Submitted:
07 July 2024
Posted:
08 July 2024
You are already at the latest version
Abstract
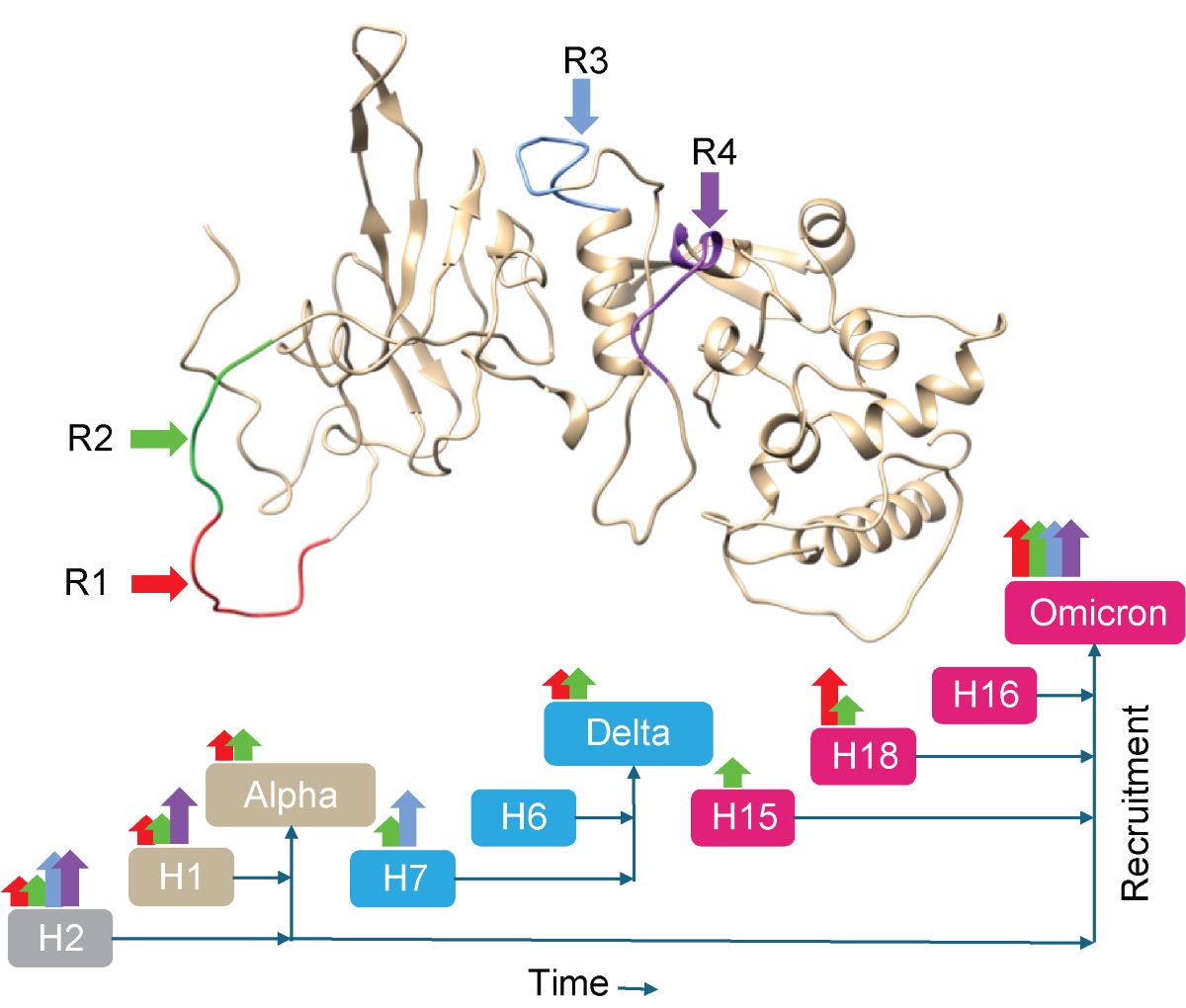
Keywords:
1. Introduction
2. Materials and Methods
3. Results
3.1. Full-Length Analysis of the N-Protein Using Backbone Root Mean Square Deviations (RMSD)
