasts can produce endogenous hydrogen sulfide, and the main enzymes in this process are sulfite reductases. Cytosolic hydrogen sulfide can also be generated from cysteine by cysteine desulfhydrase 1 (DES1/LCD1). Loss-of-function mutations in LCD1 in tomatoes [127] increase the expression of genes for ethylene synthesis (ACO1, ACO3, and ACS2), carotenoids (PSY1, PDS, and ZDS), and cell wall metabolism (CEL2, EXP, XTH5, PG, and TBG4). Knockout of the tomato D-cysteine desulfhydrase (DCD) gene results in increased expression of ripening-related genes, including NYC1, PAO, SGR1, PDS, PSY1, ACO1, ACS2, E4, CEL2, and EXP [128].
An attempt to understand hydrogen sulfide-mediated regulation of ripening is made in [129]. The authors suggest that the ubiquitin-protein ligase BRG3 undergoes persulfidation at two cysteine residues, leading to a decrease in ubiquitinating activity and its interaction with the repressor transcription factor WRKY71. This leads to increased binding of WRKY71 to the promoter of cyanoalanine synthase CAS1, which inhibits its transcription and, thus, prolongs fruit ripening.
There is also confirmation that hydrogen sulfide is a regulator of aging, which is noticeable in changes in the expression of chlorophyll degradation genes (NYC1, PAO, PPH, SGR1) and the aging-associated gene SAG [130].
5. Abiotic ripening factors
Fruit ripening is also regulated by signaling systems activated in response to abiotic stimuli. It has been reported that changes in light sensitivity and light-sensitive signaling in tomatoes can significantly change fruit development and quality characteristics. In this context, phytochromes act as molecular switches in response to light. Following light activation, phytochromes deactivate photomorphogenic response repressor proteins (e.g., COP1, CUL4, DDB1, DET1, and PIF).
Using RNAi silencing of the phytochrome genes PHYA, PHYB1, and PHYB2, it was shown that PHYA positively affects the differentiation and division of tomato plastids through changes in the expression of both light-dependent genes and cytokinin-dependent genes [131]. Regulators of carotenoid biosynthesis (GGPS, PSY1, and PDS) were also affected, resulting in decreased carotenoid biosynthesis during fruit ripening.
As for the repressor proteins mentioned above, their effect on ripening is also being studied. Thus, according to [132], overexpression of COP1 (CONSTITUTIVE PHOTOMORPHOGENIC) from Solanum melongena in tomatoes caused a delay in fruit ripening by 3-6 weeks. These transgenic plants showed decreased ethylene production due to suppressing the expression of the central genes of its biosynthesis ACO1, ACO3, and ACS2. The carotenoid biosynthesis genes PSY1, PDS, and ZDS were also downregulated. In [133], using the DDB1, DET1, and CYC-B genes as an example, the multiplex Target-AID (activation-induced cytidine deaminase) technique was developed. As a result, the authors obtained two lines of triple mutants, in which each gene had two-point substitutions, which showed a higher accumulation of carotenoids and lycopene compared to the wild type. In tomatoes, PIF-dependent light signaling has been reported to regulate fruit development and influence nutritional value and ripening time. Transient overexpression of PIF3 in tomato fruit resulted in decreased GGDR mRNA levels, which was inversely related to PIF3 transcript levels [134]. These data indicate that PIF3 mediates PHY-dependent regulation of tocopherol biosynthesis through transcriptional inhibition of geranylgeranyl diphosphate reductase GGDR expression in tomato fruit. Evidence shows that PIF4 can regulate hypocotyl elongation, plant growth, flowering, and leaf senescence in response to light and temperature [135]. The authors support this statement by obtaining tomatoes with RNAi-mediated knockdown of PIF4, which showed increased carotenoid content, accelerated fruit ripening time, and delayed leaf senescence. A small number of flowers and a decrease in vegetative mass were observed in such plants. Knockout of PIF3 using CRISPR/Cas9 led to the arrest of phase I of pollen mitosis, which was reflected in its non-viability [136]. Glutamate synthase (GLT1) and cell wall invertase (CWIN9), involved in auxin and sugar homeostasis, respectively, have also been shown to colocalize with PIF3 in anthers and are directly regulated by PIF3. Knockout lines of GLT1 and CWIN9 (cell wall invertase) showed a similar phenotype. VIGS-mediated silencing of the light-signaling transcription factors HY5 and PIF3 led to changes in glycoalkaloid levels in tomato leaves compared to wild type, suggesting their involvement in the regulation of target genes of glycoalkaloid metabolism [137].
While the most abundant antioxidant in tomato fruit is the lipophilic carotenoid lycopene, levels of water-soluble flavonoids (including anthocyanins) are suboptimal. Plants accumulate anthocyanins in response to various stress events such as low temperature, drought, UV radiation, intense light, and nutrient deficiency, acting as an antioxidant and photoprotective agent. The bZip transcription factor HY5 is believed to be a significant regulator of anthocyanin accumulation in plants in response to light [138,139]. However, research [140] has cast doubt on the accuracy of this statement. By creating HY5 knockout mutants, the authors demonstrated a reduced anthocyanin content, which suggests the presence of additional pathways for their synthesis independent of HY5. Indeed, eight candidate anthocyanin transcription factors have been identified.
A recent study has uncovered the function of the little-studied PHY-F. It turned out that PHY-F is a low-flux radiation sensor [141]. It forms dimers with PHYA and/or PHYB, with which it makes additive contributions to various processes of photomorphogenesis.
In addition to the phytochromes of red and far-red light receptors, there are also cryptochromes of blue light receptors – CRY1 and CRY2. Tomato lines overexpressing CRY1a showed significant accumulation of anthocyanins through the regulation of genes encoding key enzymes of anthocyanin biosynthesis (e.g., AN2, DFR (dihydroflavonol 4-reductase)) [142]. The same study showed that blue light consistently induced overexpressing tagged HY5 protein accumulation in tomatoes. In addition, it was shown that under the influence of blue radiation, repression of COP1 (CONSTITUTIVE PHOTOMORPHOGENIC) transcription was observed, which was confirmed by the creation of lines with RNAi-COP1. Ultimately, the silencing of HY5 and two anthocyanin biosynthesis genes (CHS1 (chalcone synthase) and DFR) in CRY1a lines was accompanied by a decrease in anthocyanin accumulation. Moreover, CRY1a was found to be critical for regulating starch accumulation in chloroplasts by inducing starch degradation through the transcription factor HY5 [143]. Induction of transcription of genes associated with starch degradation under the influence of blue radiation in CRY1a- or HY5-overexpressing plants was also confirmed.
It is known that in tomato, the R2R3-MYB group of factors regulating anthocyanin biosynthesis is represented by AN (ANANTHA) genes. Currently, their biological function is being actively clarified. For example, by generating loss-of-function mutants of AN2, the authors identified it as a positive regulator of anthocyanin biosynthesis in tomato vegetative tissues [144]. In addition to reduced anthocyanin content, the mutants had a dwarf phenotype. Overexpression of AN2 resulted in changes in multiple fruit qualities [145]. Thus, increased production of ethylene and increased content of anthocyanins, phenols, and flavonoids were observed. The content of aromatic volatiles such as aldehydes, phenylpropanoid derivatives, and terpene volatiles was also increased in these fruits. Thus, AN2 was shown here to regulate the transcription of genes in several metabolic pathways. Additionally, it was found that loss-of-function mutations in the AN2 ortholog in wild tomato impair anthocyanin synthesis [146]. Overexpression of ANT1 in tomatoes enriched the anthocyanins in leaves, contributing to more intense light absorption in the blue and red spectrum [147]. However, introducing knockout mutations into the AN2-like gene rather than ANT1 (ANTHOCYANIN) essentially eliminates the accumulation of anthocyanins [149,150]. It was found that AN2-like activated the expression of DFR; however, when AN1 was knocked out, anthocyanin pigmentation in the fruits was also eliminated. The AN2-like antagonist is the R3-MYB protein MYBATV. Meanwhile, a similar conclusion regarding MYBATV was made earlier [151]. It can be summarized that HY5 activates AN2-like, promotes the expression of AN1 and MYBATV, and MYBATV protein competes with AN2-like for binding to AN1 and thereby negatively regulates anthocyanin biosynthesis. Moreover, in [152], overexpression of AN2-like was found to increase jasmonic acid accumulation, activate the defense signaling pathway against Botrytis cinerea, and also increase fruit shelf life by inhibiting the expression of genes associated with the modification cell wall.
The previously mentioned dihydroflavonol 4-reductase (DFR) is involved in the reduction of dihydroflavonols to leukoanthocyanidins during the synthesis of the pigments pelargonidin, cyanidin, and delphinidin. The DFR gene in the tomato genome is represented by a single copy, which prompted its use in developing a natural genome editing marker based on homologous recombination with restoration of the DFR function [153]. DFR expression is also regulated by BBX20, which binds to its promoter region to activate expression [154].
6. System of regulation of tomato fruit ripening process
As a result of our literature review, we have presented a putative model of ripening factor regulatory pathways (Figure 4). We recognize five components of the maturation regulation system: transcription factors, hormones, epigenetics, external stimuli, and ncRNAs.
A significant contribution to regulation is provided by the ethylene-dependent pathway involving important polycistronic regulators like ethylene-sensitive genes, ethylene response genes, and the MADS-RIN complex. The fact that there are many additional factors with which RIN directly interacts suggests the existence of various ad hoc regulatory complexes consisting of several units of transcription factors. Quite often, functional redundancy is observed for them. There are 3 possible relationships of transcription factors: redundancy, additivity, and dependency. Redundancy is manifested in the functional identity of transcription factors. Additivity is associated with the provision of function through the joint contribution of each element. Direct dependence involves activation or repression of the role of one factor only after interaction with another. Moreover, autocatalytic regulation of the participants of regulatory cascades is possible.
