1. Introduction
Viruses are widespread in the biosphere and can infect all life forms, including humans, animals, plants, fungi, and bacteria [
1,
2,
3]. They are obligate intracellular parasites that rely on host cells for replication and survival, and can cause a wide range of diseases [
4,
5]. In recent years, several major viral outbreaks, such as Ebola, Zika, COVID-19, and monkeypox, have posed significant threats to global public health [
6,
7,
8,
9], highlighting the importance of understanding virus-host interactions and developing effective prevention and control strategies.
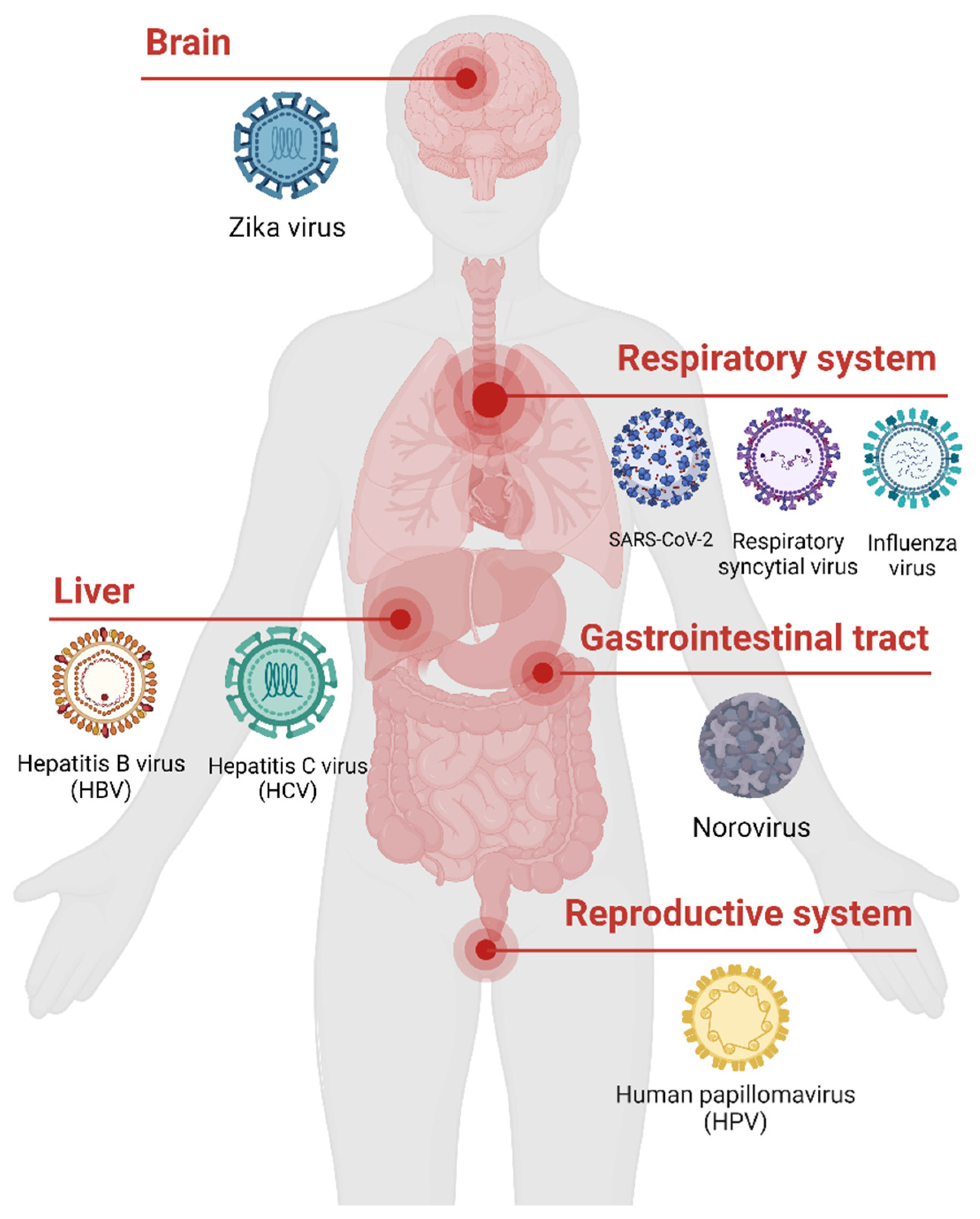
The human virome, which refers to the collection of all viruses that infect humans, is large and complex [
10,
11] (
Figure 1). It includes both pathogenic viruses that cause diseases and conditionally pathogenic viruses that may establish asymptomatic or latent infections. While most studies have focused on single virus infections, it is increasingly recognized that viral co-infections, where a host is infected by multiple viruses simultaneously, are common in humans [
12,
13]. These co-infections can occur between different variants of the same virus, between different species of viruses, or between pathogenic and conditionally pathogenic viruses [
14,
15].
Viral co-infections can have profound impacts on viral evolution, host immunity, and disease outcomes [
16]. Co-infecting viruses may interact directly or indirectly, leading to changes in viral replication, transmission, and pathogenesis. Moreover, viral co-infections can modulate host immune responses, causing enhanced or suppressed immunity, which may affect the clinical manifestations and severity of the disease [
17,
18,
19,
20,
21,
22,
23,
24,
25,
26,
27,
28,
29,
30,
31,
32]. Understanding the prevalence, types, mechanisms, and consequences of human viral co-infections is therefore crucial for the diagnosis, treatment, and prevention of viral diseases.
Despite the growing recognition of the importance of human viral co-infections, our understanding of this complex phenomenon remains limited. Current research mainly focuses on the impacts of specific viruses on the host, while the interactions between co-infecting viruses and their effects on host immunity are less well understood [
16,
19,
20,
21,
22,
23,
24,
25,
26,
27,
28,
29,
30,
31,
32,
33,
34,
35,
36,
37,
38,
39,
40,
41,
42,
43,
44,
45,
46,
47,
48,
49]. Moreover, the experimental detection and computational identification of viral co-infections can be challenging, especially for novel or emerging viruses [
21,
22,
23,
24,
50,
51,
52,
53,
54,
55,
56,
57,
58,
59,
60,
61,
62,
63,
64]. Therefore, a comprehensive review of the current knowledge, methods, and gaps in human viral co-infection research is needed to guide future studies and inform clinical practice.
In this review, we aim to provide an overview of the types, mechanisms, impacts, and identification methods of human viral co-infections. We first introduce the diversity and complexity of the human virome and the significance of studying human viral co-infections. Then, we propose a classification system for human viral co-infections based on the pathogenic properties and species of the viruses involved. Next, we discuss the molecular mechanisms of viral co-infections, focusing on virus-virus interactions, host immune responses, and clinical manifestations, and provide examples from recent studies. We also summarize the experimental and computational methods for identifying viral co-infections, including Multiplex Polymerase Chain Reaction (PCR), immunoassays, metagenomics, and bioinformatics approaches. Finally, we highlight the challenges and future directions in human viral co-infection research, and propose strategies for integrating multi-omics data, developing novel experimental models, and fostering interdisciplinary collaborations to advance our understanding and management of this important topic.
2. Types of Human Viral Co-Infections
The human virome is diverse and dynamic, consisting of both pathogenic and conditionally pathogenic viruses that can establish acute, chronic, or latent infections [
10,
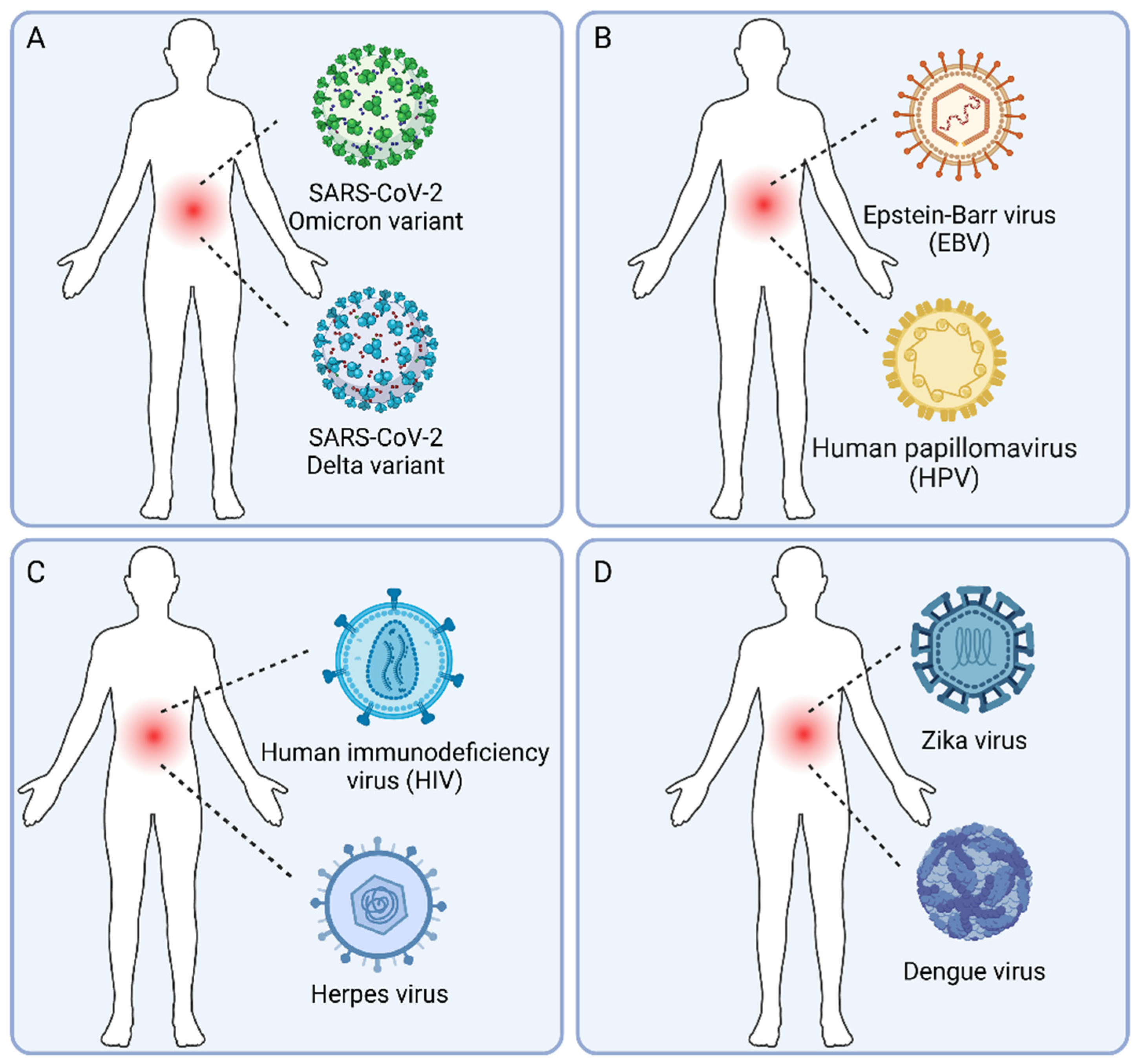
11]. Viral co-infections, where multiple viruses infect a host simultaneously, add another layer of complexity to this already intricate system. To better understand the scope and significance of human viral co-infections, we propose a classification system based on the pathogenic properties and species of the viruses involved (
Figure 2).