3.2. Regional Analysis with TM Scores Using USalign
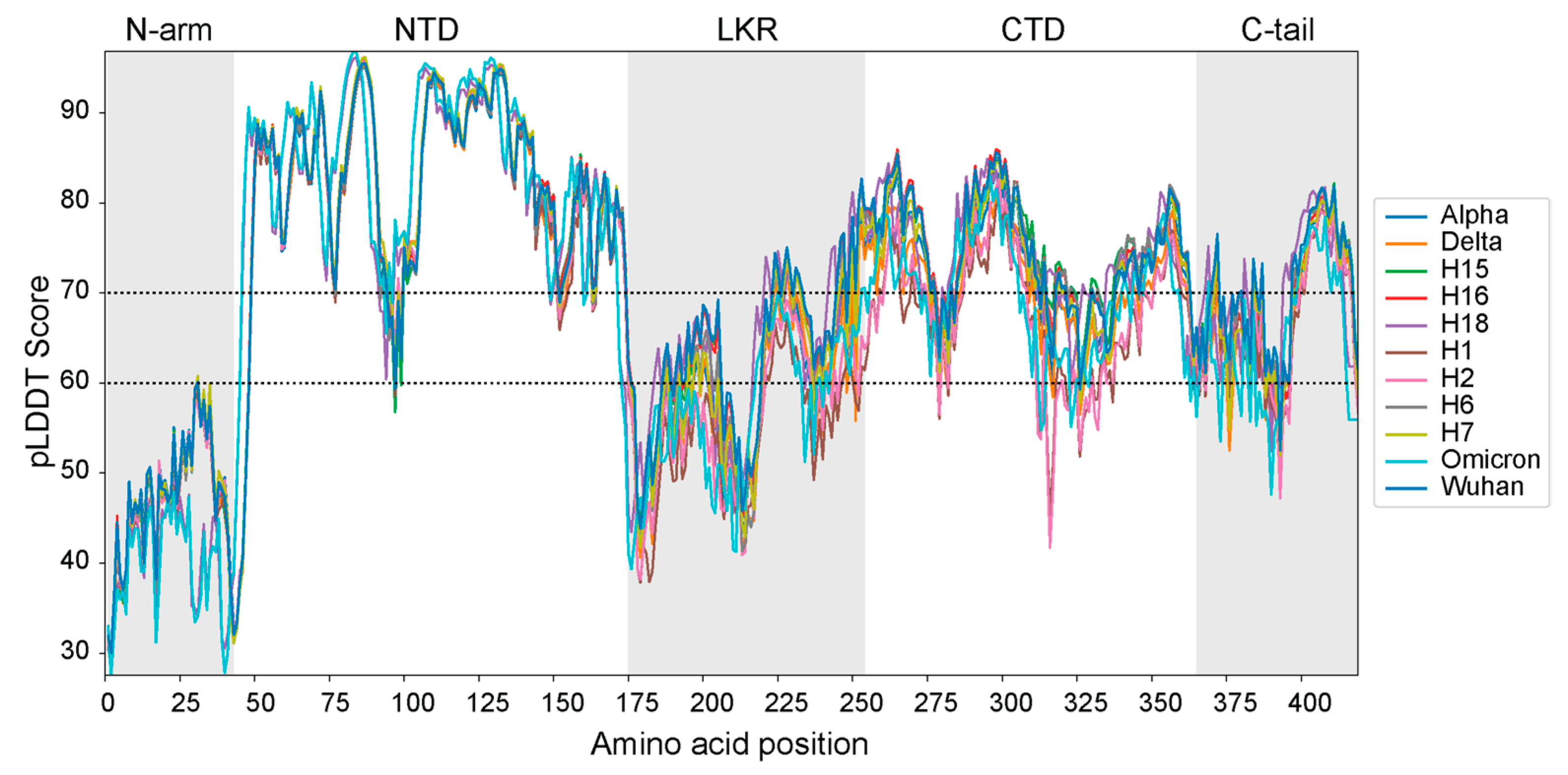
3.3. Protein Disorder and pLDDT Scores
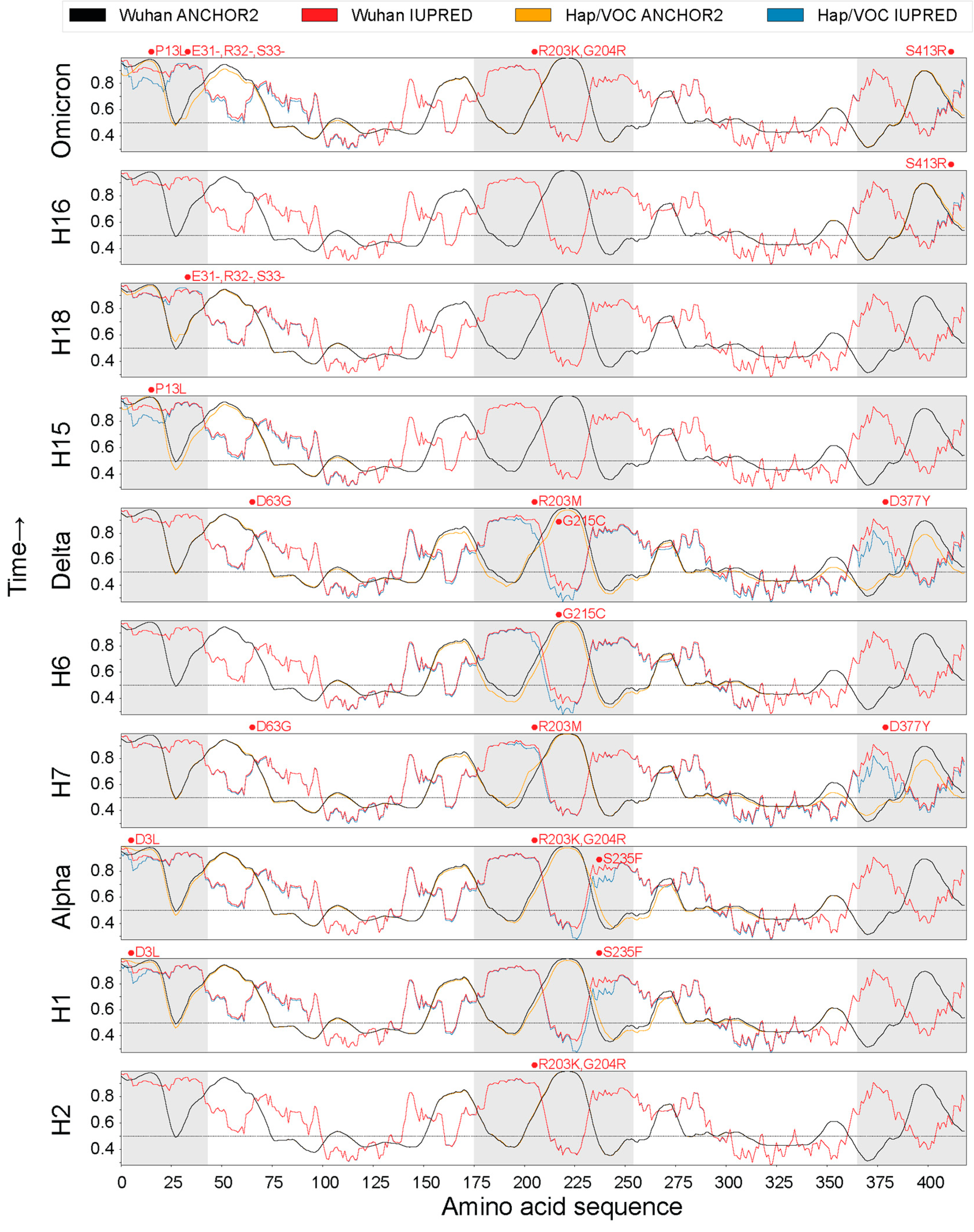
3.4. Protein Disorder and Binding Capacity across the Pandemic
3.5. Benchmarking AlphaFold2 Reference Structures against Experimental Cryo-EM Models
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- WHO COVID-19 Dashboard, COVID-19 Deaths. Available online: https://data.who.int/dashboards/covid19/cases (accessed on 1 July 2024).