As practice shows, turning off the function of any of the factors does not interrupt the entire cascade of ripening gene regulation but only leads to a delay phenotype. This is explained by the mechanism of gene compensation, i.e., the existence of homologs of the nonfunctional factor and/or other regulatory pathways. Indeed, we show that there are other regulators of the ethylene pathway, such as factors of the brassinosteroid and abscisic pathways. Although SA/JA pathways are absent in the proposed scheme, we do not exclude the presence of SA/JA regulators as additional maturation factors. Nevertheless, the number of studies aimed at clarifying their role in these processes does not allow us to conclude that they contribute significantly to the regulation of ripening-related genes.
Activation or deactivation of genes involved in ripening regulation can be mediated epigenetically, as discussed previously with specific examples. RIN-mediated regulation also requires interaction with promoters of lncRNAs, which are regulators of other genes, including those associated with maturation. This provides the so-called ethylene-independent regulatory pathway of maturation genes. Because epigenetic regulation and lncRNA regulation are potentially applicable to each element of regulatory cascades, their display on the scheme is redundant.
7. Future Prospects and Challenges
Significant progress has recently been made in understanding signal transduction systems and processes. The discovery of the function of genes and their regulatory systems in tomato ripening process can allow breeding tomatoes with an increased number of fruits and improved nutritional properties.
Although today there are attempts to generalize the cross-relationship of cascades of hormonal signaling pathways [155,156,157], they are only sometimes consistent with each other. They are united by the general concept of the participation of gibberellin- and auxin-sensitive genes at the stages of flowering, fertilization, and early fruit development, and ethylene-sensitive genes at the direct ripening of tomato fruits. Genes of mediator factors ensure the regulation of these processes. Of course, the proposed concepts still need to be completed. Thus, it was shown that ABA and BS are involved in all stages of tomato reproductive processes. It was also demonstrated with RIN and CNR that established hypotheses are not always perfect. The expanding understanding of the influence of epigenetic changes, microRNAs, and external factors on signaling regulation is also worth mentioning.
All this reminds us of a puzzle: to start putting it together, you need to find the edge of the image; otherwise, it can take infinite time to compare individual elements. But signaling pathways do not have “edges”; they must be created manually. One of the modern approaches to solving problems in this area is the use of a wide range of genetic engineering and bioengineering methods, including, among other things, the collection and processing of bioinformatics data, new sequencing technologies, targeted genome editing using CRISPR technology [158,159,160,161,162,163,164,165], use of base and prime editing for precision gene correction [166,167,168], creation of unique and simple markers for detection of transgenes [169], application transcriptomics, metabolomics and other omics technologies.
However, even here, some difficulties arise. For example, orthologous genes often have different regulatory mechanisms and are unreliable predictors of expression in related species. In addition, not all studies are field-validated, which reduces the significance of the data obtained using transgenic plants. Ideal and, at the same time, simplified laboratory conditions cannot determine all the possible subtleties of gene expression and their regulation. Moreover, changes in the expression of regulatory genes do not always entail significant changes in the transcriptome due to the possible presence in the genome of paralogs of the genes being studied. For genes higher in the regulatory cascades, pleiotropic changes occur. It is worth mentioning that the reproduction processes are discrete and appear in various tissues with excellent synchronization, determined by the life cycle stage, so their identification and description are difficult.
Undoubtedly, the existing conclusions about the regulation of plant life cycle processes obtained on model objects such as the tomato, although extensive, still need to be completed. In the future, genetic approaches will continue to make essential contributions to identifying new candidate genes involved in tomato reproductive signaling cascades. This may open up a broader cluster of regulatory signaling networks involving currently unknown factors and stimuli.
As for the perspectives of the practical application of the study of the considered ripening-related genes, these are improving fruit quality due to increased nutrient content, accelerated ripening, prolonged shelf life, and much more.
8. Conclusions
This review examined recent advances in studying tomato ripening factors using various genetic approaches. Modern research into the factors of tomato fruit ripening continues to expand our understanding of the molecular basis and physiological mechanisms of these processes, which has significant applications for improving breeding methods and growing new varieties with enhanced phenotypic traits that meet the requirements of the modern agricultural industry and consumer demand.
The abundance of current scientific reports cited in this review article reflects the relative ease of studying tomato and potential biotechnological improvements and the breadth of approaches and methods. Despite significant advances, a large number of biochemical pathways, the involvement of hundreds of genes in the fruit ripening process, and fine regulation involving transcription factors and microRNAs, at present, it is premature to talk about a complete understanding of the described processes. The complex new data presented in the review and proposed model of gene regulation will allow us to understand the principle of tomato fruit ripening better and complement the overall knowledge picture.
Author Contributions
Conceptualization, D.B. and V.T.; writing—original draft preparation, D.B.; writing—review and editing, V.T. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Russian Science Foundation, grant No. 22-14-00118.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Wang, R.; Tavano, E.C. da R.; Lammers, M.; Martinelli, A.P.; Angenent, G.C.; de Maagd, R.A. Re-Evaluation of Transcription Factor Function in Tomato Fruit Development and Ripening with CRISPR/Cas9-Mutagenesis. Sci. Rep. 2019, 9. [CrossRef]