2.1. Co-Infections with Different Variants of the Same Virus
This type of co-infection involves multiple variants or strains of the same virus species (
Figure 2A), which may differ in their genetic sequences, antigenicity, virulence, or drug resistance [
50,
51,
52,
65,
66,
67,
68,
69,
70,
71,
72,
73,
74,
75,
76,
77,
78,
79,
80,
81,
82,
83,
84,
85,
86]. Examples include co-infections with different subtypes of human immunodeficiency virus (HIV) [
65,
66,
67,
68,
69], different lineages or reassortants of influenza virus [
16,
70,
71,
72,
73,
74,
75,
76], and different variants of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) [
50,
51,
52,
77,
78,
79,
80,
81,
82,
83,
84,
85,
86]. Co-infection with different viral variants can lead to competition or complementation between the variants, and provide opportunities for recombination or reassortment, which may generate novel strains with altered biological properties [
16,
52,
66,
67,
68,
69,
72,
73,
74,
75,
76,
81,
82,
83,
84,
85,
86].
The clinical implications of co-infection with different viral variants are complex and may vary depending on the specific virus and the host immune status. For example, co-infection with different HIV subtypes has been associated with faster disease progression and increased viral load [
66,
67,
68,
69], while co-infection with different influenza virus subtypes may enhance viral shedding and transmission [
72,
73,
74,
75,
76]. In the case of SARS-CoV-2, co-infection with different variants, such as Delta and Omicron, has been detected in several studies [
50,
51,
78,
79,
80], but the clinical significance of these co-infections remains to be determined [
51].
2.2. Co-Infections with Multiple Conditionally Pathogenic Viruses
Conditionally pathogenic viruses, also known as commensal or symbiotic viruses, are commonly found in healthy individuals and do not usually cause overt diseases [
14,
15]. Examples include anelloviruses, circoviruses, and some herpesviruses [
87,
88,
89], such as Epstein-Barr virus (EBV) and human cytomegalovirus (HCMV) [
87,
90,
91,
92]. These viruses may establish latent or persistent infections and coexist with the host and other viruses in a dynamic equilibrium [
14,
15,
87,
90,
92,
93].
Co-infections with multiple conditionally pathogenic viruses are likely common (
Figure 2B), given the high prevalence of these viruses in the human population [
10,
11]. However, the interactions between these viruses and their impacts on host physiology and immunity are not well understood. Some studies have suggested that conditionally pathogenic viruses may modulate host immune responses and influence the outcome of other infections or diseases [
94,
95,
96]. For example, co-infection with EBV and human papillomavirus (HPV) has been associated with an increased risk of oral and nasopharyngeal cancers [
95,
96].
2.3. Co-Infections with Pathogenic Viruses and Conditionally Pathogenic Viruses
Pathogenic viruses, such as HIV, hepatitis B virus (HBV), and influenza virus, can cause severe and sometimes life-threatening diseases [
21,
22,
23,
33,
34,
35,
36,
37,
65]. When a pathogenic virus co-infects with a conditionally pathogenic virus (
Figure 2C), the interactions between the two viruses may have important clinical implications [
17,
18,
19,
20,
97,
98,
99]. For example, HIV infection can reactivate latent herpesviruses, such as EBV, HCMV, and Kaposi's sarcoma-associated herpesvirus (KSHV), leading to the development of opportunistic infections and cancers [
17,
18,
19,
20,
100]. Similarly, co-infection with HBV and hepatitis D virus (HDV), a satellite virus that requires HBV for replication, can lead to more severe liver disease and accelerated progression to cirrhosis and liver cancer [
101].
The mechanisms by which pathogenic viruses interact with conditionally pathogenic viruses are complex and may involve direct or indirect effects on viral replication, host cell metabolism, and immune responses [
97,
98,
99]. Understanding these interactions is important for the management of human viral co-infections and the development of new therapeutic strategies.
2.4. Co-Infections with Multiple Pathogenic Viruses
Co-infections with multiple pathogenic viruses are of particular concern (
Figure 2D), as they may lead to more severe clinical outcomes and pose challenges for diagnosis and treatment [
21,
22,
23,
24,
25,
26,
27,
28,
29,
30,
31,
32,
33,
34,
35,
36,
37]. Examples include co-infections with HIV and hepatitis C virus (HCV) [
21,
22,
24,
25], HIV and SARS-CoV-2 [
26,
27], and influenza virus and respiratory syncytial virus (RSV) [
102,
103,
104,
105].
The interactions between co-infecting pathogenic viruses can be complex and may involve competition for host resources, modulation of immune responses, and alteration of viral replication and pathogenesis [
33,
34,
35,
36,
37]. For example, co-infection with HIV and HCV is associated with faster progression of liver disease, higher HCV viral load, and increased risk of liver-related mortality [
21,
22,
24,
25]. Similarly, co-infection with RSV and other respiratory viruses can lead to more severe respiratory symptoms, prolonged hospitalization, and increased risk of mortality, especially in young children and older adults [
42,
105].
The diagnosis and treatment of co-infections with multiple pathogenic viruses can be challenging, as the symptoms may overlap and the interactions between the viruses may affect the efficacy of antiviral drugs [
26,
33,
106]. Therefore, a better understanding of the prevalence, types, and mechanisms of these co-infections is crucial for the development of effective prevention and control strategies.
3. Molecular Mechanisms and Clinical Impacts of Human Viral Co-Infections
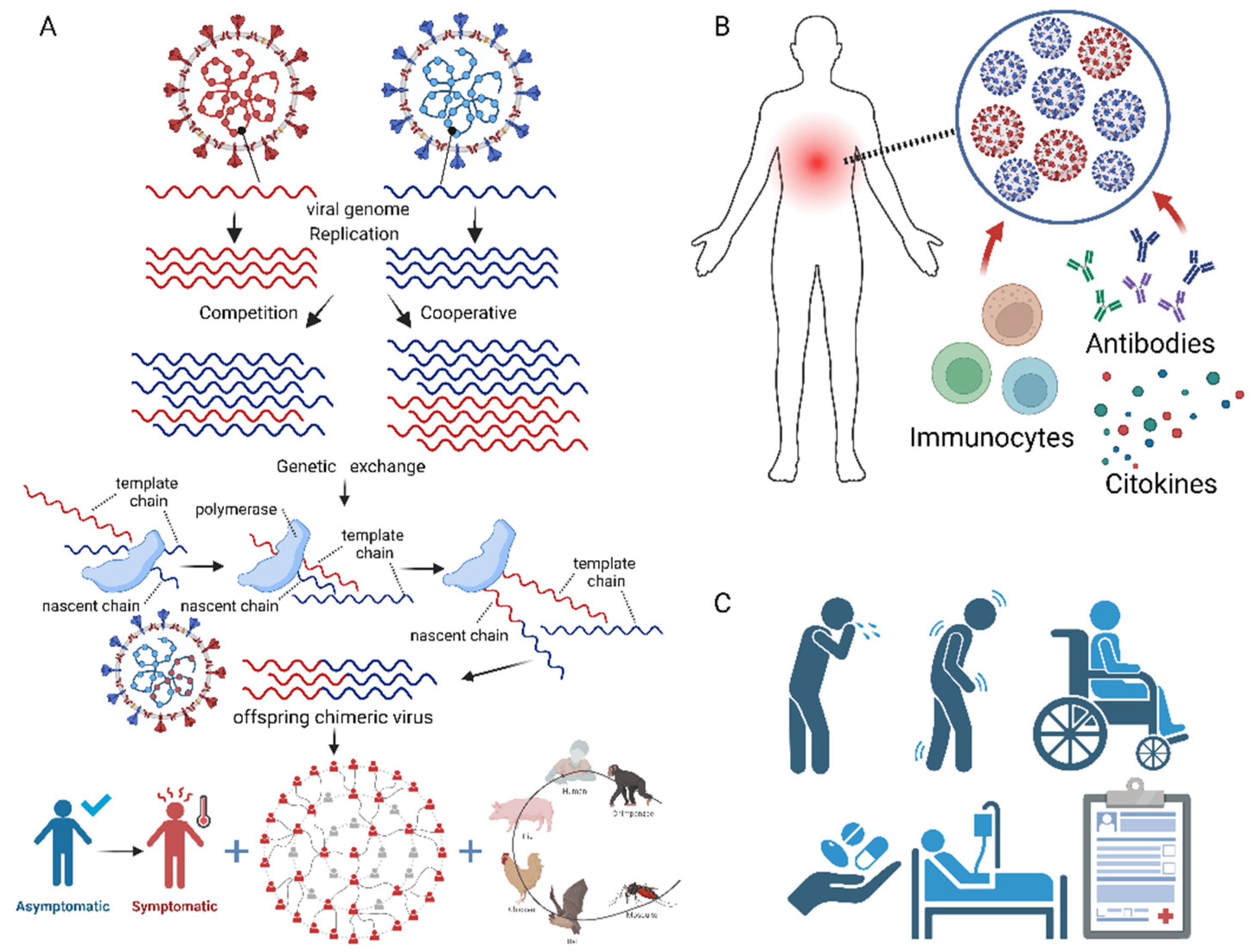
Viral co-infections involve complex interactions between the co-infecting viruses, the host cells, and the immune system, which can have profound impacts on viral replication, pathogenesis, and disease outcomes (
Figure 3). In this section, we discuss the molecular mechanisms of viral co-infections, focusing on virus-virus interactions, host immune responses, and clinical manifestations, and provide examples from recent studies.