- StatsAmerica, States in Profile, Population estimate for 2023, Ranked list. Available online: https://www.statsamerica.org/sip/rank_list.aspx?rank_label=pop1&ct=S18 (accessed on 1 July 2024).
- United Nations, Department of Economic and Social Affairs, Population Division, World Population Prospects. Available online: https://population.un.org/wpp/Download/Standard/Population/ (accessed on 1 July 2024).
- Ramadan, N.; Shaib, H. Middle East Respiratory Syndrome Coronavirus (MERS-CoV): A Review. Germs 2019, 9, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Hodgens, A.; Gupta, V. Severe Acute Respiratory Syndrome. In StatPearls; StatPearls Publishing: Treasure Island (FL), 2022. [Google Scholar]
- Summary of Probable SARS Cases with Onset of Illness from 1 November 2002 to 31 July 2003 Available online:. Available online: https://www.who.int/publications/m/item/summary-of-probable-sars-cases-with-onset-of-illness-from-1-november-2002-to-31-july-2003 (accessed on 1 July 2024).
- MERS-CoV Worldwide Overview Available online:. Available online: https://www.ecdc.europa.eu/en/middle-east-respiratory-syndrome-coronavirus-mers-cov-situation-update (accessed on 1 July 2024).
- Pormohammad, A.; Ghorbani, S.; Khatami, A.; Farzi, R.; Baradaran, B.; Turner, D.L.; Turner, R.J.; Bahr, N.C.; Idrovo, J. Comparison of Confirmed COVID-19 with SARS and MERS Cases - Clinical Characteristics, Laboratory Findings, Radiographic Signs and Outcomes: A Systematic Review and Meta-analysis. Rev Med Virol 2020, 30, e2112. [Google Scholar] [CrossRef] [PubMed]
- Pustake, M.; Tambolkar, I.; Giri, P.; Gandhi, C. SARS, MERS and CoVID-19: An Overview and Comparison of Clinical, Laboratory and Radiological Features. J Family Med Prim Care 2022, 11, 10–17. [Google Scholar] [CrossRef] [PubMed]
- Zou, L.; Ruan, F.; Huang, M.; Liang, L.; Huang, H.; Hong, Z.; Yu, J.; Kang, M.; Song, Y.; Xia, J.; et al. SARS-CoV-2 Viral Load in Upper Respiratory Specimens of Infected Patients. New England Journal of Medicine 2020, 382, 1177–1179. [Google Scholar] [CrossRef] [PubMed]
- Mizumoto, K.; Kagaya, K.; Zarebski, A.; Chowell, G. Estimating the Asymptomatic Proportion of Coronavirus Disease 2019 (COVID-19) Cases on Board the Diamond Princess Cruise Ship, Yokohama, Japan, 2020. Eurosurveillance 2020, 25, 2000180. [Google Scholar] [CrossRef]
- Liu, P.L. COVID-19 Information on Social Media and Preventive Behaviors: Managing the Pandemic through Personal Responsibility. Soc Sci Med 2021, 277, 113928. [Google Scholar] [CrossRef]
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L.; et al. A Pneumonia Outbreak Associated with a New Coronavirus of Probable Bat Origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef]
- Tomaszewski, T.; Ali, M.A.; Caetano-Anollés, K.; Caetano-Anollés, G. Seasonal Effects Decouple SARS-CoV-2 Haplotypes Worldwide 2023.