- Zhao, X.; Yuan, X.; Chen, S.; Meng, L.; Fu, D. Role of the Tomato TAGL1 Gene in Regulating Fruit Metabolites Elucidated Using RNA Sequence and Metabolomics Analyses. PLoS ONE 2018, 13, e0199083. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Yuan, X.; Chen, S.; Fu, D.-Q.; Jiang, C.-Z. Metabolomic and Transcriptomic Analyses Reveal That a MADS-Box Transcription Factor TDR4 Regulates Tomato Fruit Quality. Front. Plant Sci. 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Ito, Y.; Nishizawa-Yokoi, A.; Endo, M.; Mikami, M.; Shima, Y.; Nakamura, N.; Kotake-Nara, E.; Kawasaki, S.; Toki, S. Re-Evaluation of the Rin Mutation and the Role of RIN in the Induction of Tomato Ripening. Nature Plants 2017, 3, 866–874. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Xu, H.; Ju, Z.; Cao, D.; Zhu, H.; Fu, D.; Grierson, D.; Qin, G.; Luo, Y.; Zhu, B. The RIN-MC Fusion of MADS-Box Transcription Factors Has Transcriptional Activity and Modulates Expression of Many Ripening Genes. Plant Physiol. 2017, 176, 891–909. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Zhu, B.; Pirrello, J.; Xu, C.; Zhang, B.; Bouzayen, M.; Chen, K.; Grierson, D. Roles of RIN and Ethylene in Tomato Fruit Ripening and Ripening-associated Traits. New Phytol. 2019, 226, 460–475. [Google Scholar] [CrossRef] [PubMed]
- Upadhyay, R.K.; Tucker, M.L.; Mattoo, A.K. Ethylene and RIPENING INHIBITOR Modulate Expression of SlHSP17.7A, B Class I Small Heat Shock Protein Genes During Tomato Fruit Ripening. Front. Plant Sci. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Ito, Y.; Sekiyama, Y.; Nakayama, H.; Nishizawa-Yokoi, A.; Endo, M.; Shima, Y.; Nakamura, N.; Kotake-Nara, E.; Kawasaki, S.; Hirose, S.; et al. Allelic Mutations in the Ripening-Inhibitor Locus Generate Extensive Variation in Tomato Ripening. Plant Physiol. 2020, 183, 80–95. [Google Scholar] [CrossRef]
- Yu, T.; Tzeng, D.T.W.; Li, R.; Chen, J.; Zhong, S.; Fu, D.; Zhu, B.; Luo, Y.; Zhu, H. Genome-Wide Identification of Long Non-Coding RNA Targets of the Tomato MADS Box Transcription Factor RIN and Function Analysis. Annals of Botany 2018, 123, 469–482. [Google Scholar] [CrossRef]
- Kang, J.; Gong, J.; Zhang, L.; Gao, Z.; Xie, Q.; Hu, Z.; Chen, G. A Novel E6-like Gene, E6-2, Affects Fruit Ripening in Tomato. Plant Science 2021, 313, 111066. [Google Scholar] [CrossRef]
- Osakabe, K.; Wada, N.; Miyaji, T.; Murakami, E.; Marui, K.; Ueta, R.; Hashimoto, R.; Abe-Hara, C.; Kong, B.; Yano, K.; et al. Genome Editing in Plants Using CRISPR Type I-D Nuclease. Commun. Biol. 2020, 3. [Google Scholar] [CrossRef] [PubMed]
- Niu, Q.; Wu, S.; Li, Y.; Yang, X.; Liu, P.; Xu, Y.; Lang, Z. Expanding the Scope of CRISPR/Cas9-mediated Genome Editing in Plants Using an xCas9 and Cas9-NG Hybrid. JIPB 2020, 62, 398–402. [Google Scholar] [CrossRef] [PubMed]
- Niu, Q.; Wu, S.; Xie, H.; Wu, Q.; Liu, P.; Xu, Y.; Lang, Z. Efficient A·T to G·C Base Conversions in Dicots Using Adenine Base Editors Expressed under the Tomato EF1α Promoter. Plant Biotechnol. J. 2022, 21, 5–7. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Wei, W.; Zhao, X.; Tan, X.; Fan, Z.; Zhang, Y.; Jing, Y.; Meng, L.; Zhu, B.; Zhu, H.; et al. A NAC Transcription Factor, NOR-Like1, Is a New Positive Regulator of Tomato Fruit Ripening. Hortic. Res. 2018, 5. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Chen, H.; Nonaka, S.; Sato-Izawa, K.; Kusano, M.; Ezura, H. Ethylene Biosynthesis Controlled by NON-RIPENING: A Regulatory Conflict between Wounding and Ripening. Plant Physiol. Biochem. 2018, 132, 720–726. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Wei, W.; Fan, Z.; Zhao, X.; Zhang, Y.; Jing, Y.; Zhu, B.; Zhu, H.; Shan, W.; Chen, J.; et al. Re-Evaluation of the nor Mutation and the Role of the NAC-NOR Transcription Factor in Tomato Fruit Ripening. J. Exp. Bot. 2020, 71, 3560–3574. [Google Scholar] [CrossRef]
- Yu, Q.; Wang, B.; Li, N.; Tang, Y.; Yang, S.; Yang, T.; Xu, J.; Guo, C.; Yan, P.; Wang, Q.; et al. CRISPR/Cas9-Induced Targeted Mutagenesis and Gene Replacement to Generate Long-Shelf Life Tomato Lines. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef] [PubMed]
- Cao, X.; Wei, C.; Duan, W.; Gao, Y.; Kuang, J.; Liu, M.; Chen, K.; Klee, H.; Zhang, B. Transcriptional and Epigenetic Analysis Reveals That NAC Transcription Factors Regulate Fruit Flavor Ester Biosynthesis. Plant J. 2021, 106, 785–800. [Google Scholar] [CrossRef]
- Jian, W.; Zheng, Y.; Yu, T.; Cao, H.; Chen, Y.; Cui, Q.; Xu, C.; Li, Z. SlNAC6, A NAC Transcription Factor, Is Involved in Drought Stress Response and Reproductive Process in Tomato. J. Plant Physiol. 2021, 264, 153483. [Google Scholar] [CrossRef]
- Gong, J.; Zeng, Y.; Meng, Q.; Guan, Y.; Li, C.; Yang, H.; Zhang, Y.; Ampomah-Dwamena, C.; Liu, P.; Chen, C.; et al. Red Light-Induced Kumquat Fruit Coloration Is Attributable to Increased Carotenoid Metabolism Regulated by FcrNAC22. J. Exp. Bot. 2021, 72, 6274–6290. [Google Scholar] [CrossRef]
- Gao, Y.; Fan, Z.; Zhang, Q.; Li, H.; Liu, G.; Jing, Y.; Zhang, Y.; Zhu, B.; Zhu, H.; Chen, J.; et al. A Tomato NAC Transcription Factor, SlNAM1, Positively Regulates Ethylene Biosynthesis and the Onset of Tomato Fruit Ripening. Plant J. 2021, 108, 1317–1331. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.K.; Vishwakarma, A.; Kenea, H.D.; Galsurker, O.; Cohen, H.; Aharoni, A.; Arazi, T. CRISPR/Cas9 Mutants of Tomato MICRORNA164 Genes Uncover Their Functional Specialization in Development. Plant Physiol. 2021, 187, 1636–1652. [Google Scholar] [CrossRef] [PubMed]
- Lin, D.; Zhu, X.; Qi, B.; Gao, Z.; Tian, P.; Li, Z.; Lin, Z.; Zhang, Y.; Huang, T. SlMIR164A Regulates Fruit Ripening and Quality by Controlling SlNAM2 and SlNAM3 in Tomato. Plant Biotechnol. J. 2022, 20, 1456–1469. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Tang, M.; Huang, Z.; Song, J.; Xu, J.; Ahammed, G.J.; Yu, J.; Zhou, Y. The miR164a-NAM3 Module Confers Cold Tolerance by Inducing Ethylene Production in Tomato. Plant J. 2022, 111, 440–456. [Google Scholar] [CrossRef] [PubMed]
- Yuan, S.; Kawasaki, S.; Abdellatif, I.M.Y.; Nishida, K.; Kondo, A.; Ariizumi, T.; Ezura, H.; Miura, K. Efficient Base Editing in Tomato Using a Highly Expressed Transient System. Plant Cell Rep. 2021, 40, 667–676. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Zhang, Y.; Tureckova, V.; Xue, G.-P.; Fernie, A.R.; Mueller-Roeber, B.; Balazadeh, S. The NAC Transcription Factor SlNAP2 Regulates Leaf Senescence and Fruit Yield in Tomato. Plant Physiol. 2018, 177, 1286–1302. [Google Scholar] [CrossRef]
- Forlani, S.; Cozzi, C.; Rosa, S.; Tadini, L.; Masiero, S.; Mizzotti, C. HEBE, a Novel Positive Regulator of Senescence in Solanum lycopersicum. Sci Rep 2020, 10. [Google Scholar] [CrossRef]
- Gao, Y.; Zhu, N.; Zhu, X.; Wu, M.; Jiang, C.-Z.; Grierson, D.; Luo, Y.; Shen, W.; Zhong, S.; Fu, D.-Q.; et al. Diversity and Redundancy of the Ripening Regulatory Networks Revealed by the fruitENCODE and the New CRISPR/Cas9 CNR and NOR Mutants. Hortic. Res. 2019, 6. [Google Scholar] [CrossRef] [PubMed]
- Jiang, G.; Zeng, J.; Li, Z.; Song, Y.; Yan, H.; He, J.; Jiang, Y.; Duan, X. Redox Regulation of the NOR Transcription Factor Is Involved in the Regulation of Fruit Ripening in Tomato. Plant Physiol. 2020, 183, 671–685. [Google Scholar] [CrossRef]
- Lai, T.; Wang, X.; Ye, B.; Jin, M.; Chen, W.; Wang, Y.; Zhou, Y.; Blanks, A.M.; Gu, M.; Zhang, P.; et al. Molecular and Functional Characterization of the SBP-Box Transcription Factor SPL-CNR in Tomato Fruit Ripening and Cell Death. J. Exp. Bot. 2020, 71, 2995–3011. [Google Scholar] [CrossRef]
- Yin, W.; Hu, Z.; Cui, B.; Guo, X.; Hu, J.; Zhu, Z.; Chen, G. Suppression of the MADS-Box Gene SlMBP8 Accelerates Fruit Ripening of Tomato (Solanum lycopersicum). Plant Physiol. Biochem. 2017, 118, 235–244. [Google Scholar] [CrossRef] [PubMed]
