Submitted:
22 July 2024
Posted:
23 July 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
- (1)
- We propose a simple yet effective NER model that integrates enhanced lexicon and contrastive learning for the complex pig disease domain, making the model more sensitive to texts in this domain and improving predictions for entities. The lexicon-enhanced BERT facilitates the direct integration of external lexicon knowledge of pig diseases into BERT layers via a Lexicon Adapter layer.
- (2)
- To enrich the semantic feature representation and improve performance under data scarcity conditions, we propose a lexicon-enhanced contrastive loss layer on top of the BERT encoder. Experimental results on small sample scenarios and common public datasets demonstrate that our model outperforms other models.
- (3)
- Given the lack of an annotated corpus for the pig disease domain, we collected and annotated a new Chinese corpus and annotated datasets consisting of 7,518 entities. To address the insensitivity of word segmentation caused by the specialization of the pig disease domain, we constructed a lexicon for identifying specific terms in pig diseases using frequency statistics methods under the guidance of veterinarians.
2. Materials and Methods
2.1. Materials
2.1.1. Corpus Collection and Pre-Processing
2.1.2. Corpus Annotation
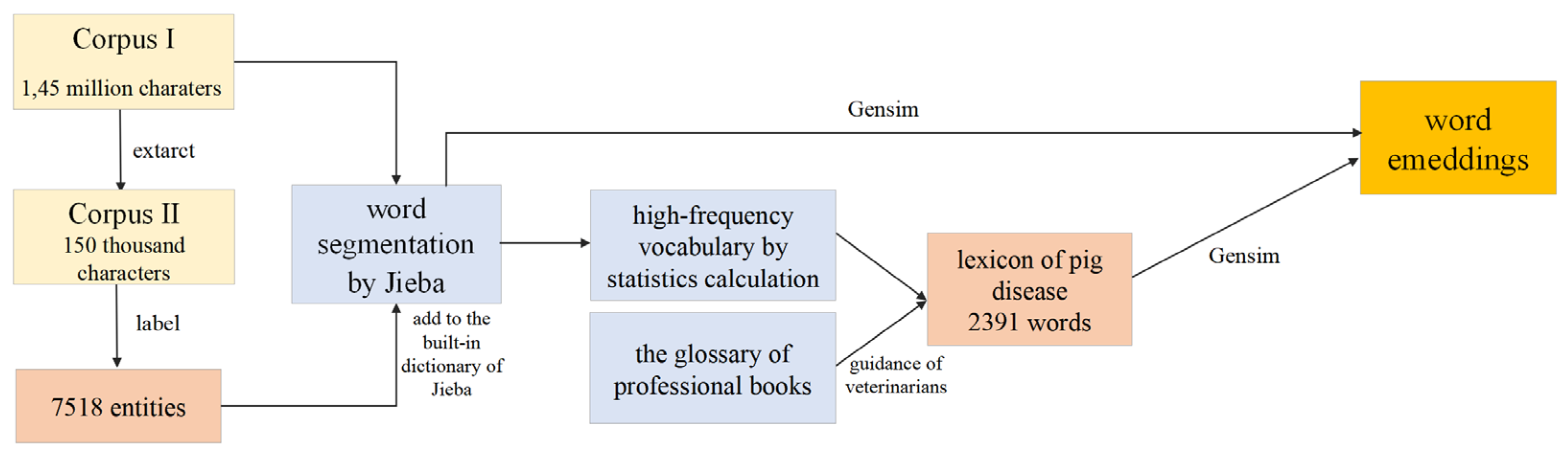
2.1.3. Construction of Lexicon and Pre-Training Word Embedding
2.2. Methods
2.2.2. Char-Words Pair Sequence
2.2.3. Lexicon Adapter
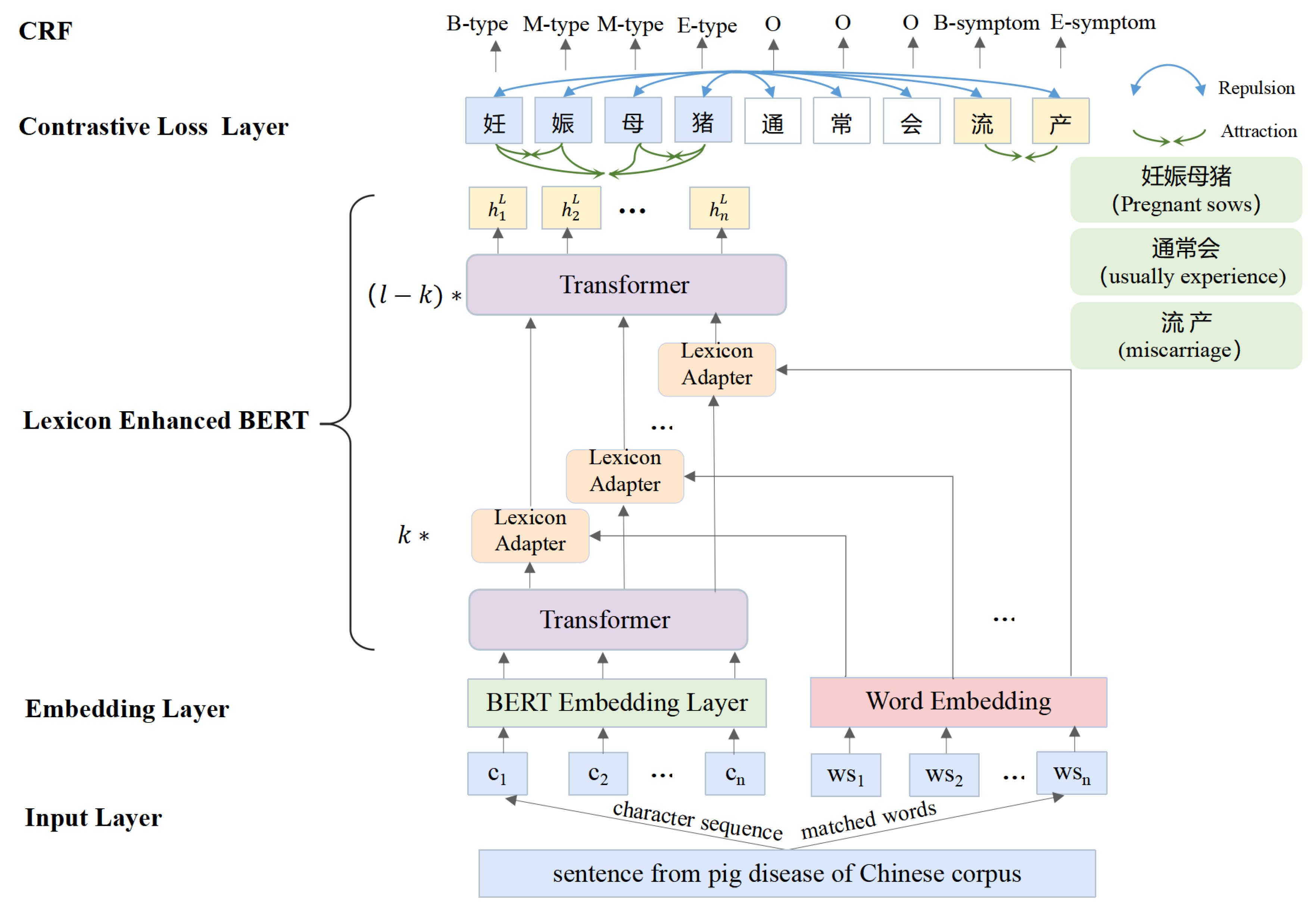
2.2.4. Lexicon Enhanced BERT
2.2.5. Lexicon Enhanced Contrastive Learning
3. Experiment and Results Analysis
3.1. Evaluation
3.2. Experimental Settings
3.3. Experiment Results and Analysis
3.3.1. Comparison with Baseline Models
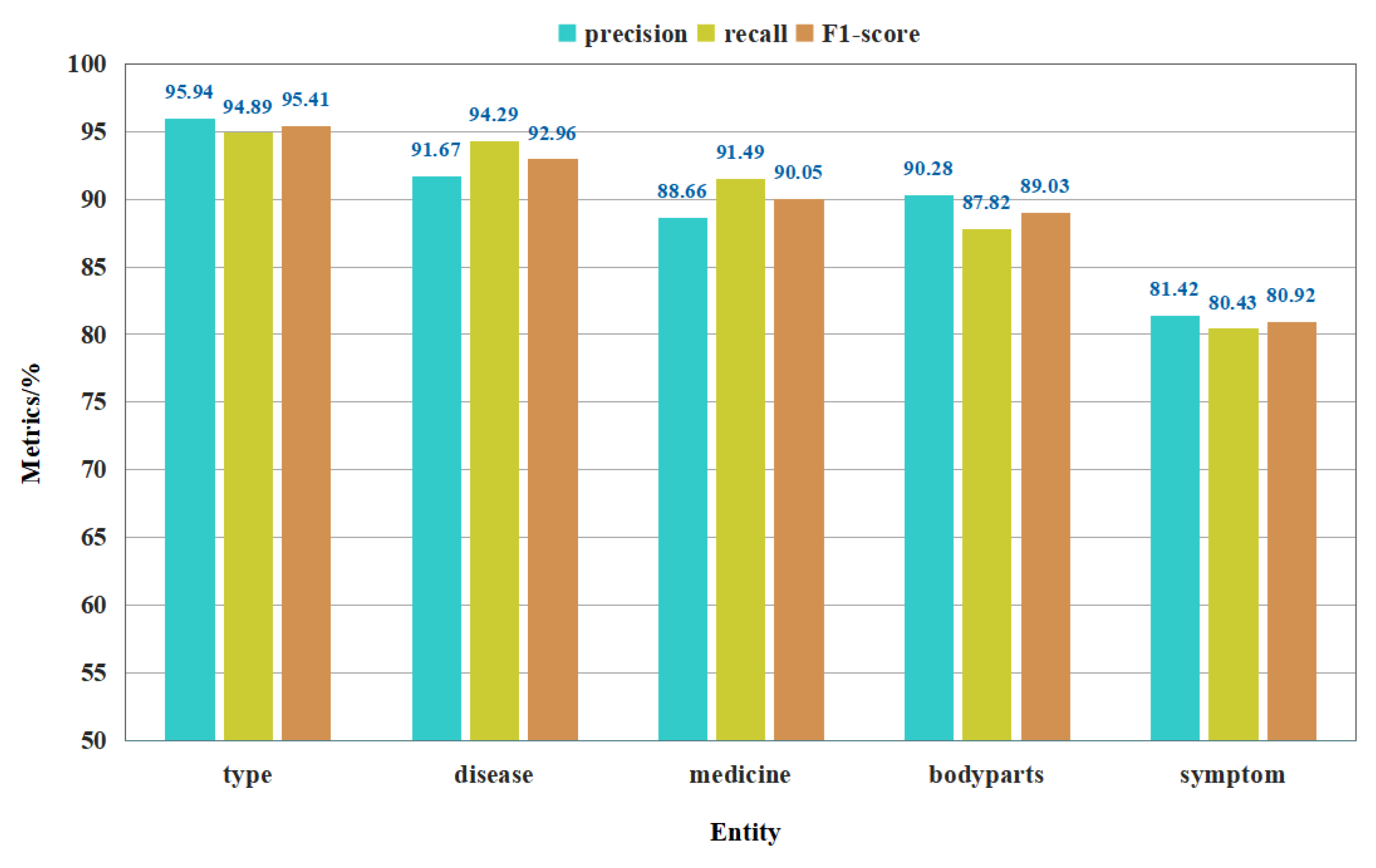
3.3.2. The Recognition Effect on Different Entities
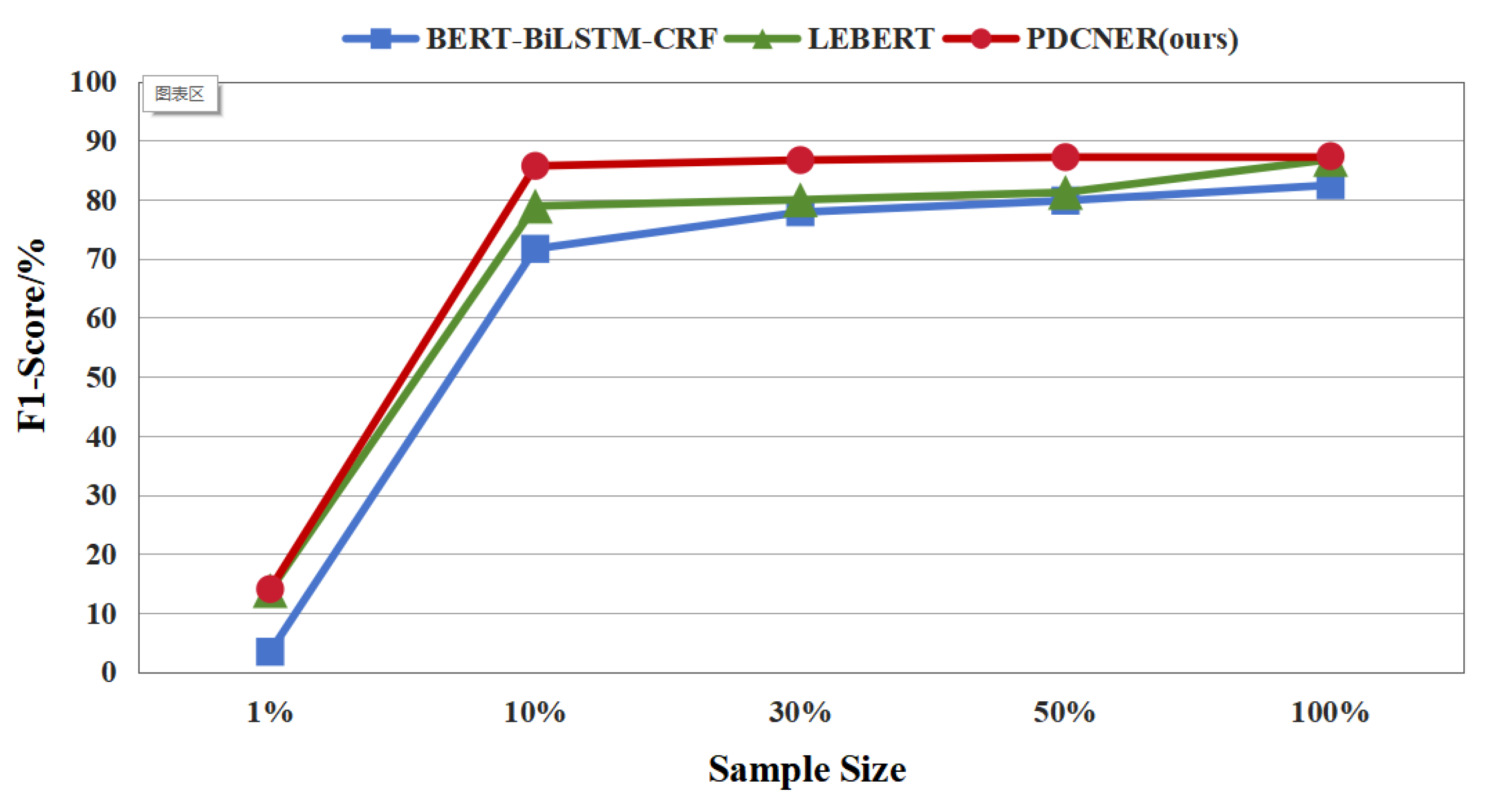
3.3.3. The Recognition Effect on Small Sample
3.3.4. The Recognition Effect on Public Datasets
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A. Data Source Details
| No | Type | Example |
| 1 | Professional books | Jeffrey J. Zimmerman, Locke A.Kerri, Alejandro Raminez.etc,editor in chief. Hanchun Yang,main translation. Disease of Swine. North United publishing media Media Co., Ltd., Liaoning science and technology publishing house:Beijing, China, 2022. Yousheng, Xu. Primary color atlas of scientific pig raising and pig disease prevention