- Ali, M.A.; Caetano-Anollés, G. AlphaFold2 Reveals Structural Patterns of Seasonal Haplotype Diversification in SARS-CoV-2 Spike Protein Variants. Biology 2024, 13, 134. [Google Scholar] [CrossRef]
- Wu, W.; Cheng, Y.; Zhou, H.; Sun, C.; Zhang, S. The SARS-CoV-2 Nucleocapsid Protein: Its Role in the Viral Life Cycle, Structure and Functions, and Use as a Potential Target in the Development of Vaccines and Diagnostics. Virol J 2023, 20, 6. [Google Scholar] [CrossRef]
- Tomaszewski, T.; DeVries, R.S.; Dong, M.; Bhatia, G.; Norsworthy, M.D.; Zheng, X.; Caetano-Anollés, G. New Pathways of Mutational Change in SARS-CoV-2 Proteomes Involve Regions of Intrinsic Disorder Important for Virus Replication and Release. Evol Bioinform Online 2020, 16, 1176934320965149. [Google Scholar] [CrossRef]
- Peng, Y.; Du, N.; Lei, Y.; Dorje, S.; Qi, J.; Luo, T.; Gao, G.F.; Song, H. Structures of the SARS-CoV-2 Nucleocapsid and Their Perspectives for Drug Design. The EMBO Journal 2020, 39, e105938. [Google Scholar] [CrossRef] [PubMed]
- Ye, Q.; West, A.M.V.; Silletti, S.; Corbett, K.D. Architecture and Self-assembly of the SARS-CoV-2 Nucleocapsid Protein. Protein Sci 2020, 29, 1890–1901. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.-K.; Hsu, Y.-L.; Chang, Y.-H.; Chao, F.-A.; Wu, M.-C.; Huang, Y.-S.; Hu, C.-K.; Huang, T.-H. Multiple Nucleic Acid Binding Sites and Intrinsic Disorder of Severe Acute Respiratory Syndrome Coronavirus Nucleocapsid Protein: Implications for Ribonucleocapsid Protein Packaging. J Virol 2009, 83, 2255–2264. [Google Scholar] [CrossRef] [PubMed]
- Hilser, V.J.; Thompson, E.B. Intrinsic Disorder as a Mechanism to Optimize Allosteric Coupling in Proteins. Proceedings of the National Academy of Sciences 2007, 104, 8311–8315. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.; Hou, M.-H.; Chang, C.-F.; Hsiao, C.-D.; Huang, T. The SARS Coronavirus Nucleocapsid Protein – Forms and Functions. Antiviral Research 2014, 103, 39–50. [Google Scholar] [CrossRef] [PubMed]
- Lu, S.; Ye, Q.; Singh, D.; Cao, Y.; Diedrich, J.K.; Yates, J.R.; Villa, E.; Cleveland, D.W.; Corbett, K.D. The SARS-CoV-2 Nucleocapsid Phosphoprotein Forms Mutually Exclusive Condensates with RNA and the Membrane-Associated M Protein. Nat Commun 2021, 12, 502. [Google Scholar] [CrossRef] [PubMed]
- Oh, S.J.; Shin, O.S. SARS-CoV-2 Nucleocapsid Protein Targets RIG-I-Like Receptor Pathways to Inhibit the Induction of Interferon Response. Cells 2021, 10, 530. [Google Scholar] [CrossRef] [PubMed]
- de Haan, C.A.M.; Rottier, P.J.M. Molecular Interactions in the Assembly of Coronaviruses. Adv Virus Res 2005, 64, 165–230. [Google Scholar] [CrossRef] [PubMed]
- Klein, S.; Cortese, M.; Winter, S.L.; Wachsmuth-Melm, M.; Neufeldt, C.J.; Cerikan, B.; Stanifer, M.L.; Boulant, S.; Bartenschlager, R.; Chlanda, P. SARS-CoV-2 Structure and Replication Characterized by in Situ Cryo-Electron Tomography 2020, 2020.06.23.167064.