- Yin, W.; Yu, X.; Chen, G.; Tang, B.; Wang, Y.; Liao, C.; Zhang, Y.; Hu, Z. Suppression of SlMBP15 Inhibits Plant Vegetative Growth and Delays Fruit Ripening in Tomato. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Wang, X.; Li, C.; Li, H.; Zhang, J.; Ye, Z. Silencing GRAS2 Reduces Fruit Weight in Tomato. JIPB 2018, 60, 498–513. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Shi, Y.; Su, D.; Lu, W.; Li, Z. SlGRAS4 Accelerates Fruit Ripening by Regulating Ethylene Biosynthesis Genes and SlMADS1 in Tomato. Hortic. Res. 2021, 8. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Peng, S.; Xian, Z.; Lin, D.; Hu, G.; Yang, L.; Ren, M.; Li, Z. Overexpression of a Tomato miR171 Target Gene SlGRAS24 Impacts Multiple Agronomical Traits via Regulating Gibberellin and Auxin Homeostasis. Plant Biotechnol. J. 2016, 15, 472–488. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Li, Y.; Fan, S.; Wen, T.; Wang, M.; Zhang, L.; Zhao, L. The Transcription Factor WRKY32 Affects Tomato Fruit Colour by Regulating YELLOW FRUITED-TOMATO 1, a Core Component of Ethylene Signal Transduction. J. Exp. Bot. 2021, 72, 4269–4282. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Zhang, X.; Wang, L.; Tian, Y.; Jia, N.; Chen, S.; Shi, N.; Huang, X.; Zhou, C.; Yu, Y.; et al. Regulation of Ethylene-Responsive SlWRKYs Involved in Color Change during Tomato Fruit Ripening. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Lang, Z.; Wang, Y.; Tang, K.; Tang, D.; Datsenka, T.; Cheng, J.; Zhang, Y.; Handa, A.K.; Zhu, J.-K. Critical Roles of DNA Demethylation in the Activation of Ripening-Induced Genes and Inhibition of Ripening-Repressed Genes in Tomato Fruit. Proc. Natl. Acad. Sci. USA 2017, 114. [Google Scholar] [CrossRef]
- Li, Z.; Pi, Y.; Fan, J.; Yang, X.; Zhai, C.; Chen, H.; Wang, F.; Ding, J.; Gu, T.; Li, Y.; et al. High Mobility Group A3 Enhances Transcription of the DNA Demethylase Gene SlDML2 to Promote Tomato Fruit Ripening. Plant Physiol. 2022, 189, 315–328. [Google Scholar] [CrossRef]
- Hollwey, E.; Out, S.; Watson, M.R.; Heidmann, I.; Meyer, P. TET3-Mediated Demethylation in Tomato Activates Expression of a CETS Gene That Stimulates Vegetative Growth. Plant Direct 2017, 1. [Google Scholar] [CrossRef]
- Yang, Y.; Tang, K.; Datsenka, T.U.; Liu, W.; Lv, S.; Lang, Z.; Wang, X.; Gao, J.; Wang, W.; Nie, W.; et al. Critical Function of DNA Methyltransferase 1 in Tomato Development and Regulation of the DNA Methylome and Transcriptome. JIPB 2019, 61, 1224–1242. [Google Scholar] [CrossRef] [PubMed]
- Yao, M.; Chen, W.; Kong, J.; Zhang, X.; Shi, N.; Zhong, S.; Ma, P.; Gallusci, P.; Jackson, S.; Liu, Y.; et al. METHYLTRANSFERASE1 and Ripening Modulate Vivipary during Tomato Fruit Development. Plant Physiol. 2020, 183, 1883–1897. [Google Scholar] [CrossRef] [PubMed]
- Wen, Y.X.; Wang, J.Y.; Zhu, H.H.; Han, G.H.; Huang, R.N.; Huang, L.; Hong, Y.G.; Zheng, S.J.; Yang, J.L.; Chen, W.W. Potential Role of Domains Rearranged Methyltransferase7 in Starch and Chlorophyll Metabolism to Regulate Leaf Senescence in Tomato. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef] [PubMed]
- Jia, H.; Jia, H.; Lu, S.; Zhang, Z.; Su, Z.; Sadeghnezhad, E.; Li, T.; Xiao, X.; Wang, M.; Pervaiz, T.; et al. DNA and Histone Methylation Regulates Different Types of Fruit Ripening by Transcriptome and Proteome Analyses. J. Agric. Food Chem. 2022, 70, 3541–3556. [Google Scholar] [CrossRef] [PubMed]
- Corem, S.; Doron-Faigenboim, A.; Jouffroy, O.; Maumus, F.; Arazi, T.; Bouché, N. Redistribution of CHH Methylation and Small Interfering RNAs across the Genome of Tomato Ddm1 Mutants. Plant Cell 2018, 30, 1628–1644. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Zhao, J.; Chen, Z.; Qiao, J.; Zhang, Y.; Shen, H.; Hu, Z. CRISPR/Cas9-Targeted Mutagenesis ofSlCMT4Causes Changes in Plant Architecture and Reproductive Organs in Tomato. Hortic. Res. 2022, 9. [Google Scholar] [CrossRef] [PubMed]
- Hu, G.; Huang, B.; Wang, K.; Frasse, P.; Maza, E.; Djari, A.; Benhamed, M.; Gallusci, P.; Li, Z.; Zouine, M.; et al. Histone Posttranslational Modifications Rather than DNA Methylation Underlie Gene Reprogramming in Pollination-dependent and Pollination-independent Fruit Set in Tomato. New Phytol. 2020, 229, 902–919. [Google Scholar] [CrossRef] [PubMed]
- Bvindi, C.; Tang, L.; Lee, S.; Patrick, R.M.; Yee, Z.R.; Mengiste, T.; Li, Y. Histone Methyltransferases SDG33 and SDG34 Regulate Organ-Specific Nitrogen Responses in Tomato. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef]
- Li, Z.; Jiang, G.; Liu, X.; Ding, X.; Zhang, D.; Wang, X.; Zhou, Y.; Yan, H.; Li, T.; Wu, K.; et al. Histone Demethylase SlJMJ6 Promotes Fruit Ripening by Removing H3K27 Methylation of Ripening-related Genes in Tomato. New Phytol. 2020, 227, 1138–1156. [Google Scholar] [CrossRef]
- Ding, X.; Zhang, D.; Gu, D.; Li, Z.; Liang, H.; Zhu, H.; Jiang, Y.; Duan, X. The Histone H3K27 Demethylase SlJMJ4 Promotes Dark- and ABA-Induced Leaf Senescence in Tomato. Hortic. Res. 2022, 9. [Google Scholar] [CrossRef]
- Yang, X.; Zhang, X.; Yang, Y.; Zhang, H.; Zhu, W.; Nie, W.-F. The Histone Variant Sl_H2A.Z Regulates Carotenoid Biosynthesis and Gene Expression during Tomato Fruit Ripening. Hortic. Res. 2021, 8. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.-E.; Hu, Z.; Li, F.; Zhang, L.; Yu, X.; Tang, B.; Chen, G. Silencing of Histone Deacetylase SlHDT3 Delays Fruit Ripening and Suppresses Carotenoid Accumulation in Tomato. Plant Sci. 2017, 265, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.-E.; Hu, Z.; Zhu, M.; Li, F.; Zhu, Z.; Lu, Y.; Chen, G. The Tomato Histone Deacetylase SlHDA1 Contributes to the Repression of Fruit Ripening and Carotenoid Accumulation. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.-E. Histone Deacetylase Gene SlHDT1 Regulates Tomato Fruit Ripening by Affecting Carotenoid Accumulation and Ethylene Biosynthesis. Plant Sci. 2022, 318, 111235. [Google Scholar] [CrossRef]
- Hawar, A.; Xiong, S.; Yang, Z.; Sun, B. Histone Acetyltransferase SlGCN5 Regulates Shoot Meristem and Flower Development in Solanum lycopersicum. Front. Plant Sci. 2022, 12. [Google Scholar] [CrossRef] [PubMed]
- Bollier, N.; Sicard, A.; Leblond, J.; Latrasse, D.; Gonzalez, N.; Gévaudant, F.; Benhamed, M.; Raynaud, C.; Lenhard, M.; Chevalier, C.; et al. At-MINI ZINC FINGER2 and Sl-INHIBITOR OF MERISTEM ACTIVITY, a Conserved Missing Link in the Regulation of Floral Meristem Termination in Arabidopsis and Tomato. Plant Cell 2018, 30, 83–100. [Google Scholar] [CrossRef] [PubMed]
- Ai, G.; Huang, R.; Zhang, D.; Li, M.; Li, G.; Li, W.; Ahiakpa, J.K.; Wang, Y.; Hong, Z.; Zhang, J. SlGH3.15, a Member of the GH3 Gene Family, Regulates Lateral Root Development and Gravitropism Response by Modulating Auxin Homeostasis in Tomato. Plant Sci. 2023, 330, 111638. [Google Scholar] [CrossRef]
- Sun, M.; Li, H.; Li, Y.; Xiang, H.; Liu, Y.; He, Y.; Qi, M.; Li, T. Tomato YABBY2b Controls Plant Height through Regulating Indole-3-Acetic Acid-Amido Synthetase (GH3.8) Expression. Plant Scie. 2020, 297, 110530. [Google Scholar] [CrossRef]
- Chen, X.; Liao, D.; Yang, X.; Ji, M.; Wang, S.; Gu, M.; Chen, A.; Xu, G. Three Cis-Regulatory Motifs, AuxRE, MYCRS1 and MYCRS2, Are Required for Modulating the Auxin- and Mycorrhiza-Responsive Expression of a Tomato GH3 Gene. Plant Cell Physiol. 2017, 58, 770–778. [Google Scholar] [CrossRef]
- Chen, X.; Chen, J.; Liao, D.; Ye, H.; Li, C.; Luo, Z.; Yan, A.; Zhao, Q.; Xie, K.; Li, Y.; et al. Auxin-mediated Regulation of Arbuscular Mycorrhizal Symbiosis: A Role of SlGH3.4 in Tomato. Plant Cell Environ. 2021, 45, 955–968. [Google Scholar] [CrossRef]