and control. China Agricultural Publishing House: Beijing, China,2017. Changyou,Li, Xiaocheng,Li. Prevention and control technology of swine epidemic disease. China Agricultural Publishing House:Beijing, China,2015. Jianxin Zhang. Diagnosis and control of herd pig epidemic disease. He ‘nan Science and Technology Press:Zhengzhou, China, 2014. Chaoying, Luo, Guibo, Wang. Prevention and treatment of pig diseases and safe medication. Chemical industry press:Beijing, China,2016, etc. 《猪病学》, 《科学养猪与猪病防制原色图谱》, 《猪群疫病防治技术》, 《群养猪疫病诊断与控制》, 《猪病防治及安全用药》等 |
| 2 | Standard specification | Technical Specification for Quarantine of Porcine Reproductive and Respiratory Syndrome (SN/T 1247-2022), Diagnostic Techniques for Mycoplasma Pneumonia in Swine (NY/T 1186-2017), Diagnostic Techniques for Infectious Pleuropneumonia in Swine (NY/T 537-2023), Diagnostic Techniques for Swine Dysentery (NY/T 545-2023), Technical Specification for Quarantine of Porcine Rotavirus Infection (SN/T 5196-2020), etc. 《猪繁殖与呼吸综合征检疫技术规范》 (SN/T 1247-2022), 《猪支原体肺炎诊断技术》 (NY/T 1186-2017), 《猪传染性胸膜肺炎诊断技术》 (NY/T 537-2023), 《猪痢疾诊断技术》(NY/T 545-2023), 《猪轮状病毒感染检疫技术规范》(SN/T 5196-2020) 等 |