- Stertz, S.; Reichelt, M.; Spiegel, M.; Kuri, T.; Martínez-Sobrido, L.; García-Sastre, A.; Weber, F.; Kochs, G. The Intracellular Sites of Early Replication and Budding of SARS-Coronavirus. Virology 2007, 361, 304–315. [Google Scholar] [CrossRef]
- Lauring, A.S.; Hodcroft, E.B. Genetic Variants of SARS-CoV-2—What Do They Mean? JAMA 2021, 325, 529–531. [Google Scholar] [CrossRef] [PubMed]
- Agnihotry, S.; Pathak, R.K.; Singh, D.B.; Tiwari, A.; Hussain, I. Chapter 11 - Protein Structure Prediction. In Bioinformatics; Singh, D.B., Pathak, R.K., Eds.; Academic Press, 2022; pp. 177–188 ISBN 978-0-323-89775-4.
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly Accurate Protein Structure Prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Kryshtafovych, A.; Schwede, T.; Topf, M.; Fidelis, K.; Moult, J. Critical Assessment of Methods of Protein Structure Prediction (CASP)—Round XIV. Proteins: Structure, Function, and Bioinformatics 2021, 89, 1607–1617. [Google Scholar] [CrossRef] [PubMed]
- Kryshtafovych, A.; Schwede, T.; Topf, M.; Fidelis, K.; Moult, J. Critical Assessment of Methods of Protein Structure Prediction (CASP)—Round XV. Proteins: Structure, Function, and Bioinformatics 2023, 91, 1539–1549. [Google Scholar] [CrossRef]
- Ford, C.T.; Jacob Machado, D.; Janies, D.A. Predictions of the SARS-CoV-2 Omicron Variant (B.1.1.529) Spike Protein Receptor-Binding Domain Structure and Neutralizing Antibody Interactions. Front. Virol. 2022, 2. [Google Scholar] [CrossRef]
- Kilim, O.; Mentes, A.; Pál, B.; Csabai, I.; Gellért, Á. SARS-CoV-2 Receptor-Binding Domain Deep Mutational AlphaFold2 Structures. Sci Data 2023, 10, 134. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Jian, X.; Syed, A.A.S.; Fahira, A.; Zheng, C.; Zhu, Z.; Wang, K.; Zhang, J.; Wen, Y.; Li, Z.; et al. Structural Comparison and Drug Screening of Spike Proteins of Ten SARS-CoV-2 Variants. Research 2022, 2022. [Google Scholar] [CrossRef]
- Tomaszewski, T.; Gurtler, V.; Caetano-Anollés, K.; Caetano-Anollés, G. Chapter 8 - The Emergence of SARS-CoV-2 Variants of Concern in Australia by Haplotype Coalescence Reveals a Continental Link to COVID-19 Seasonality. In Methods in Microbiology; Pavia, C.S., Gurtler, V., Eds.; Covid-19: Biomedical Perspectives; Academic Press, 2022; Vol. 50, pp. 233–268.
- Burra, P.; Soto-Díaz, K.; Chalen, I.; Gonzalez-Ricon, R.J.; Istanto, D.; Caetano-Anollés, G. Temperature and Latitude Correlate with SARS-CoV-2 Epidemiological Variables but Not with Genomic Change Worldwide. Evol Bioinform Online 2021, 17, 1176934321989695. [Google Scholar] [CrossRef] [PubMed]
- Hernandez, N.; Caetano-Anollés, G. Worldwide Correlations Support COVID-19 Seasonal Behavior and Impact of Global Change. Evolutionary Bioinformatics 2023, 19, 11769343231169377. [Google Scholar] [CrossRef]
- Mirdita, M.; Schütze, K.; Moriwaki, Y.; Heo, L.; Ovchinnikov, S.; Steinegger, M. ColabFold: Making Protein Folding Accessible to All. Nat Methods 2022, 19, 679–682. [Google Scholar] [CrossRef]
- Mariani, V.; Biasini, M.; Barbato, A.; Schwede, T. lDDT: A Local Superposition-Free Score for Comparing Protein Structures and Models Using Distance Difference Tests. Bioinformatics 2013, 29, 2722–2728. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera--a Visualization System for Exploratory Research and Analysis. J Comput Chem 2004, 25, 1605–1612. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Skolnick, J. Scoring Function for Automated Assessment of Protein Structure Template Quality. Proteins 2004, 57, 702–710. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Shine, M.; Pyle, A.M.; Zhang, Y. US-Align: Universal Structure Alignments of Proteins, Nucleic Acids, and Macromolecular Complexes. Nat Methods 2022, 19, 1109–1115. [Google Scholar] [CrossRef] [PubMed]
- Zemla, A. LGA: A Method for Finding 3D Similarities in Protein Structures. Nucleic Acids Res 2003, 31, 3370–3374. [Google Scholar] [CrossRef] [PubMed]
- Zemla, A.; Zhou, C.E.; Slezak, T.; Kuczmarski, T.; Rama, D.; Torres, C.; Sawicka, D.; Barsky, D. AS2TS System for Protein Structure Modeling and Analysis. Nucleic Acids Res 2005, 33, W111–115. [Google Scholar] [CrossRef] [PubMed]
- LGA - Protein Structure Comparison Facility Available online:. Available online: http://proteinmodel.org/AS2TS/LGA/lga.html (accessed on 2 July 2024).