- Sravankumar, T.; Akash; Naik, N.; Kumar, R. A Ripening-Induced SlGH3-2 Gene Regulates Fruit Ripening via Adjusting Auxin-Ethylene Levels in Tomato (Solanum lycopersicum L.). Plant Mol. Biol. 2018, 98, 455–469. [CrossRef] [PubMed]
- Shi, Z.; Jiang, Y.; Han, X.; Liu, X.; Cao, R.; Qi, M.; Xu, T.; Li, T. SlPIN1 Regulates Auxin Efflux to Affect Flower Abscission Process. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef] [PubMed]
- Li, A.; Chen, G.; Yu, X.; Zhu, Z.; Zhang, L.; Zhou, S.; Hu, Z. The Tomato MADS-Box Gene SlMBP9 Negatively Regulates Lateral Root Formation and Apical Dominance by Reducing Auxin Biosynthesis and Transport. Plant Cell Rep. 2019, 38, 951–963. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Zhang, Y.; Feng, Q.; Qin, L.; Pan, C.; Lamin-Samu, A.T.; Lu, G. Tomato AUXIN RESPONSE FACTOR 5 Regulates Fruit Set and Development via the Mediation of Auxin and Gibberellin Signaling. Sci. Rep. 2018, 8. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; Mei, L.; Wu, M.; Wei, W.; Shan, W.; Gong, Z.; Zhang, Q.; Yang, F.; Yan, F.; Zhang, Q.; et al. SlARF10, an Auxin Response Factor, Is Involved in Chlorophyll and Sugar Accumulation during Tomato Fruit Development. J. Exp. Bot. 2018. [Google Scholar] [CrossRef]
- Israeli, A.; Capua, Y.; Shwartz, I.; Tal, L.; Meir, Z.; Levy, M.; Bar, M.; Efroni, I.; Ori, N. Multiple Auxin-Response Regulators Enable Stability and Variability in Leaf Development. Curr. Biol. 2019, 29, 1746–1759.e5. [Google Scholar] [CrossRef] [PubMed]
- Abe-Hara, C.; Yamada, K.; Wada, N.; Ueta, R.; Hashimoto, R.; Osakabe, K.; Osakabe, Y. Effects of the Sliaa9 Mutation on Shoot Elongation Growth of Tomato Cultivars. Front. Plant Sci. 2021, 12. [Google Scholar] [CrossRef]
- Wu, L.; Tian, Z.; Zhang, J. Functional Dissection of Auxin Response Factors in Regulating Tomato Leaf Shape Development. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Damodharan, S.; Corem, S.; Gupta, S.K.; Arazi, T. Tuning of SlARF10A Dosage by sly-miR160a Is Critical for Auxin-mediated Compound Leaf and Flower Development. Plant J. 2018, 96, 855–868. [Google Scholar] [CrossRef]
- Tomlinson, L.; Yang, Y.; Emenecker, R.; Smoker, M.; Taylor, J.; Perkins, S.; Smith, J.; MacLean, D.; Olszewski, N.E.; Jones, J.D.G. Using CRISPR/Cas9 Genome Editing in Tomato to Create a Gibberellin-responsive Dominant Dwarf DELLA Allele. Plant Biotechnol. J. 2018, 17, 132–140. [Google Scholar] [CrossRef]
- Shinozaki, Y.; Ezura, K.; Hu, J.; Okabe, Y.; Bénard, C.; Prodhomme, D.; Gibon, Y.; Sun, T.; Ezura, H.; Ariizumi, T. Identification and Functional Study of a Mild Allele of SlDELLA Gene Conferring the Potential for Improved Yield in Tomato. Sci. Rep. 2018, 8. [Google Scholar] [CrossRef] [PubMed]
- Cheng, W.; Yin, S.; Tu, Y.; Mei, H.; Wang, Y.; Yang, Y. SlCAND1, Encoding Cullin-Associated Nedd8-Dissociated Protein 1, Regulates Plant Height, Flowering Time, Seed Germination, and Root Architecture in Tomato. Plant Mol. Biol. 2020, 102, 537–551. [Google Scholar] [CrossRef] [PubMed]
- Illouz-Eliaz, N.; Ramon, U.; Shohat, H.; Blum, S.; Livne, S.; Mendelson, D.; Weiss, D. Multiple Gibberellin Receptors Contribute to Phenotypic Stability under Changing Environments. Plant Cell 2019, 31, 1506–1519. [Google Scholar] [CrossRef] [PubMed]
- Nir, I.; Shohat, H.; Panizel, I.; Olszewski, N.; Aharoni, A.; Weiss, D. The Tomato DELLA Protein PROCERA Acts in Guard Cells to Promote Stomatal Closure. Plant Cell 2017, 29, 3186–3197. [Google Scholar] [CrossRef] [PubMed]
- Shohat, H.; Illouz-Eliaz, N.; Kanno, Y.; Seo, M.; Weiss, D. The Tomato DELLA Protein PROCERA Promotes Abscisic Acid Responses in Guard Cells by Upregulating an Abscisic Acid Transporter. Plant Physiol. 2020, 184, 518–528. [Google Scholar] [CrossRef] [PubMed]
- Silva, G.F.F.; Silva, E.M.; Correa, J.P.O.; Vicente, M.H.; Jiang, N.; Notini, M.M.; Junior, A.C.; De Jesus, F.A.; Castilho, P.; Carrera, E.; et al. Tomato Floral Induction and Flower Development Are Orchestrated by the Interplay between Gibberellin and Two Unrelated microRNA-controlled Modules. New Phytol. 2018, 221, 1328–1344. [Google Scholar] [CrossRef]
- Naeem, M.; Waseem, M.; Zhu, Z.; Zhang, L. Downregulation of SlGRAS15 Manipulates Plant Architecture in Tomato (Solanum lycopersicum). Dev. Genes. Evol. 2019, 230, 1–12. [Google Scholar] [CrossRef]
- Zhu, Z.; Liang, H.; Chen, G.; Li, F.; Wang, Y.; Liao, C.; Hu, Z. The bHLH Transcription Factor SlPRE2 Regulates Tomato Fruit Development and Modulates Plant Response to Gibberellin. Plant Cell Rep. 2019, 38, 1053–1064. [Google Scholar] [CrossRef]
- Zhu, Z.; Chen, G.; Guo, X.; Yin, W.; Yu, X.; Hu, J.; Hu, Z. Overexpression of SlPRE2, an Atypical bHLH Transcription Factor, Affects Plant Morphology and Fruit Pigment Accumulation in Tomato. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Li, J.; Gong, J.; Zhang, L.; Shen, H.; Chen, G.; Xie, Q.; Hu, Z. Overexpression of SlPRE5, an Atypical bHLH Transcription Factor, Affects Plant Morphology and Chlorophyll Accumulation in Tomato. Journal of Plant Physiol. 2022, 273, 153698. [Google Scholar] [CrossRef]
- Glanz-Idan, N.; Lach, M.; Tarkowski, P.; Vrobel, O.; Wolf, S. Delayed Leaf Senescence by Upregulation of Cytokinin Biosynthesis Specifically in Tomato Roots. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef]
- Gan, L.; Song, M.; Wang, X.; Yang, N.; Li, H.; Liu, X.; Li, Y. Cytokinins Are Involved in Regulation of Tomato Pericarp Thickness and Fruit Size. Hortic. Res. 2022, 9. [Google Scholar] [CrossRef] [PubMed]
- Jia, M.; Du, P.; Ding, N.; Zhang, Q.; Xing, S.; Wei, L.; Zhao, Y.; Mao, W.; Li, J.; Li, B.; et al. Two FERONIA-Like Receptor Kinases Regulate Apple Fruit Ripening by Modulating Ethylene Production. Front. Plant Sci. 2017, 8. [Google Scholar] [CrossRef]
- Yanping, Z.; Yuqing, H.; Chen, W.; Qian, M.; Songtao, J.; Xudong, Z.; Ting, Z.; Kekun, Z.; Haifeng, J.; Tariq, P.; et al. Characterization and Identification of PpEIN3 during the Modulation of Fruit Ripening Process by Ectopic Expressions in Tomato. Plant Genome 2019, 12. [Google Scholar] [CrossRef]
- Wang, H.; Zhang, H.; Liang, F.; Cong, L.; Song, L.; Li, X.; Zhai, R.; Yang, C.; Wang, Z.; Ma, F.; et al. PbEIL1 Acts Upstream ofPbCysp1to Regulate Ovule Senescence in Seedless Pear. Hortic. Res. 2021, 8. [Google Scholar] [CrossRef]
- Althiab-Almasaud, R.; Chen, Y.; Maza, E.; Djari, A.; Frasse, P.; Mollet, J.; Mazars, C.; Jamet, E.; Chervin, C. Ethylene Signaling Modulates Tomato Pollen Tube Growth through Modifications of Cell Wall Remodeling and Calcium Gradient. Plant J. 2021, 107, 893–908. [Google Scholar] [CrossRef]
- Chen, Y.; Hu, G.; Rodriguez, C.; Liu, M.; Binder, B.M.; Chervin, C. Roles of SlETR7, a Newly Discovered Ethylene Receptor, in Tomato Plant and Fruit Development. Hortic. Res. 2020, 7. [Google Scholar] [CrossRef]
- Shimatani, Z.; Kashojiya, S.; Takayama, M.; Terada, R.; Arazoe, T.; Ishii, H.; Teramura, H.; Yamamoto, T.; Komatsu, H.; Miura, K.; et al. Targeted Base Editing in Rice and Tomato Using a CRISPR-Cas9 Cytidine Deaminase Fusion. Nat. Biotechnol. 2017, 35, 441–443. [Google Scholar] [CrossRef]
- Kashojiya, S.; Lu, Y.; Takayama, M.; Komatsu, H.; Minh, L.H.T.; Nishida, K.; Shirasawa, K.; Miura, K.; Nonaka, S.; Masuda, J.; et al. Modification of Tomato Breeding Traits and Plant Hormone Signaling by Target-AID, the Genome-Editing System Inducing Efficient Nucleotide Substitution. Hortic. Res. 2022, 9. [Google Scholar] [CrossRef]