| 3 | Technological specification | Technical specification for prevention and control of highly pathogenic blue ear disease in pigs, technical specification for prevention and control of foot-and-mouth disease, technical specification for prevention and control of classical swine fever, etc. 《高致病性猪蓝耳病防治技术规范》, 《口蹄疫防治技术规范》, 《猪瘟防治技术规范》等 |
| 4 | Policy paper | Ministry of Agriculture and Rural Affairs “List of Class I, II and III Animal Diseases”, The Ministry of Agriculture issued the “Guiding Opinions on Prevention and Control of Highly Pathogenic Porcine Blue Ear Disease (2017-2020)”, Notice of National Guiding Opinions on Prevention and Control of Classical Swine Fever (2017-2020), etc. 农业农村部《一, 二, 三类动物疫病病种名录》, 农业部关于印发《国家高致病性猪蓝耳病防治指导意见(2017—2020年)》, 《国家猪瘟防治指导意见 (2017—2020年)》的通知 |
| 5 | Relevant industry website. | China Veterinary Website(https://www.cadc.net.cn/sites/MainSite/), Big Animal Husbandry Website(https://www.dxumu.com/), Huinong Website(https://www.cnhnb.com/), etc. 中国兽医网, 大畜牧网, 惠农网等 |
References
- Li, J.; Sun, A.; Han, J.; Li, C. A Survey on Deep Learning for Named Entity Recognition. IEEE Transactions on Knowledge and Data Engineering, 2022, 34, 50–70. [Google Scholar] [CrossRef]
- Cheng, J.; Liu, J.; Xu, X.; Xia, D.; Liu, L.; Sheng, V.S. A review of Chinese named entity recognition. KSII Trans. Internet Inf. Syst. 2021, 15, 2012–2030. [Google Scholar] [CrossRef]