- Chothia, C.; Lesk, A.M. The Relation between the Divergence of Sequence and Structure in Proteins. The EMBO Journal 1986, 5, 823–826. [Google Scholar] [CrossRef] [PubMed]
- Reva, B.A.; Finkelstein, A.V.; Skolnick, J. What Is the Probability of a Chance Prediction of a Protein Structure with an Rmsd of 6 å? Folding and Design 1998, 3, 141–147. [Google Scholar] [CrossRef] [PubMed]
- Smith, C.C.; Olsen, K.S.; Gentry, K.M.; Sambade, M.; Beck, W.; Garness, J.; Entwistle, S.; Willis, C.; Vensko, S.; Woods, A.; et al. Landscape and Selection of Vaccine Epitopes in SARS-CoV-2. Genome Med 2021, 13, 101. [Google Scholar] [CrossRef]
- TM-Score: Quantitative Assessment of Similarity between Protein Structures Available online:. Available online: https://zhanggroup.org/TM-score/ (accessed on 1 July 2024).
- Akdel, M.; Pires, D.E.V.; Pardo, E.P.; Jänes, J.; Zalevsky, A.O.; Mészáros, B.; Bryant, P.; Good, L.L.; Laskowski, R.A.; Pozzati, G.; et al. A Structural Biology Community Assessment of AlphaFold2 Applications. Nat Struct Mol Biol 2022, 29, 1056–1067. [Google Scholar] [CrossRef]
- Bruley, A.; Mornon, J.-P.; Duprat, E.; Callebaut, I. Digging into the 3D Structure Predictions of AlphaFold2 with Low Confidence: Disorder and Beyond. Biomolecules 2022, 12, 1467. [Google Scholar] [CrossRef] [PubMed]
- Wilson, C.J.; Choy, W.-Y.; Karttunen, M. AlphaFold2: A Role for Disordered Protein/Region Prediction? Int J Mol Sci 2022, 23, 4591. [Google Scholar] [CrossRef]
- Alderson, T.R.; Pritišanac, I.; Kolarić, Đ.; Moses, A.M.; Forman-Kay, J.D. Systematic Identification of Conditionally Folded Intrinsically Disordered Regions by AlphaFold2. Proc Natl Acad Sci U S A 120, e2304302120. [CrossRef] [PubMed]
- Mészáros, B.; Erdős, G.; Dosztányi, Z. IUPred2A: Context-Dependent Prediction of Protein Disorder as a Function of Redox State and Protein Binding. Nucleic Acids Research 2018, 46, W329–W337. [Google Scholar] [CrossRef]
- Cubuk, J.; Alston, J.J.; Incicco, J.J.; Singh, S.; Stuchell-Brereton, M.D.; Ward, M.D.; Zimmerman, M.I.; Vithani, N.; Griffith, D.; Wagoner, J.A.; et al. The SARS-CoV-2 Nucleocapsid Protein Is Dynamic, Disordered, and Phase Separates with RNA. Nat Commun 2021, 12, 1936. [Google Scholar] [CrossRef] [PubMed]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Research 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Bank, R.P.D. RCSB PDB: Homepage Available online:. Available online: https://www.rcsb.org/ (accessed on 1 July 2024).