- Guo, H.; Mao, M.; Deng, Y.; Sun, L.; Chen, R.; Cao, P.; Lai, J.; Zhang, Y.; Wang, C.; Li, C.; et al. Multi-Omics Analysis Reveals That SlERF.D6 Synergistically Regulates SGAs and Fruit Development. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef]
- Gambhir, P.; Singh, V.; Parida, A.; Raghuvanshi, U.; Kumar, R.; Sharma, A.K. Ethylene Response Factor ERF.D7 Activates Auxin Response Factor 2 Paralogs to Regulate Tomato Fruit Ripening. Plant Physiol. 2022, 190, 2775–2796. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Gao, M.; Zhao, Y.; Chen, Y.; Wu, L.; Yin, H.; Yang, J.; Xiong, S.; Wang, S.; Wang, J.; et al. LcERF19, an AP2/ERF Transcription Factor from Litsea Cubeba, Positively Regulates Geranial and Neral Biosynthesis. Hortic. Res. 2022, 9. [Google Scholar] [CrossRef]
- Li, G.; Wang, J.; Zhang, C.; Ai, G.; Zhang, D.; Wei, J.; Cai, L.; Li, C.; Zhu, W.; Larkin, R.M.; et al. L2, a Chloroplast Metalloproteinase, Regulates Fruit Ripening by Participating in Ethylene Autocatalysis under the Control of Ethylene Response Factors. J. Exp. Bot. 2021, 72, 7035–7048. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Feng, P.; Tang, B.; Hu, Z.; Xie, Q.; Zhou, S.; Chen, G. The AP2/ERF Transcription Factor SlERF.F5 Functions in Leaf Senescence in Tomato. Plant Cell Rep. 2022, 41, 1181–1195. [Google Scholar] [CrossRef]
- Hu, C.; Wu, S.; Li, J.; Dong, H.; Zhu, C.; Sun, T.; Hu, Z.; Foyer, C.H.; Yu, J. Herbivore-induced Ca2+ Signals Trigger a Jasmonate Burst by Activating ERF16-mediated Expression in Tomato. New Phytol. 2022, 236, 1796–1808. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Hu, K.; Wei, S.; Yao, G.; Zhang, H. ETHYLENE RESPONSE FACTORS 4.1/4.2 with an EAR Motif Repress Anthocyanin Biosynthesis in Red-Skinned Pears. Plant Physiol. 2023, 192, 1892–1912. [Google Scholar] [CrossRef] [PubMed]
- Liu, A.; Cheng, C. Pathogen-induced ERF68 Regulates Hypersensitive Cell Death in Tomato. Molecular Plant Pathology 2016, 18, 1062–1074. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Yuan, L.; Guo, W.; Wu, W. Transcription Factor TERF1 Promotes Seed Germination under Osmotic Conditions by Activating Gibberellin Acid Signaling. Plant Sci. 2022, 322, 111350. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Yang, H.; Tang, B.; Li, F.; Xie, Q.; Chen, G.; Hu, Z. The AP2/ERF Transcription Factor SlERF.J2 Functions in Hypocotyl Elongation and Plant Height in Tomato. Plant Cell Rep. 2022. [Google Scholar] [CrossRef]
- Chen, Y.; Cai, X.; Tang, B.; Xie, Q.; Chen, G.; Chen, X.; Hu, Z. SlERF.J2 Reduces Chlorophyll Accumulation and Inhibits Chloroplast Biogenesis and Development in Tomato Leaves. Plant Sci. 2023, 328, 111578. [Google Scholar] [CrossRef]
- Gupta, A.; Upadhyay, R.K.; Prabhakar, R.; Tiwari, N.; Garg, R.; Sane, V.A.; Sane, A.P. SlDREB3, a Negative Regulator of ABA Responses, Controls Seed Germination, Fruit Size and the Onset of Ripening in Tomato. Plant Sci. 2022, 319, 111249. [Google Scholar] [CrossRef] [PubMed]
- Fang, P.; Wang, Y.; Wang, M.; Wang, F.; Chi, C.; Zhou, Y.; Zhou, J.; Shi, K.; Xia, X.; Foyer, C.H.; et al. Crosstalk between Brassinosteroid and Redox Signaling Contributes to the Activation of CBF Expression during Cold Responses in Tomato. Antioxidants 2021, 10, 509. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Cao, J.-J.; Wang, K.-X.; Xia, X.-J.; Shi, K.; Zhou, Y.-H.; Yu, J.-Q.; Zhou, J. BZR1 Mediates Brassinosteroid-Induced Autophagy and Nitrogen Starvation in Tomato. Plant Physiol. 2018, 179, 671–685. [Google Scholar] [CrossRef] [PubMed]
- Sang, K.; Li, J.; Qian, X.; Yu, J.; Zhou, Y.; Xia, X. The APETALA2a/DWARF/BRASSINAZOLE-RESISTANT 1 Module Contributes to Carotenoid Synthesis in Tomato Fruits. Plant J. 2022, 112, 1238–1251. [Google Scholar] [CrossRef]
- Hu, S.; Liu, L.; Li, S.; Shao, Z.; Meng, F.; Liu, H.; Duan, W.; Liang, D.; Zhu, C.; Xu, T.; et al. Regulation of Fruit Ripening by the Brassinosteroid Biosynthetic Gene SlCYP90B3 via an Ethylene-Dependent Pathway in Tomato. Hortic. Res. 2020, 7. [Google Scholar] [CrossRef] [PubMed]
- Nie, S.; Huang, S.; Wang, S.; Cheng, D.; Liu, J.; Lv, S.; Li, Q.; Wang, X. Enhancing Brassinosteroid Signaling via Overexpression of Tomato (Solanum lycopersicum) SlBRI1 Improves Major Agronomic Traits. Front. Plant Sci. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Liu, J.; Zhao, T.; Du, C.; Nie, S.; Zhang, Y.; Lv, S.; Huang, S.; Wang, X. Modification of Threonine-1050 of SlBRI1 Regulates BR Signalling and Increases Fruit Yield of Tomato. BMC Plant Biol. 2019, 19. [Google Scholar] [CrossRef]
- Wang, D.; Yang, Z.; Wu, M.; Wang, W.; Wang, Y.; Nie, S. Enhanced Brassinosteroid Signaling via the Overexpression of SlBRI1 Positively Regulates the Chilling Stress Tolerance of Tomato. Plant Sci. 2022, 320, 111281. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.; Luo, B.; Hu, M.; Fu, S.; Liu, J.; Jiang, M.; Zhao, Y.; Huang, S.; Wang, S.; Wang, X. Brassinosteroid Signaling Downstream Suppressor BIN2 Interacts with SLFRIGIDA-LIKE to Induce Early Flowering in Tomato. IJMS 2022, 23, 11264. [Google Scholar] [CrossRef]
- Lee, J.; Han, S.; Lee, H.-Y.; Jeong, B.; Heo, T.-Y.; Hyun, T.K.; Kim, K.; Je, B.I.; Lee, H.; Shim, D.; et al. Brassinosteroids Facilitate Xylem Differentiation and Wood Formation in Tomato. Planta 2019, 249, 1391–1403. [Google Scholar] [CrossRef]
- Mori, K.; Lemaire-Chamley, M.; Jorly, J.; Carrari, F.; Conte, M.; Asamizu, E.; Mizoguchi, T.; Ezura, H.; Rothan, C. The Conserved Brassinosteroid-Related Transcription Factor BIM1a Negatively Regulates Fruit Growth in Tomato. J. Exp. Bot. 2020, 72, 1181–1197. [Google Scholar] [CrossRef] [PubMed]
- Yu, T.; Ai, G.; Xie, Q.; Wang, W.; Song, J.; Wang, J.; Tao, J.; Zhang, X.; Hong, Z.; Lu, Y.; et al. Regulation of Tomato Fruit Elongation by Transcription Factor BZR1.7 through Promotion of SUN Gene Expression. Hortic. Res. 2022, 9. [Google Scholar] [CrossRef] [PubMed]
- Xia, X.; Dong, H.; Yin, Y.; Song, X.; Gu, X.; Sang, K.; Zhou, J.; Shi, K.; Zhou, Y.; Foyer, C.H.; et al. Brassinosteroid Signaling Integrates Multiple Pathways to Release Apical Dominance in Tomato. Proc. Natl. Acad. Sci. U.S.A. 2021, 118. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Liu, L.; Liang, D.; Zhang, M.; Jia, C.; Qi, M.; Liu, Y.; Shao, Z.; Meng, F.; Hu, S.; et al. SlBES1 Promotes Tomato Fruit Softening through Transcriptional Inhibition of PMEU1. iScience 2021, 24, 102926. [Google Scholar] [CrossRef] [PubMed]
- Sierra-Orozco, E.; Shekasteband, R.; Illa-Berenguer, E.; Snouffer, A.; van der Knaap, E.; Lee, T.G.; Hutton, S.F. Identification and Characterization of GLOBE, a Major Gene Controlling Fruit Shape and Impacting Fruit Size and Marketability in Tomato. Hortic Res 2021, 8. [Google Scholar] [CrossRef] [PubMed]
- Hou, J.; Sun, Q.; Li, J.; Ahammed, G.J.; Yu, J.; Fang, H.; Xia, X. Glutaredoxin S25 and Its Interacting TGACG Motif-Binding Factor TGA2 Mediate Brassinosteroid-Induced Chlorothalonil Metabolism in Tomato Plants. Environ. Pollut. 2019, 255, 113256. [Google Scholar] [CrossRef] [PubMed]
- Hou, J.; Zhang, Q.; Zhou, Y.; Ahammed, G.J.; Zhou, Y.; Yu, J.; Fang, H.; Xia, X. Glutaredoxin GRXS16 Mediates Brassinosteroid-Induced Apoplastic H2O2 Production to Promote Pesticide Metabolism in Tomato. Environ. Pollut. 2018, 240, 227–234. [Google Scholar] [CrossRef] [PubMed]
- An, S.; Liu, Y.; Sang, K.; Wang, T.; Yu, J.; Zhou, Y.; Xia, X. Brassinosteroid Signaling Positively Regulates Abscisic Acid Biosynthesis in Response to Chilling Stress in Tomato. JIPB 2023, 65, 10–24. [Google Scholar] [CrossRef]