3.1. Virus-Virus Interactions
Co-infecting viruses can interact directly or indirectly, leading to changes in viral replication, gene expression, and evolution [
53,
102,
107,
108,
109,
110,
111,
112] (
Figure 3A). These interactions can be competitive, whereby one virus suppresses the replication or pathogenesis of the other [
13], or cooperative, whereby one virus enhances the replication or pathogenesis of the other [
53,
107]. For example, co-infections with human rhinovirus and other respiratory viruses can lead to competitive interactions, with rhinovirus suppressing the replication of other respiratory viruses [
102]. In contrast, co-infection with HIV and HCV can lead to cooperative interactions, with HIV enhancing the replication and pathogenesis of HCV [
21,
22].
The mechanisms of virus-virus interactions are complex and may involve direct physical interactions between viral proteins, competition for host cell receptors or replication machinery, or modulation of host cell signaling pathways. For example, co-infection with human cytomegalovirus (HCMV) and herpes simplex virus-1 (HSV-1) can lead to enhanced replication of both viruses, possibly through direct interactions between viral proteins or modulation of host cell metabolism [
107]. Similarly, co-infection with human parainfluenza virus type 2 (hPIV2) and influenza A virus (IAV) can lead to enhanced IAV replication, possibly through hPIV2-induced changes in host cell fusion and membrane permeability [
53].
Viral co-infections also provide opportunities for genetic exchange between the co-infecting viruses, through mechanisms such as recombination or reassortment [
108,
109,
110,
111,
112] (
Figure 3A). Recombination involves the exchange of genetic material between two closely related viruses, while reassortment involves the exchange of entire gene segments between two viruses with segmented genomes, such as influenza viruses [
110,
112]. These processes can generate novel viral strains with altered antigenicity, virulence, or host range, and contribute to viral evolution and emergence [
108,
109,
110,
111,
112]. For example, recombination between different subtypes of HIV can lead to the emergence of novel strains with increased fitness and drug resistance [
108], while reassortment between human and avian influenza viruses can lead to the emergence of pandemic strains [
112].
3.2. Host Immune Responses
Viral co-infections can have complex and sometimes opposing effects on host immune responses, leading to enhanced or suppressed immunity [
103,
113,
114] (
Figure 3B). The specific immune responses elicited by viral co-infections depend on the types of viruses involved, the timing and order of the infections, and the host immune status.
In some cases, co-infection with two viruses can lead to enhanced immune responses, possibly through cross-reactive T cell or antibody responses [
103,
113]. For example, co-infection with influenza A virus and respiratory syncytial virus (RSV) can lead to enhanced activation of natural killer (NK) cells and CD8+ T cells, which may provide cross-protection against subsequent infections [
103,
115]. Similarly, co-infection with SARS-CoV-2 and influenza virus can lead to enhanced activation of neutrophils and production of pro-inflammatory cytokines, which may contribute to the development of severe disease [
113].
In other cases, viral co-infections can lead to suppressed or dysregulated immune responses, possibly through the induction of immunosuppressive cytokines or the exhaustion of antiviral T cells [
17,
18,
19,
20,
114]. For example, co-infection with HIV and herpesviruses, such as EBV, HCMV, and KSHV, can lead to the reactivation of the herpesviruses and the development of opportunistic infections and cancers, due to the immunosuppressive effects of HIV [
17,
18,
19,
20]. Similarly, co-infection with HCV and other hepatitis viruses can lead to the dysregulation of innate and adaptive immune responses in the liver, contributing to the development of chronic liver disease and cancer [
114].
The long-term effects of viral co-infections on host immunity are not well understood and may depend on the specific viruses and host factors involved. Some studies have suggested that viral co-infections may lead to the development of chronic inflammation or autoimmunity, while others have suggested that they may lead to the induction of regulatory T cells or the establishment of viral persistence. Further research is needed to elucidate the complex interactions between viral co-infections and host immunity, and to develop new strategies for the prevention and treatment of co-infection-associated diseases.
3.3. Clinical Manifestations
The clinical manifestations of viral co-infections are highly variable and depend on the types of viruses involved, the timing and order of the infections, and the host immune status (
Figure 3C). In some cases, viral co-infections can lead to more severe or prolonged symptoms compared to single infections, while in other cases, they may have no apparent effect on disease outcomes [
17,
18,
19,
20,
38,
39,
40,
41,
42,
43,
44,
45,
46,
47,
48,
49].
Studies have shown that co-infections with multiple respiratory viruses, such as influenza virus, RSV, and rhinovirus, can lead to more severe respiratory symptoms, prolonged hospitalization, and increased risk of mortality, especially in young children and older adults [
33,
34,
35,
36,
37,
40,
42,
43]. Similarly, co-infections with HIV and conditionally pathogenic viruses, such as herpesviruses and human papillomavirus (HPV), can lead to the development of AIDS-defining illnesses and cancers [
17,
18,
19,
20].
In contrast, some studies have found no significant differences in disease severity between single and co-infections, or even reduced severity in co-infected patients [
46,
47,
48,
49]. For example, a study of children with bronchiolitis found no significant differences in clinical outcomes between those with single virus infections and those with multiple virus infections [
47]. Another study of children with acute respiratory infections found that co-infections with multiple respiratory viruses were associated with reduced severity of symptoms compared to single respiratory virus infection [
49].
The discrepancies in clinical outcomes of human viral co-infections may reflect the complex interactions between the viruses, the host, and the environment, as well as the limitations of current diagnostic and classification methods. For example, the use of different diagnostic tests, such as PCR or serological assays, may affect the detection and interpretation of viral co-infections [
21,
22,
23,
24,
53,
54,
55,
56,
57]. Similarly, the lack of standardized definitions and classification systems for viral co-infections may hinder the comparison and generalization of results across studies.
To better understand the clinical impacts of human viral co-infections, future studies should use standardized diagnostic methods and classification systems, and consider the potential confounding effects of host and environmental factors. Moreover, the development of novel experimental models and computational tools for studying viral co-infections may provide new insights into the mechanisms and outcomes of these complex co-infections.
4. Experimental and Computational Identification of Human Viral Co-Infections
The accurate and timely identification of human viral co-infections is crucial for the diagnosis, treatment, and prevention of associated diseases. However, the detection and characterization of multiple viruses in a single sample can be challenging, due to the limitations of traditional diagnostic methods and the complexity of viral genomes and interactions [
21,
22,
23,
24,
53,
54,
55,
56,
57]. In this section, we review the current experimental and computational methods for identifying viral co-infections, and discuss their advantages, limitations, and future perspectives.
4.1. Experimental Methods
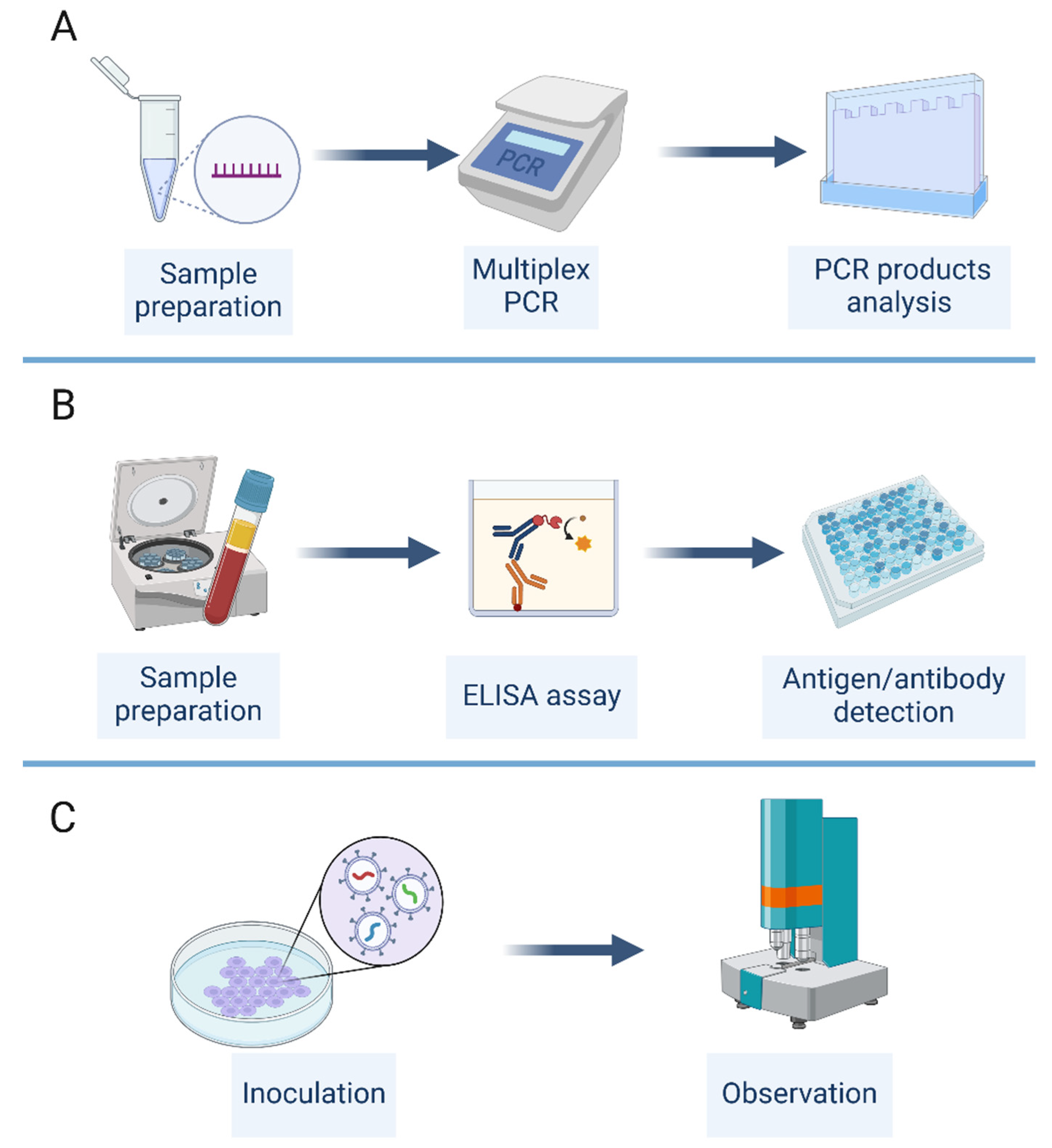
Several experimental methods have been used for the detection and characterization of viral co-infections, including multiplex PCR, immunological assays, and virus culture (
Figure 4). Each of these methods has its own advantages and limitations, and the choice of method depends on the specific viruses and samples involved, as well as the available resources and expertise.