- Mi, B.G.; Fan, Y. A review: development of named entity recognition (NER) technology for aeronautical information intelligence. Artif. Intell. Rev. 2022, 56, 1515–1542. [Google Scholar]
- Liu, P.; Guo, Y.; Wang, F.; Li, G. Chinese named entity recognition: The state of the art. Neuro Comput. 2022, 473, 37–53. [Google Scholar] [CrossRef]
- Qiu, X.; Sun, T.; Xu, Y.; Shao, Y.; Dai, N.; Huang, X. Pre-trained models for natural language processing: A survey. Sci. China Technol. Sci. 2020, 63, 1872–1897. [Google Scholar] [CrossRef]
- Zhang, S.; Elhadad, N. Unsupervised biomedical named entity recognition: Experiments with clinical and biological texts. J. Biomed. Informatics 2013, 46, 1088–1098. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.-J.; Zhang, T. Research on Me-based Chinese NER model. In Proceeding of 7th International Conference on Machine Learning and Cybernetics(ICMLC), IEEE Kunming, China, 12-15 July, 2008; 2597-2602.
- Hu, H.; Zhang, H. Chinese Named Entity Recognition with CRFs: Two Levels. In Proceeding of International Conference on Computational Intelligence&Security, IEEE, Suzhou, China, 2008; 6, 1-6.