- Liang, B.; Sun, Y.; Wang, J.; Zheng, Y.; Zhang, W.; Xu, Y.; Li, Q.; Leng, P. Tomato Protein Phosphatase 2C Influences the Onset of Fruit Ripening and Fruit Glossiness. J. Exp. Bot. 2020, 72, 2403–2418. [Google Scholar] [CrossRef]
- Wang, J.; Xu, Y.; Zhang, W.; Zheng, Y.; Yuan, B.; Li, Q.; Leng, P. Tomato SlPP2C5 Is Involved in the Regulation of Fruit Development and Ripening. Plant Cell Physiol. 2021, 62, 1760–1769. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, Q.; Jiang, L.; Kai, W.; Liang, B.; Wang, J.; Du, Y.; Zhai, X.; Wang, J.; Zhang, Y.; et al. Suppressing Type 2C Protein Phosphatases Alters Fruit Ripening and the Stress Response in Tomato. Plant Cell Physiol. 2017, 59, 142–154. [Google Scholar] [CrossRef]
- Sun, Y.; Ji, K.; Liang, B.; Du, Y.; Jiang, L.; Wang, J.; Kai, W.; Zhang, Y.; Zhai, X.; Chen, P.; et al. Suppressing ABA Uridine Diphosphate Glucosyltransferase (SlUGT75C1) Alters Fruit Ripening and the Stress Response in Tomato. Plant J. 2017, 91, 574–589. [Google Scholar] [CrossRef] [PubMed]
- Kai, W.; Wang, J.; Liang, B.; Fu, Y.; Zheng, Y.; Zhang, W.; Li, Q.; Leng, P. PYL9 Is Involved in the Regulation of ABA Signaling during Tomato Fruit Ripening. J. Exp. Bot. 2019, 70, 6305–6319. [Google Scholar] [CrossRef]
- Lei, L.; Zhang, J.; Pu, D.; Liu, B.; Meng, X.; Shang, Q.; Duan, Y.; Zhang, F.; Zhang, M.; Dong, C. ABA-responsive AREB1/ABI3-1/ABI5 Cascade Regulates IAA Oxidase Gene SlDAO2 to Inhibit Hypocotyl Elongation in Tomato. Plant Cell Environ. 2022, 46, 498–517. [Google Scholar] [CrossRef]
- Song, J.; Shang, L.; Wang, X.; Xing, Y.; Xu, W.; Zhang, Y.; Wang, T.; Li, H.; Zhang, J.; Ye, Z. MAPK11 Regulates Seed Germination and ABA Signaling in Tomato by Phosphorylating SnRKs. J. Exp. Bot. 2020, 72, 1677–1690. [Google Scholar] [CrossRef]
- Wang, W.; Wang, X.; Wang, Y.; Zhou, G.; Wang, C.; Hussain, S.; Adnan; Lin, R.; Wang, T.; Wang, S. SlEAD1, an EAR Motif-Containing ABA down-Regulated Novel Transcription Repressor Regulates ABA Response in Tomato. GM Crops Food 2020, 11, 275–289. [CrossRef]
- Hu, K.-D.; Zhang, X.-Y.; Yao, G.-F.; Rong, Y.-L.; Ding, C.; Tang, J.; Yang, F.; Huang, Z.-Q.; Xu, Z.-M.; Chen, X.-Y.; et al. A Nuclear-Localized Cysteine Desulfhydrase Plays a Role in Fruit Ripening in Tomato. Hortic. Res. 2020, 7. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.-Q.; Hu, K.-D.; Yao, G.-F.; Wang, S.-Y.; Peng, X.-J.; Zhang, H. A D-Cysteine Desulfhydrase, SlDCD2, Participates in Tomato Fruit Ripening by Modulating ROS Homoeostasis and Ethylene Biosynthesis. Hortic. Res. 2023, 10. [Google Scholar] [CrossRef] [PubMed]
- Sun, C.; Yao, G.; Li, L.; Li, T.; Zhao, Y.; Hu, K.; Zhang, C.; Zhang, H. E3 Ligase BRG3 Persulfidation Delays Tomato Ripening by Reducing Ubiquitination of the Repressor WRKY71. Plant Physiol. 2023, 192, 616–632. [Google Scholar] [CrossRef]
- Hu, K.; Peng, X.; Yao, G.; Zhou, Z.; Yang, F.; Li, W.; Zhao, Y.; Li, Y.; Han, Z.; Chen, X.; et al. Roles of a Cysteine Desulfhydrase LCD1 in Regulating Leaf Senescence in Tomato. IJMS 2021, 22, 13078. [Google Scholar] [CrossRef]
- Ernesto Bianchetti, R.; Silvestre Lira, B.; Santos Monteiro, S.; Demarco, D.; Purgatto, E.; Rothan, C.; Rossi, M.; Freschi, L. Fruit-Localized Phytochromes Regulate Plastid Biogenesis, Starch Synthesis, and Carotenoid Metabolism in Tomato. J. Exp. Bot. 2018, 69, 3573–3586. [Google Scholar] [CrossRef]
- Naeem, M.; Muqarab, R.; Waseem, M. The Solanum Melongena COP1 Delays Fruit Ripening and Influences Ethylene Signaling in Tomato. Journal of Plant Physiol. 2019, 240, 152997. [Google Scholar] [CrossRef] [PubMed]
- Hunziker, J.; Nishida, K.; Kondo, A.; Kishimoto, S.; Ariizumi, T.; Ezura, H. Multiple Gene Substitution by Target-AID Base-Editing Technology in Tomato. Sci. Rep. 2020, 10. [Google Scholar] [CrossRef] [PubMed]
- Gramegna, G.; Rosado, D.; Sánchez Carranza, A.P.; Cruz, A.B.; Simon-Moya, M.; Llorente, B.; Rodríguez-Concepcíon, M.; Freschi, L.; Rossi, M. PHYTOCHROME-INTERACTING FACTOR 3 Mediates Light-dependent Induction of Tocopherol Biosynthesis during Tomato Fruit Ripening. Plant Cell Environ. 2018, 42, 1328–1339. [Google Scholar] [CrossRef] [PubMed]
- Rosado, D.; Trench, B.; Bianchetti, R.; Zuccarelli, R.; Rodrigues Alves, F.R.; Purgatto, E.; Segal Floh, E.I.; Silveira Nogueira, F.T.; Freschi, L.; Rossi, M. Downregulation of PHYTOCHROME-INTERACTING FACTOR 4 Influences Plant Development and Fruit Production. Plant Physiol. 2019, 181, 1360–1370. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Liu, Y.; Ali, M.; Ye, L.; Pan, C.; Li, M.; Zhao, X.; Yu, F.; Zhao, X.; Lu, G. Phytochrome Interacting Factor 3 Regulates Pollen Mitotic Division through Auxin Signalling and Sugar Metabolism Pathways in Tomato. New Phytol. 2021, 234, 560–577. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Meng, L.; Gao, Y.; Grierson, D.; Fu, D. Manipulation of Light Signal Transduction Factors as a Means of Modifying Steroidal Glycoalkaloids Accumulation in Tomato Leaves. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Wang, X.; Zhang, Y.; Yan, J.; Ahammed, G.J.; Bu, X.; Sun, X.; Liu, Y.; Xu, T.; Qi, H.; et al. SlFHY3 and SlHY5 Act Compliantly to Enhance Cold Tolerance through the Integration of Myo-inositol and Light Signaling in Tomato. New Phytol. 2022, 233, 2127–2143. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Zhang, C.; Dong, H.; Liu, X.; Guo, H.; Tong, B.; Fang, F.; Zhao, Y.; Yu, Y.; Liu, Y.; et al. Activation and Negative Feedback Regulation ofSlHY5Transcription by the SlBBX20/21–SlHY5 Transcription Factor Module in UV-B Signaling. Plant Cell 2022, 34, 2038–2055. [Google Scholar] [CrossRef]
- Qiu, Z.; Wang, H.; Li, D.; Yu, B.; Hui, Q.; Yan, S.; Huang, Z.; Cui, X.; Cao, B. Identification of Candidate HY5-Dependent and -Independent Regulators of Anthocyanin Biosynthesis in Tomato. Plant Cell Physiol. 2018, 60, 643–656. [Google Scholar] [CrossRef]
- Balderrama, D.; Barnwell, S.; Carlson, K.D.; Salido, E.; Guevara, R.; Nguyen, C.; Madlung, A. Phytochrome F Mediates Red Light Responsiveness Additively with Phytochromes B1 and B2 in Tomato. Plant Physiol. 2023, 191, 2353–2366. [Google Scholar] [CrossRef]
- Liu, C.; Chi, C.; Jin, L.; Zhu, J.; Yu, J.; Zhou, Y. The bZip Transcription Factor HY5 Mediates CRY1a-induced Anthocyanin Biosynthesis in Tomato. Plant Cell Environ. 2018, 41, 1762–1775. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Hu, C.; Liu, C.; Wang, J.; Zhou, Y.; Yu, J. ELONGATED HYPOCOTYL 5 Mediates Blue Light-Induced Starch Degradation in Tomato. J. Exp. Bot. 2020, 72, 2627–2641. [Google Scholar] [CrossRef]
- Zhi, J.; Liu, X.; Li, D.; Huang, Y.; Yan, S.; Cao, B.; Qiu, Z. CRISPR/Cas9-Mediated SlAN2 Mutants Reveal Various Regulatory Models of Anthocyanin Biosynthesis in Tomato Plant. Plant Cell Rep. 2020, 39, 799–809. [Google Scholar] [CrossRef] [PubMed]
- Jian, W.; Cao, H.; Yuan, S.; Liu, Y.; Lu, J.; Lu, W.; Li, N.; Wang, J.; Zou, J.; Tang, N.; et al. SlMYB75, an MYB-Type Transcription Factor, Promotes Anthocyanin Accumulation and Enhances Volatile Aroma Production in Tomato Fruits. Hortic. Res. 2019, 6. [Google Scholar] [CrossRef] [PubMed]
- Heo, J.; Bang, W.Y.; Jeong, J.C.; Park, S.-C.; Lee, J.M.; Choi, S.; Lee, B.; Lee, Y.K.; Kim, K.; Park, S.J. The Comparisons of Expression Pattern Reveal Molecular Regulation of Fruit Metabolites in S. nigrum and S. lycopersicum. Sci. Rep. 2022, 12. [Google Scholar] [CrossRef]