Multiplex PCR is a widely used method for the simultaneous detection of multiple viral targets in a single reaction [
54,
55,
56] (
Figure 4A). This method involves the use of multiple primer sets that are specific for different viral genes or regions, allowing the amplification and detection of multiple viruses in a single assay. Multiplex PCR can be performed using conventional or real-time PCR platforms, and can detect a wide range of viruses, including RNA and DNA viruses [
54,
55,
56]. The advantages of multiplex PCR include its high sensitivity, specificity, and throughput, as well as its ability to detect multiple viruses in a single sample. However, the design and optimization of multiplex PCR assays can be challenging, due to the potential for primer-primer interactions and competition for reagents, which may affect the sensitivity and specificity of the assay.
Immunological assays, such as enzyme-linked immunosorbent assay (ELISA), western blot, and immunofluorescence assay (IFA), are also commonly used for the detection of viral co-infections [
21,
22,
23,
24,
53,
57] (
Figure 4B). These assays involve the use of antibodies that are specific for viral antigens or proteins, allowing the detection and quantification of multiple viruses in a single sample [
57]. The advantages of immunological assays include their high specificity, ease of use, and ability to detect both active and past infections. However, the sensitivity of these assays may be lower than that of PCR-based methods, and the interpretation of results may be complicated by cross-reactivity between different viral antigens.
Virus culture is a traditional method for the isolation and characterization of viruses from clinical samples [
13,
53] (
Figure 4C). This method involves the inoculation of a sample onto a permissive cell line, followed by the observation of cytopathic effects and the identification of the virus using specific antibodies or molecular methods. Virus culture can be used to detect multiple viruses in a single sample, and can provide information on the infectivity and replication kinetics of the viruses [
53]. However, virus culture is time-consuming, labor-intensive, and requires specialized expertise and facilities [
13]. Moreover, some viruses may not grow well in cell culture, or may require the use of specific cell lines or growth conditions [
13].
4.2. Computational Methods
In recent years, the development of high-throughput sequencing technologies and bioinformatics tools has revolutionized the identification and characterization of viral co-infections [
58,
59,
60,
61,
62,
63,
64] (
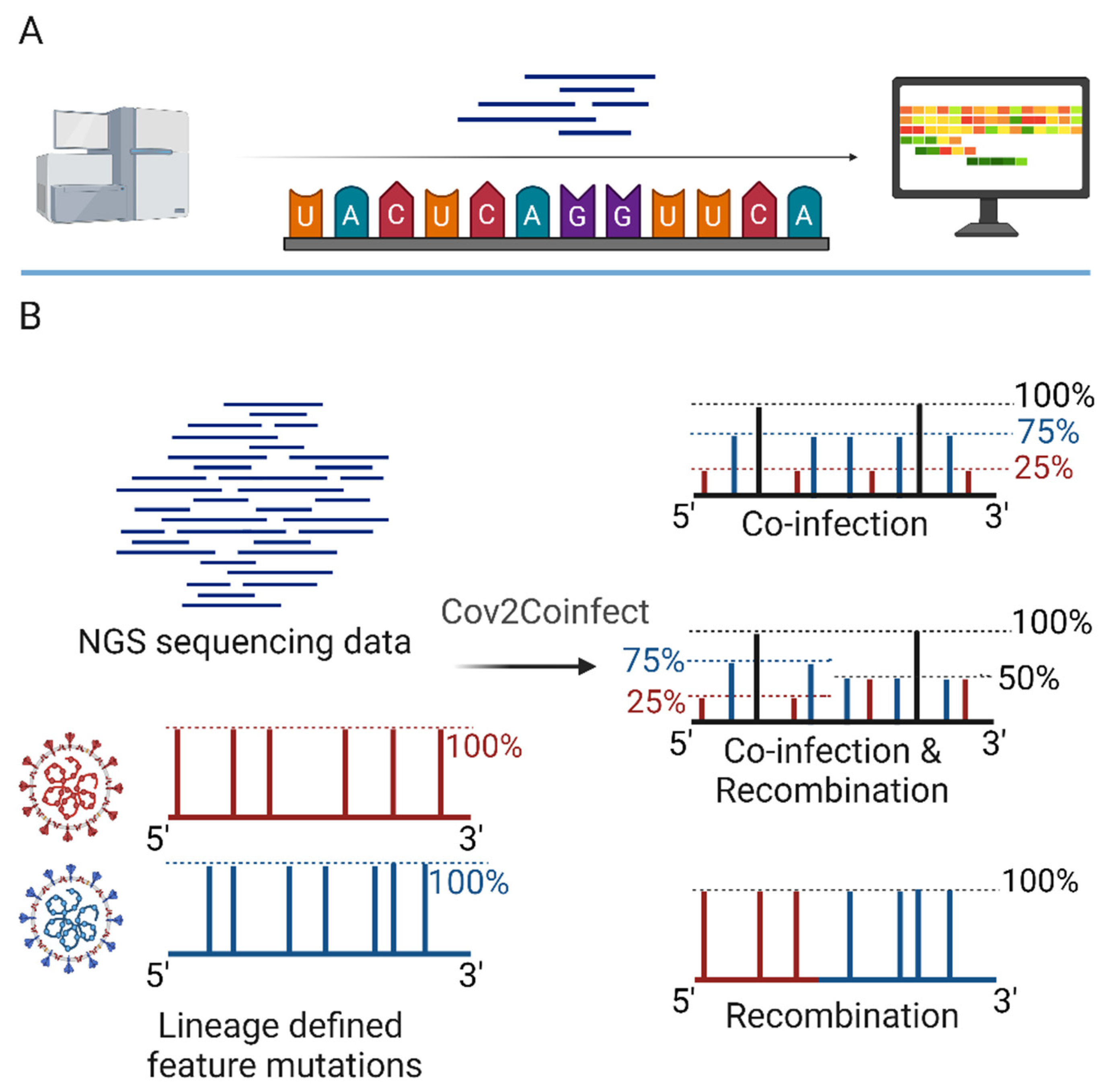
Figure 5). These methods allow the unbiased detection and analysis of all viruses present in a sample, including novel and emerging viruses.
Metagenomics is a powerful approach for the identification of viral co-infections, as it involves the sequencing of all nucleic acids present in a sample (
Figure 5A), without the need for prior knowledge of the viral genomes. This approach can detect both known and novel viruses, as well as their genetic variants and recombinants [
62,
63,
64]. Metagenomics studies have revealed the high prevalence and diversity of viral co-infections in various human tissues and body fluids, including the respiratory tract, gut, and blood. However, the analysis of metagenomic data can be challenging, due to the large amount of data generated, the presence of host and bacterial sequences, and the need for specialized bioinformatics tools and databases.
Several bioinformatics tools and platforms have been developed for the analysis of viral metagenomic data, including GISAID [
58,
59,
60], Virus Variation Resource [
61], and 2019nCoVR [
63]. These platforms provide access to large databases of viral genomes and annotations, as well as tools for the assembly, alignment, and phylogenetic analysis of viral sequences [
58,
59,
60,
61,
62,
63,
64]. These tools can be used to identify and characterize viral co-infections, as well as to track the evolution and transmission of viruses over time and space [
61,
62,
63,
64].
Computational methods have also been developed for the specific identification of viral co-infections and recombinants from sequencing data [
50,
51,
52]. For example, the Cov2Coinfect algorithm uses a hypergeometric distribution test to identify the co-occurrence of lineage-defining feature mutations in SARS-CoV-2 genomes (
Figure 5B), allowing the detection of co-infections with different viral variants [
50]. Similarly, a method developed by Pipek et al. uses the unique defining mutations of viral variants and mutually exclusive defining mutations in variant combinations to identify co-infections and intra-host recombination in SARS-CoV-2 samples [
51]. These methods provide a rapid and accurate way to detect viral co-infections and recombinants, and can be applied to large datasets of viral genomes.
Despite the advances in experimental and computational methods for identifying viral co-infections [
54,
55,
56,
57,
58,
59,
60,
61,
62,
63,
64], several challenges and limitations remain. For example, the sensitivity and specificity of these methods may be affected by the quality and quantity of the viral nucleic acids in the sample, as well as by the presence of PCR inhibitors or contaminating sequences [
54,
55,
56]. Moreover, the interpretation of results may be complicated by the potential for cross-reactivity between different viral targets, or by the presence of defective or incomplete viral genomes [
61,
62,
63,
64]. Further development and validation of these methods, as well as the integration of multiple approaches, may help to overcome these challenges and improve the accuracy and reliability of viral co-infection identification.
5. Discussion and Future Perspectives
Human viral co-infections are a complex and understudied phenomenon that have important implications for viral evolution, host immunity, and disease outcomes. Despite the growing recognition of the prevalence and significance of viral co-infections, our understanding of their types, mechanisms, impacts, and identification methods remains limited. In this review, we have provided an overview of the current knowledge and gaps in the field of human viral co-infections, and highlighted the need for further research and collaboration to address this important public health challenge.
We have proposed a classification system for human viral co-infections based on the pathogenic properties and species of the viruses involved, which includes co-infections with different variants of the same virus, co-infections with multiple conditionally pathogenic viruses, co-infections with pathogenic and conditionally pathogenic viruses, and co-infections with multiple pathogenic viruses. This classification system provides a framework for studying the different types and consequences of viral co-infections, and for developing targeted prevention and control strategies.