- Kang, Y.; Sun, L.; Zhu, R.; Meng, L. Survey on Chinese named entity recognition with deep learning. J. Huazhong Univ. Sci. Technol. Nat. Sci. Ed. 2022, 50, 44–53. [Google Scholar]
- Yue, Z.; Ji, Y. Chinese NER using lattice LSTM. In Proceedings of the 56th Annual Meeting of the Association for Computational Linguistics, Melbourne, Australia, 2018; 1554-1564.
- Tao, G.; Ruotian, M.; Qi, Z.; et al. CNN-Based Chinese NER with lexicon rethinking. In Proceedings of the 28th International Joint Conference on Artificial Intelligence, Macao, China, AAAI Press, 2019; 8, 4982-4988.
- Gui, T.; Zou, Y.; Zhang, Q.; Peng, M.; Fu, J.; Wei, Z. ; Huang, X-J. A Lexicon-Based Graph Neural Network for Chinese NER. In Proceeding. of the 2019 Conference on Empirical Methods in Natural Language Processing and the 9th International Joint Conference on Natural Language Processing (EMNLP-IJCNLP), Hong Kong, China, 3-7 Nov, 2019; 1040-1050.
- Liu, W.; Fu, X.; Zhang, Y.; Xiao, W. Lexicon enhanced Chinese sequence labeling using BERT adapter. In Proceeding of 59th Annual Meeting of the Association for Computational Linguistics and the 11th International Joint Conference on Natural Language Processing, Bangkok, Thailand, 1-6 August 2021; arXiv2021.
- Ruotian, M.; Minling, P.; Qi, Z.; et al. Simplify the usage of lexicon in Chinese NER. In Proceedings of the 58th Annual Meeting of the Association for Computational Linguistics. Online: Association for Computational Linguistics, 2020, 5951-5960.
- Xiaonan, L.; Huang, Y.; Xipeng, Q.; et al. FLAT: Chinese NER using flat-lattice transformer. In Proceedings of the 58th Annual Meeting of the Association for Computational Linguistics, ACL Press, 2020; 6836–6842.
- Mengge, X.; Yu, B.; Liu, T.; Zhang, Y.; Meng, E.; Wang, B. Porous lattice transformer encoder for Chinese NER. In Proceedings of the 28th International Conference on Computational Linguistics. International Committee on Computational Linguistics, Barcelona, Spain, 13-18 Sep 2020; pp: 3831-3841.
- Devlin, J.; Chang, M.-W.; Lee, K.; Toutanova, K. BERT: Pre-training of Deep Bidirectional Transformers for Language Understanding. arXiv 2018, arXiv:1810.04805. [Google Scholar]
- Peters, M.; Neumann, M.; Iyyer, M.; Gardner, M.; Clark, C.; Lee, K.; Zettlemoyer, L. Deep contextualized word representations. arXiv 2018, arXiv:1802.05365. [Google Scholar]
- Liu, Y.; Ott, M.; Goyal, N.; Du, J.; Joshi, M.; Chen, D.; Levy, O.; Lewis, M.; Zettlemoyer, L.; Stoyanov, V. RoBERTa: A robustly optimized BERT pretraining approach. arXiv 2019, arXiv:1907.11692. [Google Scholar]
- Sun, Y.; Wang, S.; Li, Y.; Feng, S.; Chen, X.; Zhang, H.; Tian, X.; Zhu, D.; Tian, H.; Wu, H. ERNIE: enhanced representation through knowledge integration. arXiv 2019, arXiv:1904.09223v1. [Google Scholar]
- Lan, Z.; Chen, M.; Goodman, S.; Gimpel, K.; Sharma, P.; Soricut, R. ALBERT: a lite BERT for self-supervised learning of language representations. 8th International Conference on Learning Representations(ICLR), Addis Ababa, Ethiopia, 26-30 April 2020.