- Cerqueira, J.V.A.; Zhu, F.; Mendes, K.; Nunes-Nesi, A.; Martins, S.C.V.; Benedito, V.; Fernie, A.R.; Zsogon, A. Promoter Replacement of ANT1 Induces Anthocyanin Accumulation and Triggers the Shade Avoidance Response through Developmental, Physiological and Metabolic Reprogramming in Tomato. Hortic. Res. 2022, 10. [Google Scholar] [CrossRef] [PubMed]
- Deng, L.; Wang, H.; Sun, C.; Li, Q.; Jiang, H.; Du, M.; Li, C.-B.; Li, C. Efficient Generation of Pink-Fruited Tomatoes Using CRISPR/Cas9 System. J. Genet. Genomics 2018, 45, 51–54. [Google Scholar] [CrossRef]
- Sun, C.; Deng, L.; Du, M.; Zhao, J.; Chen, Q.; Huang, T.; Jiang, H.; Li, C.-B.; Li, C. A Transcriptional Network Promotes Anthocyanin Biosynthesis in Tomato Flesh. Mol. Plant 2020, 13, 42–58. [Google Scholar] [CrossRef]
- Yan, S.; Chen, N.; Huang, Z.; Li, D.; Zhi, J.; Yu, B.; Liu, X.; Cao, B.; Qiu, Z. Anthocyanin Fruit Encodes an R2R3-MYB Transcription Factor, SlAN2-like, Activating the Transcription of SlMYBATV to Fine-tune Anthocyanin Content in Tomato Fruit. New Phytol. 2019, 225, 2048–2063. [Google Scholar] [CrossRef]
- Colanero, S.; Perata, P.; Gonzali, S. The Atroviolacea Gene Encodes an R3-MYB Protein Repressing Anthocyanin Synthesis in Tomato Plants. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Zhang, Z.; Xu, Z.; Wang, L.; Chen, C.; Ren, Z. Overexpression of SlMYB75 Enhances Resistance to Botrytis Cinerea and Prolongs Fruit Storage Life in Tomato. Plant Cell Rep. 2020, 40, 43–58. [Google Scholar] [CrossRef] [PubMed]
- Danilo, B.; Perrot, L.; Botton, E.; Nogué, F.; Mazier, M. The DFR Locus: A Smart Landing Pad for Targeted Transgene Insertion in Tomato. PLoS ONE 2018, 13, e0208395. [Google Scholar] [CrossRef] [PubMed]
- Luo, D.; Xiong, C.; Lin, A.; Zhang, C.; Sun, W.; Zhang, J.; Yang, C.; Lu, Y.; Li, H.; Ye, Z.; et al. SlBBX20 Interacts with the COP9 Signalosome Subunit SlCSN5-2 to Regulate Anthocyanin Biosynthesis by Activating SlDFR Expression in Tomato. Hortic. Res. 2021, 8. [Google Scholar] [CrossRef] [PubMed]
- Quinet, M.; Angosto, T.; Yuste-Lisbona, F.J.; Blanchard-Gros, R.; Bigot, S.; Martinez, J.-P.; Lutts, S. Tomato Fruit Development and Metabolism. Front. Plant Sci. 2019, 10. [Google Scholar] [CrossRef]
- Li, S.; Chen, K.; Grierson, D. Molecular and Hormonal Mechanisms Regulating Fleshy Fruit Ripening. Cells 2021, 10, 1136. [Google Scholar] [CrossRef] [PubMed]
- Fenn, M.A.; Giovannoni, J.J. Phytohormones in Fruit Development and Maturation. Plant J. 2021, 105, 446–458. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Andersson, M.; Granell, A.; Cardi, T.; Hofvander, P.; Nicolia, A. Establishment of a DNA-Free Genome Editing and Protoplast Regeneration Method in Cultivated Tomato (Solanum lycopersicum). Plant Cell Rep. 2022, 41, 1843–1852. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Wang, X.; Li, Q.; Peng, W.; Zhang, Z.; Chu, P.; Guo, S.; Fan, Y.; Lyu, S. AtGCS Promoter-Driven Clustered Regularly Interspaced Short Palindromic Repeats/Cas9 Highly Efficiently Generates Homozygous/Biallelic Mutations in the Transformed Roots by Agrobacterium rhizogenes–Mediated Transformation. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef]
- Zhang, N.; Roberts, H.M.; Van Eck, J.; Martin, G.B. Generation and Molecular Characterization of CRISPR/Cas9-Induced Mutations in 63 Immunity-Associated Genes in Tomato Reveals Specificity and a Range of Gene Modifications. Front. Plant Sci. 2020, 11. [Google Scholar] [CrossRef]
- Vu, T.V.; Doan, D.T.H.; Tran, M.T.; Sung, Y.W.; Song, Y.J.; Kim, J.-Y. Improvement of the LbCas12a-crRNA System for Efficient Gene Targeting in Tomato. Front. Plant Sci. 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Kirke, J.; Kaplan, N.; Velez, S.; Jin, X.-L.; Vichyavichien, P.; Zhang, X.-H. Tissue-Preferential Activity and Induction of the Pepper Capsaicin Synthase PUN1 Promoter by Wounding, Heat and Metabolic Pathway Precursor in Tobacco and Tomato Plants. Mol. Biotechnol. 2018, 60, 194–202. [Google Scholar] [CrossRef] [PubMed]
- Vu, T.V.; Sivankalyani, V.; Kim, E.; Doan, D.T.H.; Tran, M.T.; Kim, J.; Sung, Y.W.; Park, M.; Kang, Y.J.; Kim, J. Highly Efficient Homology-directed Repair Using CRISPR/Cpf1-geminiviral Replicon in Tomato. Plant Biotechnol. J. 2020, 18, 2133–2143. [Google Scholar] [CrossRef] [PubMed]
- de Maagd, R.A.; Loonen, A.; Chouaref, J.; Pele, A.; Meijer-Dekens, F.; Fransz, P.; Bai, Y. CRISPR/Cas Inactivation of RECQ4 Increases Homeologous Crossovers in an Interspecific Tomato Hybrid. Plant Biotechnol. J. 2019, 18, 805–813. [Google Scholar] [CrossRef] [PubMed]
- Wada, N.; Osakabe, K.; Osakabe, Y. Type I-D CRISPR System-Mediated Genome Editing in Plants. Methods Mol. Biol. 2023, 21–38. [Google Scholar] [CrossRef]
- Veillet, F.; Perrot, L.; Guyon-Debast, A.; Kermarrec, M.-P.; Chauvin, L.; Chauvin, J.-E.; Gallois, J.-L.; Mazier, M.; Nogue, F. Expanding the CRISPR Toolbox in P. Patens Using SpCas9-NG Variant and Application for Gene and Base Editing in Solanaceae Crops. IJMS 2020, 21, 1024. [Google Scholar] [CrossRef] [PubMed]
- Shimatani, Z.; Kashojiya, S.; Takayama, M.; Terada, R.; Arazoe, T.; Ishii, H.; Teramura, H.; Yamamoto, T.; Komatsu, H.; Miura, K.; et al. Targeted Base Editing in Rice and Tomato Using a CRISPR-Cas9 Cytidine Deaminase Fusion. Nat. Biotechnol. 2017, 35, 441–443. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; He, Y.; Sretenovic, S.; Liu, S.; Cheng, Y.; Han, Y.; Liu, G.; Bao, Y.; Fang, Q.; Zheng, X.; et al. CRISPR-BETS: A Base-editing Design Tool for Generating Stop Codons. Plant Biotechnol. J. 2021, 20, 499–510. [Google Scholar] [CrossRef]
- Matsumoto, A.; Schlüter, T.; Melkonian, K.; Takeda, A.; Nakagami, H.; Mine, A. A Versatile Tn7 Transposon-Based Bioluminescence Tagging Tool for Quantitative and Spatial Detection of Bacteria in Plants. Plant Commun. 2022, 3, 100227. [Google Scholar] [CrossRef]
Figure 1.
CRISPR/Cas9-related studies on tomato by year.

Figure 2.
Topics of studied tomato genes (2017-2023) by CRISPR/Cas9.

Figure 3.
The number of publications devoted to exploring the processes of flowering and ripening in tomatoes. Using CRISPR/Cas9 technology (a), using over- and heterologous expression approach (b), and using gene silencing technologies (c).
Figure 3.
The number of publications devoted to exploring the processes of flowering and ripening in tomatoes. Using CRISPR/Cas9 technology (a), using over- and heterologous expression approach (b), and using gene silencing technologies (c).

Figure 4.
A molecular interaction network model of ripening-related genes in tomato.

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Copyright: This open access article is published under a Creative Commons CC BY 4.0 license, which permit the free download, distribution, and reuse, provided that the author and preprint are cited in any reuse.
Characterization of RIN Isoforms and Their Expression in Tomato Fruit Ripening
Maria Slugina
et al.
Cells,
2021
Effect of Ethylene Sletr1-2 Receptor Allele on Flowering, Fruit Phenotype, Yield, and Shelf-Life of Four F1 Generations of Tropical Tomatoes (Solanum lycopersicum L.)
Anas Anas
et al.
Horticulturae,
2022
Identification of Genes of Molecular Marker TGS0892 on Chromosome 6 and Its Mechanism of Soluble Solids Metabolism in Tomato
Rong-Rong Zhang
et al.
Horticulturae,
2023
MDPI Initiatives
Important Links
© 2024 MDPI (Basel, Switzerland) unless otherwise stated