We have also discussed the molecular mechanisms of viral co-infections, including virus-virus interactions, host immune responses, and clinical manifestations. Virus-virus interactions can be competitive or cooperative, and can involve direct physical interactions, competition for host resources, or modulation of host cell signaling pathways. These interactions can affect viral replication, evolution, and pathogenesis, and provide opportunities for the emergence of novel viral strains [
108,
109,
110,
111,
112]. Host immune responses to viral co-infections can be enhanced or suppressed, depending on the types of viruses involved, the timing and order of the infections, and the host immune status [
17,
18,
19,
20,
103,
113,
114]. These responses can affect the susceptibility to and severity of viral diseases, as well as the development of chronic inflammation or autoimmunity. The clinical manifestations of viral co-infections are highly variable, ranging from asymptomatic infections to severe and life-threatening diseases [
17,
18,
19,
20,
38,
39,
40,
41,
42,
43,
44,
45,
46,
47,
48,
49]. The accurate diagnosis and management of viral co-infections require a better understanding of their types, mechanisms, and impacts, as well as the development of standardized diagnostic and classification methods.
We have reviewed the current experimental and computational methods for identifying viral co-infections, including multiplex PCR, immunological assays, virus culture, metagenomics, and bioinformatics tools [
54,
55,
56,
57,
58,
59,
60,
61,
62,
63,
64]. These methods have different advantages and limitations, and the choice of method depends on the specific viruses and samples involved, as well as the available resources and expertise [
54,
55,
56,
57]. The integration of multiple approaches, such as the combination of PCR and sequencing methods, may help to improve the accuracy and reliability of viral co-infection identification. Moreover, the development of novel experimental models, such as organoids and animal models, may provide new insights into the mechanisms and consequences of viral co-infections.
Despite the advances in viral co-infection research, several challenges and opportunities remain. First, the complex interactions between co-infecting viruses, host cells, and immune responses are still poorly understood, and require further investigation using systems biology and multi-omics approaches. Second, the long-term impacts of viral co-infections on host health and disease risk are largely unknown, and require longitudinal studies and surveillance programs. Third, the development of effective vaccines and therapies for viral co-infections is hindered by the lack of suitable animal models and the potential for drug-drug interactions. Fourth, the public health and socioeconomic burden of viral co-infections, especially in low- and middle-income countries, is underestimated and requires urgent attention and action.
To address these challenges and opportunities, we propose several future research directions and strategies. First, the integration of multi-omics data, such as genomics, transcriptomics, proteomics, and metabolomics, may provide a comprehensive view of the molecular mechanisms and consequences of viral co-infections. Second, the development of novel experimental models, such as organoids, humanized mice, and non-human primates, may enable the study of viral co-infections in a more physiologically relevant context. Third, the establishment of international collaborations and data sharing platforms, such as GISAID and the Global Virome Project, may facilitate the rapid detection, characterization, and response to viral co-infections, especially during outbreaks and pandemics. Fourth, the engagement of multidisciplinary teams, including virologists, immunologists, clinicians, epidemiologists, and bioinformaticians, may accelerate the translation of basic research findings into clinical and public health applications.
In conclusion, human viral co-infections represent a significant challenge and opportunity for biomedical research and public health. By integrating cutting-edge experimental and computational methods, and fostering collaboration and data sharing across disciplines and borders, we can improve our understanding and management of these complex infections, and develop more effective strategies for their prevention, diagnosis, and treatment. This review provides a comprehensive and up-to-date overview of the field of human viral co-infections, and highlights the need for further research and investment in this important area.
Author Contributions
Conceptualization, A.W. and H.W.; investigation, H.W.; writing—original draft preparation, H.W.; writing—review and editing, H.-Y.Z.; supervision, H.Z. and A.W.; funding acquisition, A.W. and H.-Y.Z. All authors have read and agreed to the published version of the manuscript.
Funding
A.W. was supported by the National Key Research and Development Program of China [grant number 2021YFC2301305]; the National Natural Science Foundation of China [grant number 92169106 and 9216910042], Capital’s Funds for Health Improvement and Research [grant number 2022-1G-1131]; Suzhou Science and Technology Plan Project [grant number szs2020311] ,the CAMS Innovation Fund for Medical Sciences [grant number 2021-I2M-1-061], and Non-profit Central Research Institute Fund of Chinese Academy of Medical Sciences [grant number 2021-PT180-001]; H.-Y.Z. was supported by the Natural Science Foundation of Jiangsu Province [grant number BK20220278].
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
The final draft of this paper was refined and optimized using the large language model to improve readability and expression.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Forterre, P. Defining life: the virus viewpoint. Orig Life Evol Biosph 2010, 40, 151–160. [Google Scholar] [CrossRef] [PubMed]
- Suttle, C. A. Marine viruses--major players in the global ecosystem. Nat Rev Microbiol 2007, 5, 801–812. [Google Scholar] [CrossRef] [PubMed]
- Ryan, F. P. Viruses as symbionts. Symbiosis 2007, 44, 11–21. [Google Scholar]
- Nichol, S. T.; Arikawa, J.; Kawaoka, Y. Emerging viral diseases. Proc Natl Acad Sci U S A 2000, 97, 12411–12412. [Google Scholar] [CrossRef]
- Marston, H. D.; Folkers, G. K.; Morens, D. M.; Fauci, A. S. Emerging viral diseases: confronting threats with new technologies. Sci Transl Med 2014, 6, 253ps210. [Google Scholar] [CrossRef] [PubMed]
- Dixon, M. G.; Schafer, I. J.; Centers for Disease, C.; Prevention. Ebola viral disease outbreak--West Africa, 2014. MMWR Morb Mortal Wkly Rep 2014, 63, 548–551. [Google Scholar] [PubMed]
- WHO Ebola Response Team. Ebola virus disease in West Africa--the first 9 months of the epidemic and forward projections. N Engl J Med 2014, 371, 1481–1495. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Dickens, B. L.; Jit, M.; Cook, A. R.; Carrasco, L. R. Mapping the cryptic spread of the 2015-2016 global Zika virus epidemic. BMC Med 2020, 18, 399. [Google Scholar] [CrossRef] [PubMed]
- Taylor, L. Monkeypox: WHO declares a public health emergency of international concern. BMJ 2022, 378, o1874. [Google Scholar] [CrossRef] [PubMed]
- Liang, G.; Bushman, F. D. The human virome: assembly, composition and host interactions. Nat Rev Microbiol 2021, 19, 514–527. [Google Scholar] [CrossRef] [PubMed]
- Ye, S.; Lu, C.; Qiu, Y.; Zheng, H.; Ge, X.; Wu, A.; Xia, Z.; Jiang, T.; Zhu, H.; Peng, Y. An atlas of human viruses provides new insights into diversity and tissue tropism of human viruses. Bioinformatics 2022, 38, 3087–3093. [Google Scholar] [CrossRef] [PubMed]
- Waner, J. L. Mixed viral infections: detection and management. Clin Microbiol Rev 1994, 7, 143–151. [Google Scholar] [CrossRef] [PubMed]
- Kumar, N.; Sharma, S.; Barua, S.; Tripathi, B. N.; Rouse, B. T. Virological and Immunological Outcomes of Coinfections. Clin Microbiol Rev 2018, 31. [Google Scholar] [CrossRef] [PubMed]
- Lisco, A.; Vanpouille, C.; Margolis, L. War and peace between microbes: HIV-1 interactions with coinfecting viruses. Cell Host Microbe 2009, 6, 403–408. [CrossRef] [PubMed]
- Roossinck, M. J. The good viruses: viral mutualistic symbioses. Nat Rev Microbiol 2011, 9, 99–108. [Google Scholar] [CrossRef] [PubMed]