- Bai, Z.; Cao, Y.; Liu, W.; Li, J. The SARS-CoV-2 Nucleocapsid Protein and Its Role in Viral Structure, Biological Functions, and a Potential Target for Drug or Vaccine Mitigation. Viruses 2021, 13, 1115. [Google Scholar] [CrossRef]
- Peng, T.-Y.; Lee, K.-R.; Tarn, W.-Y. Phosphorylation of the Arginine/Serine Dipeptide-Rich Motif of the Severe Acute Respiratory Syndrome Coronavirus Nucleocapsid Protein Modulates Its Multimerization, Translation Inhibitory Activity and Cellular Localization. The FEBS Journal 2008, 275, 4152–4163. [Google Scholar] [CrossRef] [PubMed]
- Dinesh, D.C.; Chalupska, D.; Silhan, J.; Koutna, E.; Nencka, R.; Veverka, V.; Boura, E. Structural Basis of RNA Recognition by the SARS-CoV-2 Nucleocapsid Phosphoprotein. PLoS Pathog 2020, 16, e1009100. [Google Scholar] [CrossRef]
- Takeda, M.; Chang, C.; Ikeya, T.; Güntert, P.; Chang, Y.; Hsu, Y.; Huang, T.; Kainosho, M. Solution Structure of the C-Terminal Dimerization Domain of SARS Coronavirus Nucleocapsid Protein Solved by the SAIL-NMR Method. Journal of Molecular Biology 2008, 380, 608–622. [Google Scholar] [CrossRef]
- Chen, C.-Y.; Chang, C.; Chang, Y.-W.; Sue, S.-C.; Bai, H.-I.; Riang, L.; Hsiao, C.-D.; Huang, T. Structure of the SARS Coronavirus Nucleocapsid Protein RNA-Binding Dimerization Domain Suggests a Mechanism for Helical Packaging of Viral RNA. Journal of Molecular Biology 2007, 368, 1075–1086. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.; Chen, C.-M.M.; Chiang, M.; Hsu, Y.; Huang, T. Transient Oligomerization of the SARS-CoV N Protein--Implication for Virus Ribonucleoprotein Packaging. PLoS One 2013, 8, e65045. [Google Scholar] [CrossRef] [PubMed]
- Surjit, M.; Kumar, R.; Mishra, R.N.; Reddy, M.K.; Chow, V.T.K.; Lal, S.K. The Severe Acute Respiratory Syndrome Coronavirus Nucleocapsid Protein Is Phosphorylated and Localizes in the Cytoplasm by 14-3-3-Mediated Translocation. Journal of Virology 2005, 79, 11476–11486. [Google Scholar] [CrossRef] [PubMed]
| State-Chain | GDT-TS (AS2S) | TM Score/L (US-Align Server) | Superimposed RMSD < 5 Å (AS2S) |
|---|---|---|---|
| NTD-A (7CDZ) | 91.99*(128/131) = 89.88 |
*L1: 0.93398, L2: 0.91375 |
1.293/128 |
| NTD-B (7CDZ) | 90.35*(127/131) = 87.59 |
0.92756, 0.90091 |
1.366/127 |
| NTD-C (7CDZ) | 95.24*(126/131) = 91.60 |
0.95983, 0.92447 |
0.941/126 |
| NTD-D (7CDZ) | 94.26*(122/131) = 87.78 |
0.93300, 0.90592 |
1.1108/122 |
| CTD-A (7CE0) | 69.73*(109/110) = 69.10 |
0.74156 | 2.508/109 |
| CTD-B (7CE0) | 69.50*(109/110) = 68.87 |
0.73968 | 2.519/109 |
| CTD-C (7CE0) | 69.39*(107/110) = 67.50 |
0.73391 | 2.476/107 |
| CTD-D (7CE0) | 68.93*(107/110) = 67.05 |
0.73302 | 2.490/107 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