- Li, S.; Bao, Z.; Zhao, S.; Jiang, G.; Shan, L.; Zhang, L. A LEBERT-based model for named entity recognition. In Proceedings of the 2021 3rd International Conference on Artificial Intelligence and Advanced Manufacture(AIAM), ACM International Conference Proceeding Series, Manchester, UK, 23-25 Oct 2021; pp: 980-983. [Google Scholar]
- Yan, Y.; Li, R.; Wang, S.; Zhang, F.; Wu, W.; Xu, W. ConSERT: A contrastive framework for self-supervised sentence representation transfer. arXiv 2021, arXiv:2105.11741. [Google Scholar]
- Gao, T.; Yao, X.; Chen, D. SimCSE: Simple Contrastive Learning of Sentence Embeddings. EMNLP, Conference on Empirical Methods in Natural Language Processing. Punta Cana, Dominican Republic, 7-11 Nov 2021; 6894-6910.
- Huang, Y.; He, K.; Wang, Y.; et al. COPNER: Contrastive learning with prompt guiding for few-shot named entity recognition. In Proceedings of the 29th International Conference on Computational Linguistics. Gyeongju, Republic of Korea, 12-17 Oct 2022; pp: 2515-2527.
- He, K.; Mao, R.; Huang, Y.; Gong, T.; Li, C.; Cambria, E. Template-Free Prompting for Few-Shot Named Entity Recognition via Semantic-Enhanced Contrastive Learning. IEEE transactions on neural networks and learning systems, PP. [CrossRef]
- Li, X.; Li, X.; Zhao, M.; Yang, M.; Yu, R.; Yu, M.; Yu, J. CLINER: exploring task-relevant features and label semantic for few-shot named entity recognition. Neural Comput. Appl. 2023, 36, 4679–4691. [Google Scholar] [CrossRef]
- Chen, P.; Wang, J.; Lin, H.; Di, Z.; Yang, Z. Few-shot biomedical named entity recognition via knowledge-guided instance generation and prompt contrastive learning. Bioinformatics 2023, 39. [Google Scholar] [CrossRef]
- Sahadevan, S.; Hofmann-Apitius, M.; Schellander, K.; Tesfaye, D.; Fluck, J.; Friedrich, C.M. Text mining in livestock animal science: Introducing the potential of text mining to animal sciences 1. J. Anim. Sci. 2012, 90, 3666–3676. [Google Scholar] [CrossRef] [PubMed]
- Oh, H.-S.; Lee, H. Named Entity Recognition for Pet Disease Q&A System. J. Digit. Contents Soc. 2022, 23, 765–771. [Google Scholar]
- Kung, H.; Yu, R.; Chen, C.; Tsai, C.; Lin, C. Intelligent pig-raising knowledge question-answering system based on neural network schemes. Agron. J. 2021, 113, 906–922. [Google Scholar] [CrossRef]
- Zhang, D.; Zheng, G.; Liu, H.; Ma, X.; Xi, L. AWdpCNER: Automated Wdp Chinese Named Entity Recognition from Wheat Diseases and Pests Text. Agriculture 2023, 13, 1220. [Google Scholar] [CrossRef]
- Veena, G.; Kanjirangat, V.; Gupta, D. AGRONER: An unsupervised agriculture named entity recognition using weighted distributional semantic model. Expert Syst. Appl. 2023, 229. [Google Scholar] [CrossRef]
- Zhang, L.; Nie, X.; Zhang, M.; Gu, M.; Geissen, V.; Ritsema, C.J.; Niu, D.; Zhang, H. Lexicon and attention-based named entity recognition for kiwifruit diseases and pests: A Deep learning approach. Front. Plant Sci. 2022, 13, 1053449. [Google Scholar] [CrossRef]
- Guo, X.; Lu, S.; Tang, Z.; Bai, Z.; Diao, L.; Zhou, H.; Li, L. CG-ANER: Enhanced contextual embeddings and glyph features-based agricultural named entity recognition. Comput. Electron. Agric. 2022, 194, 106776. [Google Scholar] [CrossRef]