- Ghedin, E.; Fitch, A.; Boyne, A.; Griesemer, S.; DePasse, J.; Bera, J.; Zhang, X.; Halpin, R. A.; Smit, M.; Jennings, L.; St George, K.; Holmes, E. C.; Spiro, D. J. Mixed infection and the genesis of influenza virus diversity. J Virol 2009, 83, 8832–8841. [Google Scholar] [CrossRef] [PubMed]
- Carbone, A.; Cesarman, E.; Spina, M.; Gloghini, A.; Schulz, T. F. HIV-associated lymphomas and gamma-herpesviruses. Blood 2009, 113, 1213–1224. [Google Scholar] [CrossRef] [PubMed]
- Steininger, C. Clinical relevance of cytomegalovirus infection in patients with disorders of the immune system. Clin Microbiol Infect 2007, 13, 953–963. [Google Scholar] [CrossRef] [PubMed]
- Schacker, T.; Zeh, J.; Hu, H. L.; Hill, E.; Corey, L. Frequency of symptomatic and asymptomatic herpes simplex virus type 2 reactivations among human immunodeficiency virus-infected men. J Infect Dis 1998, 178, 1616–1622. [Google Scholar] [CrossRef] [PubMed]
- Strickler, H. D.; Burk, R. D.; Fazzari, M.; Anastos, K.; Minkoff, H.; Massad, L. S.; Hall, C.; Bacon, M.; Levine, A. M.; Watts, D. H.; Silverberg, M. J.; Xue, X.; Schlecht, N. F.; Melnick, S.; Palefsky, J. M. Natural history and possible reactivation of human papillomavirus in human immunodeficiency virus-positive women. J Natl Cancer Inst 2005, 97, 577–586. [Google Scholar] [CrossRef]
- Sulkowski, M. S. Viral hepatitis and HIV coinfection. J Hepatol 2008, 48, 353–367. [Google Scholar] [CrossRef] [PubMed]
- Taylor, L. E.; Swan, T.; Mayer, K. H. HIV coinfection with hepatitis C virus: evolving epidemiology and treatment paradigms. Clin Infect Dis 2012, 55 Suppl 1, S33–42. [Google Scholar] [CrossRef]
- Chun, H. M.; Roediger, M. P.; Hullsiek, K. H.; Thio, C. L.; Agan, B. K.; Bradley, W. P.; Peel, S. A.; Jagodzinski, L. L.; Weintrob, A. C.; Ganesan, A.; Wortmann, G.; Crum-Cianflone, N. F.; Maguire, J. D.; Landrum, M. L.; Infectious Disease Clinical Research Program, H. I. V. W. G. Hepatitis B virus coinfection negatively impacts HIV outcomes in HIV seroconverters. J Infect Dis 2012, 205, 185–193. [Google Scholar] [CrossRef] [PubMed]
- Matthews, G. V.; Dore, G. J. HIV and hepatitis C coinfection. J Gastroenterol Hepatol 2008, 23, 1000–1008. [Google Scholar] [CrossRef] [PubMed]
- Lacombe, K.; Rockstroh, J. HIV and viral hepatitis coinfections: advances and challenges. Gut 2012, 61 Suppl 1, i47–58. [Google Scholar] [CrossRef]
- Byrd, K. M.; Beckwith, C. G.; Garland, J. M.; Johnson, J. E.; Aung, S.; Cu-Uvin, S.; Farmakiotis, D.; Flanigan, T.; Gillani, F. S.; Macias-Gil, R.; Mileno, M.; Ramratnam, B.; Rybak, N. R.; Sanchez, M.; Tashima, K.; Mylonakis, E.; Kantor, R. SARS-CoV-2 and HIV coinfection: clinical experience from Rhode Island, United States. J Int AIDS Soc 2020, 23, e25573. [Google Scholar] [CrossRef] [PubMed]
- Kanwugu, O. N.; Adadi, P. HIV/SARS-CoV-2 coinfection: A global perspective. J Med Virol 2021, 93, 726–732. [Google Scholar] [CrossRef] [PubMed]
- Adland, E.; Klenerman, P.; Goulder, P.; Matthews, P. C. Ongoing burden of disease and mortality from HIV/CMV coinfection in Africa in the antiretroviral therapy era. Front Microbiol 2015, 6, 1016. [Google Scholar] [CrossRef] [PubMed]
- Effros, R. B. The silent war of CMV in aging and HIV infection. Mech Ageing Dev 2016, 158, 46–52. [Google Scholar] [CrossRef] [PubMed]
- Freeman, M. L.; Lederman, M. M.; Gianella, S. Partners in Crime: The Role of CMV in Immune Dysregulation and Clinical Outcome During HIV Infection. Curr HIV/AIDS Rep 2016, 13, 10–19. [Google Scholar] [CrossRef] [PubMed]
- Desai, D. V.; Kulkarni, S. S. Herpes Simplex Virus: The Interplay Between HSV, Host, and HIV-1. Viral Immunol 2015, 28, 546–555. [Google Scholar] [CrossRef] [PubMed]
- Looker, K. J.; Elmes, J. A. R.; Gottlieb, S. L.; Schiffer, J. T.; Vickerman, P.; Turner, K. M. E.; Boily, M. C. Effect of HSV-2 infection on subsequent HIV acquisition: an updated systematic review and meta-analysis. Lancet Infect Dis 2017, 17, 1303–1316. [Google Scholar] [CrossRef] [PubMed]
- Musuuza, J. S.; Watson, L.; Parmasad, V.; Putman-Buehler, N.; Christensen, L.; Safdar, N. Prevalence and outcomes of co-infection and superinfection with SARS-CoV-2 and other pathogens: A systematic review and meta-analysis. PLoS One 2021, 16, e0251170. [Google Scholar] [CrossRef]
- Cuadrado-Payán, E.; Montagud-Marrahi, E.; Torres-Elorza, M.; Bodro, M.; Blasco, M.; Poch, E.; Soriano, A.; Piñeiro, G. J. J. L. SARS-CoV-2 and influenza virus co-infection. 2020, 395, e84. [Google Scholar] [CrossRef] [PubMed]
- Touzard-Romo, F.; Tape, C.; Lonks, J. R. Co-infection with SARS-CoV-2 and Human Metapneumovirus. R I Med J (2013) 2020, 103, 75–76. [Google Scholar]
- Xiang, X.; Wang, Z. H.; Ye, L. L.; He, X. L.; Wei, X. S.; Ma, Y. L.; Li, H.; Chen, L.; Wang, X. R.; Zhou, Q. Co-infection of SARS-COV-2 and Influenza A Virus: A Case Series and Fast Review. Curr Med Sci 2021, 41, 51–57. [Google Scholar] [CrossRef] [PubMed]
- Massey, B. W.; Jayathilake, K.; Meltzer, H. Y. Respiratory Microbial Co-infection With SARS-CoV-2. Front Microbiol 2020, 11, 2079. [Google Scholar] [CrossRef] [PubMed]
- Cohen, A. L.; McMorrow, M.; Walaza, S.; Cohen, C.; Tempia, S.; Alexander-Scott, M.; Widdowson, M. A. Potential Impact of Co-Infections and Co-Morbidities Prevalent in Africa on Influenza Severity and Frequency: A Systematic Review. PLoS One 2015, 10, e0128580. [Google Scholar] [CrossRef]
- Mercado-Reyes, M.; Acosta-Reyes, J.; Navarro-Lechuga, E.; Corchuelo, S.; Rico, A.; Parra, E.; Tolosa, N.; Pardo, L.; Gonzalez, M.; Martin-Rodriguez-Hernandez, J.; Karime-Osorio, L.; Ospina-Martinez, M.; Rodriguez-Perea, H.; Del Rio-Pertuz, G.; Viasus, D. Dengue, chikungunya and zika virus coinfection: results of the national surveillance during the zika epidemic in Colombia. Epidemiol Infect 2019, 147, e77. [Google Scholar] [CrossRef] [PubMed]
- Goka, E. A.; Vallely, P. J.; Mutton, K. J.; Klapper, P. E. Single, dual and multiple respiratory virus infections and risk of hospitalization and mortality. Epidemiol Infect 2015, 143, 37–47. [Google Scholar] [CrossRef] [PubMed]
- Tang, M. B.; Yu, C. P.; Chen, S. C.; Chen, C. H. Co-Infection of Adenovirus, Norovirus and Torque Teno Virus in Stools of Patients with Acute Gastroenteritis. Southeast Asian J Trop Med Public Health 2014, 45, 1326–1336. [Google Scholar] [PubMed]
- Semple, M. G.; Cowell, A.; Dove, W.; Greensill, J.; McNamara, P. S.; Halfhide, C.; Shears, P.; Smyth, R. L.; Hart, C. A. Dual infection of infants by human metapneumovirus and human respiratory syncytial virus is strongly associated with severe bronchiolitis. J Infect Dis 2005, 191, 382–386. [Google Scholar] [CrossRef] [PubMed]
- Goka, E.; Vallely, P.; Mutton, K.; Klapper, P. Influenza A viruses dual and multiple infections with other respiratory viruses and risk of hospitalisation and mortality. Influenza Other Respir Viruses 2013, 7, 1079–1087. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez Alvarez, D. A.; Lopez Cortes, L. F.; Cordero, E. Impact of HIV on the severity of influenza. Expert Rev Respir Med 2016, 10, 463–472. [Google Scholar] [CrossRef] [PubMed]
- Griesbeck, M.; Valantin, M. A.; Lacombe, K.; Samri-Hassimi, A.; Bottero, J.; Blanc, C.; Sbihi, Z.; Zoorob, R.; Katlama, C.; Guiguet, M.; Altfeld, M.; Autran, B.; Hep, A. C. T. V. I. H. s. g. Hepatitis C virus drives increased type I interferon-associated impairments associated with fibrosis severity in antiretroviral treatment-treated HIV-1-hepatitis C virus-coinfected individuals. AIDS 2017, 31, 1223–1234. [Google Scholar] [CrossRef] [PubMed]
- Martin, E. T.; Fairchok, M. P.; Stednick, Z. J.; Kuypers, J.; Englund, J. A. Epidemiology of multiple respiratory viruses in childcare attendees. J Infect Dis 2013, 207, 982–989. [Google Scholar] [CrossRef] [PubMed]
- Brand, H. K.; de Groot, R.; Galama, J. M.; Brouwer, M. L.; Teuwen, K.; Hermans, P. W.; Melchers, W. J.; Warris, A. Infection with multiple viruses is not associated with increased disease severity in children with bronchiolitis. Pediatr Pulmonol 2012, 47, 393–400. [Google Scholar] [CrossRef]
- Rotzen-Ostlund, M.; Eriksson, M.; Tiveljung Lindell, A.; Allander, T.; Zweygberg Wirgart, B.; Grillner, L. Children with multiple viral respiratory infections are older than those with single viruses. Acta Paediatr 2014, 103, 100–104. [Google Scholar] [CrossRef] [PubMed]