- Huang, B.; Lin, Y.; Pang, S.; Fu, L. Named Entity Recognition in Government Audit Texts Based on ChineseBERT and Character-Word Fusion. Appl. Sci. 2024, 14, 1425. [Google Scholar] [CrossRef]
- Guo, Y.; Feng, S.; Liu, F.; Lin, W.; Liu, H.; Wang, X.; Su, J.; Gao, Q. Enhanced Chinese Domain Named Entity Recognition: An Approach with Lexicon Boundary and Frequency Weight Features. Appl. Sci. 2023, 14, 354. [Google Scholar] [CrossRef]
- Jia, Y.C.; Zhu, D.J. Medical Named Entity Recognition Based on Deep Learning. Comput. Systems Applications 2022, 31, 70–81. [Google Scholar]
- Jin-hua, D.U.; Hao, Y.I.N.; Song, F. Research and Development of Named Entity Recognition in Chinese Electronic Medical Record. Acta Electron. Sin. 2022, 50, 3030–3053. [Google Scholar]
- Chen, T.; Kornblith, S.; Norouzi, M.; Hinton, G. A simple framework for contrastive learning of visual representations. arXiv 2020, arXiv:2002.05709. [Google Scholar]

| Category | Category definition | Examples | Numbers | Proportion of the total |
|---|---|---|---|---|
| Type | Name of different types of pig | 妊娠母猪, 仔猪 (Pregnant sows,piglets) | 735 | 9.78% |
| Disease | Name of pig disease | 猪丹毒, 胸膜炎 (Porcine erysipelas, pleurisy) | 958 | 12.74% |
| Body parts | Body position, organs and system of pigs | 心脏, 巨噬细胞 (Heart, Macrophages) | 2063 | 27.44% |
| Symptom | External performance | 气喘, 咳嗽, 水肿 (Asthma, cough,swollen) | 2973 | 39.55% |
| caused by diseases | ||||
| Medicine | Medications for treating diseases | 替米考星, 克林霉素 (Timicosin, clindamycin) | 789 | 10.49% |
| Total | 7518 | 100% |
| Model category | Model | P(%) | R(%) | F1(%) |
|---|---|---|---|---|
| baseline model without pre-trained | BILSTM_CRF | 71.58 | 67.51 | 69.49 |
| pre-trained model | BERT-BiLSTM-CRF | 80.29 | 84.73 | 82.45 |
| BERT-CRF | 81.62 | 84.39 | 82.98 | |
| BERT-CNN-CRF | 82.44 | 80.55 | 81.48 | |
| BERT-WWM-ext-BiLSTM-CRF | 81.73 | 85.47 | 83.56 | |
| RoBERTa-BiLSTM-CRF | 81.64 | 85.31 | 83.43 | |
| pre-trained model with lexicon | BERT-BILSTM-CRF-SoftLexicon | 82.99 | 84.73 | 83.85 |
| LEBERT | 87.18 | 86.45 | 86.81 | |
| PDCNER(ours) | 87.76 | 86.97 | 87.36 |
| Model | 1% | 10% | 30% | ||||||
| P | R | F1 | P | R | F1 | P | R | F1 | |
| BERT-BiLSTM-CRF | 31.79 | 1.80 | 3.41 | 67.92 | 75.84 | 71.66 | 74.59 | 81.44 | 77.87 |
| LEBERT | 17.39 | 11.43 | 13.79 | 74.81 | 83.47 | 78.91 | 74.05 | 80.14 | 76.97 |
| PDCNER(ours) | 18.18 | 11.43 | 14.04 | 85.00 | 86.44 | 85.71 | 87.22 | 86.17 | 86.69 |
| Model | 50% | 100% | |||||||
| P | R | F1 | P | R | F1 | ||||
| BERT-BiLSTM-CRF | 79.18 | 83.45 | 81.26 | 80.29 | 84.73 | 82.45 | |||
| LEBERT | 81.09 | 83.4 | 82.23 | 87.18 | 86.45 | 86.81 | |||
| PDCNER(ours) | 87.68 | 86.71 | 87.19 | 87.76 | 86.97 | 87.36 | |||
| Model | Ontonotes | Resume | |||||||
|---|---|---|---|---|---|---|---|---|---|
| P | R | F1 | P | R | F1 | P | R | F1 | |
| BERT-BiLSTM-CRF | 71.29 | 67.1 | 69.13 | 84.11 | 80.2 | 82.11 | 96.56 | 97.23 | 95.89 |
| LEBERT | 78.2 | 74.64 | 76.38 | 89.76 | 82.68 | 86.07 | 95.89 | 97.42 | 96.65 |
| PDCNER(ours) | 81.42 | 76.56 | 78.91 | 89.68 | 83.43 | 86.44 | 96.07 | 97.36 | 96.71 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