- Martin, E. T.; Kuypers, J.; Wald, A.; Englund, J. A. Multiple versus single virus respiratory infections: viral load and clinical disease severity in hospitalized children. Influenza Other Respir Viruses 2012, 6, 71–77. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H. Y.; Cheng, Y. X.; Xu, L.; Li, J. Y.; Tao, C. Y.; Ji, C. Y.; Han, N.; Yang, R.; Wu, H.; Li, Y.; Wu, A. Genomic evidence for divergent co-infections of co-circulating SARS-CoV-2 lineages. Comput Struct Biotechnol J 2022, 20, 4015–4024. [Google Scholar] [CrossRef] [PubMed]
- Pipek, O. A.; Medgyes-Horvath, A.; Steger, J.; Papp, K.; Visontai, D.; Koopmans, M.; Nieuwenhuijse, D.; Oude Munnink, B. B.; Group, V. E. O. T. W.; Csabai, I. Systematic detection of co-infection and intra-host recombination in more than 2 million global SARS-CoV-2 samples. Nat Commun 2024, 15, 517. [Google Scholar] [CrossRef] [PubMed]
- Bolze, A.; Basler, T.; White, S.; Dei Rossi, A.; Wyman, D.; Dai, H.; Roychoudhury, P.; Greninger, A. L.; Hayashibara, K.; Beatty, M.; Shah, S.; Stous, S.; McCrone, J. T.; Kil, E.; Cassens, T.; Tsan, K.; Nguyen, J.; Ramirez, J.; Carter, S.; Cirulli, E. T.; Schiabor Barrett, K.; Washington, N. L.; Belda-Ferre, P.; Jacobs, S.; Sandoval, E.; Becker, D.; Lu, J. T.; Isaksson, M.; Lee, W.; Luo, S. Evidence for SARS-CoV-2 Delta and Omicron co-infections and recombination. Med 2022, 3, 848–859 e844. [Google Scholar] [CrossRef] [PubMed]
- Goto, H.; Ihira, H.; Morishita, K.; Tsuchiya, M.; Ohta, K.; Yumine, N.; Tsurudome, M.; Nishio, M. Enhanced growth of influenza A virus by coinfection with human parainfluenza virus type 2. Med Microbiol Immunol 2016, 205, 209–218. [Google Scholar] [CrossRef] [PubMed]
- Ching, N. S.; Kotsanas, D.; Easton, M. L.; Francis, M. J.; Korman, T. M.; Buttery, J. P. Respiratory virus detection and co-infection in children and adults in a large Australian hospital in 2009-2015. J Paediatr Child Health 2018, 54, 1321–1328. [Google Scholar] [CrossRef] [PubMed]
- Roest, R. W.; Maertzdorf, J.; Kant, M.; van der Meijden, W. I.; Osterhaus, A. D.; Verjans, G. M. High incidence of genotypic variance between sequential herpes simplex virus type 2 isolates from HIV-1-seropositive patients with recurrent genital herpes. J Infect Dis 2006, 194, 1115–1118. [Google Scholar] [CrossRef] [PubMed]
- van Baarle, D.; Hovenkamp, E.; Kersten, M. J.; Klein, M. R.; Miedema, F.; van Oers, M. H. Direct Epstein-Barr virus (EBV) typing on peripheral blood mononuclear cells: no association between EBV type 2 infection or superinfection and the development of acquired immunodeficiency syndrome-related non-Hodgkin's lymphoma. Blood 1999, 93, 3949–3955. [Google Scholar] [PubMed]
- Evans, L. A.; Moreau, J.; Odehouri, K.; Seto, D.; Thomson-Honnebier, G.; Legg, H.; Barboza, A.; Cheng-Mayer, C.; Levy, J. A. Simultaneous isolation of HIV-1 and HIV-2 from an AIDS patient. Lancet 1988, 2, 1389–1391. [Google Scholar] [CrossRef] [PubMed]
- Shu, Y.; McCauley, J. GISAID: Global initiative on sharing all influenza data - from vision to reality. Euro Surveill 2017, 22. [Google Scholar] [CrossRef] [PubMed]
- Elbe, S.; Buckland-Merrett, G. Data, disease and diplomacy: GISAID's innovative contribution to global health. Glob Chall 2017, 1, 33–46. [Google Scholar] [CrossRef]
- Khare, S.; Gurry, C.; Freitas, L.; Schultz, M. B.; Bach, G.; Diallo, A.; Akite, N.; Ho, J.; Lee, R. T.; Yeo, W.; Curation Team, G. C.; Maurer-Stroh, S. GISAID's Role in Pandemic Response. China CDC Wkly 2021, 3, 1049–1051. [Google Scholar] [CrossRef] [PubMed]
- Hatcher, E. L.; Zhdanov, S. A.; Bao, Y.; Blinkova, O.; Nawrocki, E. P.; Ostapchuck, Y.; Schaffer, A. A.; Brister, J. R. Virus Variation Resource - improved response to emergent viral outbreaks. Nucleic Acids Res 2017, 45, D482–D490. [Google Scholar] [CrossRef] [PubMed]
- consortiumcontact@cogconsortium.uk, C.-G. U. An integrated national scale SARS-CoV-2 genomic surveillance network. Lancet Microbe 2020, 1, e99–e100. [Google Scholar] [CrossRef] [PubMed]
- Song, S.; Ma, L.; Zou, D.; Tian, D.; Li, C.; Zhu, J.; Chen, M.; Wang, A.; Ma, Y.; Li, M.; Teng, X.; Cui, Y.; Duan, G.; Zhang, M.; Jin, T.; Shi, C.; Du, Z.; Zhang, Y.; Liu, C.; Li, R.; Zeng, J.; Hao, L.; Jiang, S.; Chen, H.; Han, D.; Xiao, J.; Zhang, Z.; Zhao, W.; Xue, Y.; Bao, Y. The Global Landscape of SARS-CoV-2 Genomes, Variants, and Haplotypes in 2019nCoVR. Genomics Proteomics Bioinformatics 2020, 18, 749–759. [Google Scholar] [CrossRef] [PubMed]
- Gong, Z.; Zhu, J. W.; Li, C. P.; Jiang, S.; Ma, L. N.; Tang, B. X.; Zou, D.; Chen, M. L.; Sun, Y. B.; Song, S. H.; Zhang, Z.; Xiao, J. F.; Xue, Y. B.; Bao, Y. M.; Du, Z. L.; Zhao, W. M. An online coronavirus analysis platform from the National Genomics Data Center. Zool Res 2020, 41, 705–708. [Google Scholar] [CrossRef] [PubMed]
- Klimas, N.; Koneru, A. O.; Fletcher, M. A. Overview of HIV. Psychosom Med 2008, 70, 523–530. [Google Scholar] [CrossRef] [PubMed]
- Gottlieb, G. S.; Nickle, D. C.; Jensen, M. A.; Wong, K. G.; Grobler, J.; Li, F.; Liu, S. L.; Rademeyer, C.; Learn, G. H.; Karim, S. S.; Williamson, C.; Corey, L.; Margolick, J. B.; Mullins, J. I. Dual HIV-1 infection associated with rapid disease progression. Lancet 2004, 363, 619–622. [Google Scholar] [CrossRef] [PubMed]
- van der Kuyl, A. C.; Cornelissen, M. Identifying HIV-1 dual infections. Retrovirology 2007, 4, 67. [Google Scholar] [CrossRef] [PubMed]
- Goulder, P. J.; Walker, B. D. HIV-1 superinfection--a word of caution. N Engl J Med 2002, 347, 756–758. [Google Scholar] [CrossRef] [PubMed]
- Blackard, J. T.; Cohen, D. E.; Mayer, K. H. Human immunodeficiency virus superinfection and recombination: current state of knowledge and potential clinical consequences. Clin Infect Dis 2002, 34, 1108–1114. [Google Scholar] [CrossRef]
- Hutchinson, E. C. Influenza Virus. Trends Microbiol 2018, 26, 809–810. [Google Scholar] [CrossRef] [PubMed]
- Kawaoka, Y.; Neumann, G. Influenza viruses: an introduction. Methods Mol Biol 2012, 865, 1–9. [Google Scholar] [PubMed]
- Ghedin, E.; Laplante, J.; DePasse, J.; Wentworth, D. E.; Santos, R. P.; Lepow, M. L.; Porter, J.; Stellrecht, K.; Lin, X.; Operario, D.; Griesemer, S.; Fitch, A.; Halpin, R. A.; Stockwell, T. B.; Spiro, D. J.; Holmes, E. C.; St George, K. Deep sequencing reveals mixed infection with 2009 pandemic influenza A (H1N1) virus strains and the emergence of oseltamivir resistance. J Infect Dis 2011, 203, 168–174. [Google Scholar] [CrossRef] [PubMed]
- Vijaykrishna, D.; Mukerji, R.; Smith, G. J. RNA Virus Reassortment: An Evolutionary Mechanism for Host Jumps and Immune Evasion. PLoS Pathog 2015, 11, e1004902. [Google Scholar] [CrossRef] [PubMed]
- Tao, H.; Li, L.; White, M. C.; Steel, J.; Lowen, A. C. Influenza A Virus Coinfection through Transmission Can Support High Levels of Reassortment. J Virol 2015, 89, 8453–8461. [Google Scholar] [CrossRef] [PubMed]
- Peacey, M.; Hall, R. J.; Sonnberg, S.; Ducatez, M.; Paine, S.; Nicol, M.; Ralston, J. C.; Bandaranayake, D.; Hope, V.; Webby, R. J.; Huang, S. Pandemic (H1N1) 2009 and seasonal influenza A (H1N1) co-infection, New Zealand, 2009. Emerg Infect Dis 2010, 16, 1618–1620. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X. S.; De Angelis, D.; White, P. J.; Charlett, A.; Pebody, R. G.; McCauley, J. Co-circulation of influenza A virus strains and emergence of pandemic via reassortment: the role of cross-immunity. Epidemics 2013, 5, 20–33. [Google Scholar] [CrossRef] [PubMed]
- Wu, A.; Wang, L.; Zhou, H. Y.; Ji, C. Y.; Xia, S. Z.; Cao, Y.; Meng, J.; Ding, X.; Gold, S.; Jiang, T.; Cheng, G. One year of SARS-CoV-2 evolution. Cell Host Microbe 2021, 29, 503–507. [Google Scholar] [CrossRef] [PubMed]
- Bal, A.; Simon, B.; Destras, G.; Chalvignac, R.; Semanas, Q.; Oblette, A.; Queromes, G.; Fanget, R.; Regue, H.; Morfin, F.; Valette, M.; Lina, B.; Josset, L. Detection and prevalence of SARS-CoV-2 co-infections during the Omicron variant circulation in France. Nat Commun 2022, 13, 6316. [Google Scholar] [CrossRef]
- Combes, P.; Bisseux, M.; Bal, A.; Marin, P.; Latour, J.; Archimbaud, C.; Brebion, A.; Chabrolles, H.; Regagnon, C.; Lafolie, J.; Destras, G.; Simon, B.; Izopet, J.; Laurence, J.; Henquell, C.; Mirand, A. Evidence of co-infections during Delta and Omicron SARS-CoV-2 variants co-circulation through prospective screening and sequencing. Clin Microbiol Infect 2022, 28, 1503 e1505–1503 e1508. [Google Scholar] [CrossRef] [PubMed]
- Dezordi, F. Z.; Resende, P. C.; Naveca, F. G.; do Nascimento, V. A.; de Souza, V. C.; Dias Paixao, A. C.; Appolinario, L.; Lopes, R. S.; da Fonseca Mendonca, A. C.; Barreto da Rocha, A. S.; Martins Venas, T. M.; Pereira, E. C.; Paiva, M. H. S.; Docena, C.; Bezerra, M. F.; Machado, L. C.; Salvato, R. S.; Gregianini, T. S.; Martins, L. G.; Pereira, F. M.; Rovaris, D. B.; Fernandes, S. B.; Ribeiro-Rodrigues, R.; Costa, T. O.; Sousa, J. C.; Miyajima, F.; Delatorre, E.; Graf, T.; Bello, G.; Siqueira, M. M.; Wallau, G. L. Unusual SARS-CoV-2 intrahost diversity reveals lineage superinfection. Microb Genom 2022, 8. [Google Scholar] [CrossRef] [PubMed]
- Su, S.; Wong, G.; Shi, W.; Liu, J.; Lai, A. C. K.; Zhou, J.; Liu, W.; Bi, Y.; Gao, G. F. Epidemiology, Genetic Recombination, and Pathogenesis of Coronaviruses. Trends Microbiol 2016, 24, 490–502. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Meng, K.; Meng, G. Genomic recombination events may reveal the evolution of coronavirus and the origin of SARS-CoV-2. Sci Rep 2020, 10, 21617. [Google Scholar] [CrossRef] [PubMed]
- Neches, R. Y.; McGee, M. D.; Kyrpides, N. C. Recombination should not be an afterthought. Nat Rev Microbiol 2020, 18, 606. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Ma, W.; Dang, S.; Chen, L.; Zhang, R.; Mei, S.; Wei, X.; Lv, Q.; Peng, B.; Chen, J.; Kong, D.; Sun, Y.; Tang, X.; Wu, W.; Chen, Z.; Li, S.; Wan, J.; Zou, X.; Li, M.; Feng, T.; Ren, L.; Wang, J. Possible recombination between two variants of concern in a COVID-19 patient. Emerg Microbes Infect 2022, 11, 552–555. [Google Scholar] [CrossRef] [PubMed]
- Jackson, B.; Boni, M. F.; Bull, M. J.; Colleran, A.; Colquhoun, R. M.; Darby, A. C.; Haldenby, S.; Hill, V.; Lucaci, A.; McCrone, J. T.; Nicholls, S. M.; O'Toole, A.; Pacchiarini, N.; Poplawski, R.; Scher, E.; Todd, F.; Webster, H. J.; Whitehead, M.; Wierzbicki, C.; Consortium, C.-G. U.; Loman, N. J.; Connor, T. R.; Robertson, D. L.; Pybus, O. G.; Rambaut, A. Generation and transmission of interlineage recombinants in the SARS-CoV-2 pandemic. Cell 2021, 184, 5179–5188 e5178. [Google Scholar] [CrossRef] [PubMed]
- Varabyou, A.; Pockrandt, C.; Salzberg, S. L.; Pertea, M. Rapid detection of inter-clade recombination in SARS-CoV-2 with Bolotie. Genetics 2021, 218. [Google Scholar] [CrossRef] [PubMed]
- Whitley, R. J.; Roizman, B. Herpes simplex virus infections. Lancet 2001, 357, 1513–1518. [Google Scholar] [CrossRef] [PubMed]
- Kaczorowska, J.; van der Hoek, L. Human anelloviruses: diverse, omnipresent and commensal members of the virome. FEMS Microbiol Rev 2020, 44, 305–313. [Google Scholar] [CrossRef] [PubMed]
- Biagini, P. Human circoviruses. Vet Microbiol 2004, 98, 95–101. [Google Scholar] [CrossRef] [PubMed]
- Griffiths, P.; Baraniak, I.; Reeves, M. The pathogenesis of human cytomegalovirus. J Pathol 2015, 235, 288–297. [Google Scholar] [CrossRef] [PubMed]
- Landolfo, S.; Gariglio, M.; Gribaudo, G.; Lembo, D. The human cytomegalovirus. Pharmacol Ther 2003, 98, 269–297. [Google Scholar] [CrossRef] [PubMed]
- Cohen, J. I. Epstein-Barr virus infection. N Engl J Med 2000, 343, 481–492. [Google Scholar] [CrossRef]
- Baseman, J. G.; Koutsky, L. A. The epidemiology of human papillomavirus infections. J Clin Virol 2005, 32 Suppl 1, S16–24. [Google Scholar] [CrossRef]
- Sand, L.; Jalouli, J. Viruses and oral cancer. Is there a link? Microbes Infect 2014, 16, 371–378. [Google Scholar] [CrossRef] [PubMed]
- Polz-Gruszka, D.; Stec, A.; Dworzanski, J.; Polz-Dacewicz, M. EBV, HSV, CMV and HPV in laryngeal and oropharyngeal carcinoma in Polish patients. Anticancer Res 2015, 35, 1657–1661. [Google Scholar] [PubMed]
- Al Moustafa, A. E.; Chen, D.; Ghabreau, L.; Akil, N. Association between human papillomavirus and Epstein-Barr virus infections in human oral carcinogenesis. Med Hypotheses 2009, 73, 184–186. [Google Scholar] [CrossRef] [PubMed]
- Virgin, H. W.; Wherry, E. J.; Ahmed, R. Redefining chronic viral infection. Cell 2009, 138, 30–50. [Google Scholar] [CrossRef] [PubMed]
- Thein, H. H.; Yi, Q.; Dore, G. J.; Krahn, M. D. Natural history of hepatitis C virus infection in HIV-infected individuals and the impact of HIV in the era of highly active antiretroviral therapy: a meta-analysis. AIDS 2008, 22, 1979–1991. [Google Scholar] [CrossRef] [PubMed]
- Masur, H.; Kaplan, J. Current status of opportunistic infections in patients with HIV infection in the era of highly active antiretroviral therapy. Curr Clin Top Infect Dis 2001, 21, 64–82. [Google Scholar] [PubMed]
- Whitby, D.; Howard, M. R.; Tenant-Flowers, M.; Brink, N. S.; Copas, A.; Boshoff, C.; Hatzioannou, T.; Suggett, F. E.; Aldam, D. M.; Denton, A. S.; et al. Detection of Kaposi sarcoma associated herpesvirus in peripheral blood of HIV-infected individuals and progression to Kaposi's sarcoma. Lancet 1995, 346, 799–802. [Google Scholar] [CrossRef] [PubMed]
- Urban, S.; Neumann-Haefelin, C.; Lampertico, P. Hepatitis D virus in 2021: virology, immunology and new treatment approaches for a difficult-to-treat disease. Gut 2021, 70, 1782–1794. [Google Scholar] [CrossRef] [PubMed]
- Pinky, L.; Dobrovolny, H. M. Coinfections of the Respiratory Tract: Viral Competition for Resources. PLoS One 2016, 11, e0155589. [Google Scholar] [CrossRef] [PubMed]
- Piret, J.; Boivin, G. Viral Interference between Respiratory Viruses. Emerg Infect Dis 2022, 28, 273–281. [Google Scholar] [CrossRef] [PubMed]
- Topoulos, S.; Giesa, C.; Gatermann, S.; Fussen, R.; Lemmen, S.; Ewig, S. Analysis of acute respiratory infections due to influenza virus A, B and RSV during an influenza epidemic 2018. Infection 2019, 47, 425–433. [Google Scholar] [CrossRef] [PubMed]
- Welliver, T. P.; Reed, J. L.; Welliver, R. C., Sr. Respiratory syncytial virus and influenza virus infections: observations from tissues of fatal infant cases. Pediatr Infect Dis J 2008, 27, S92–96. [Google Scholar] [CrossRef] [PubMed]
- Griffiths, E. C.; Pedersen, A. B.; Fenton, A.; Petchey, O. L. The nature and consequences of coinfection in humans. J Infect 2011, 63, 200–206. [Google Scholar] [CrossRef] [PubMed]
- Rinaldo, C. R., Jr.; Richter, B. S.; Black, P. H.; Callery, R.; Chess, L.; Hirsch, M. S. Replication of herpes simplex virus and cytomegalovirus in human leukocytes. J Immunol 1978, 120, 130–136. [Google Scholar] [CrossRef]
- Parrish, C. R.; Holmes, E. C.; Morens, D. M.; Park, E. C.; Burke, D. S.; Calisher, C. H.; Laughlin, C. A.; Saif, L. J.; Daszak, P. Cross-species virus transmission and the emergence of new epidemic diseases. Microbiol Mol Biol Rev 2008, 72, 457–470. [Google Scholar] [CrossRef]
- Perez-Losada, M.; Arenas, M.; Galan, J. C.; Palero, F.; Gonzalez-Candelas, F. Recombination in viruses: mechanisms, methods of study, and evolutionary consequences. Infect Genet Evol 2015, 30, 296–307. [Google Scholar] [CrossRef]
- McDonald, S. M.; Nelson, M. I.; Turner, P. E.; Patton, J. T. Reassortment in segmented RNA viruses: mechanisms and outcomes. Nat Rev Microbiol 2016, 14, 448–460. [Google Scholar] [CrossRef]
- Focosi, D.; Maggi, F. Recombination in Coronaviruses, with a Focus on SARS-CoV-2. Viruses 2022, 14. [Google Scholar] [CrossRef] [PubMed]
- Schweiger, B.; Bruns, L.; Meixenberger, K. Reassortment between human A(H3N2) viruses is an important evolutionary mechanism. Vaccine 2006, 24, 6683–6690. [Google Scholar] [CrossRef] [PubMed]
- Guan, Z.; Chen, C.; Li, Y.; Yan, D.; Zhang, X.; Jiang, D.; Yang, S.; Li, L. Impact of Coinfection With SARS-CoV-2 and Influenza on Disease Severity: A Systematic Review and Meta-Analysis. Front Public Health 2021, 9, 773130. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Verslype, C.; van Pelt, J. F.; van Ranst, M.; Fevery, J. Viral interaction and clinical implications of coinfection of hepatitis C virus with other hepatitis viruses. Eur J Gastroenterol Hepatol 2006, 18, 1311–1319. [Google Scholar] [CrossRef] [PubMed]
- Chan, K. F.; Carolan, L. A.; Korenkov, D.; Druce, J.; McCaw, J.; Reading, P. C.; Barr, I. G.; Laurie, K. L. Investigating Viral Interference Between Influenza A Virus and Human Respiratory Syncytial Virus in a Ferret Model of Infection. J Infect Dis 2018, 218, 406–417. [Google Scholar] [CrossRef] [PubMed]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).