Submitted:
07 November 2024
Posted:
08 November 2024
You are already at the latest version
Abstract
Keywords:
Method
The Start of SARS-CoV-2: Data Descriptions, Results, and Interpretations
The Data
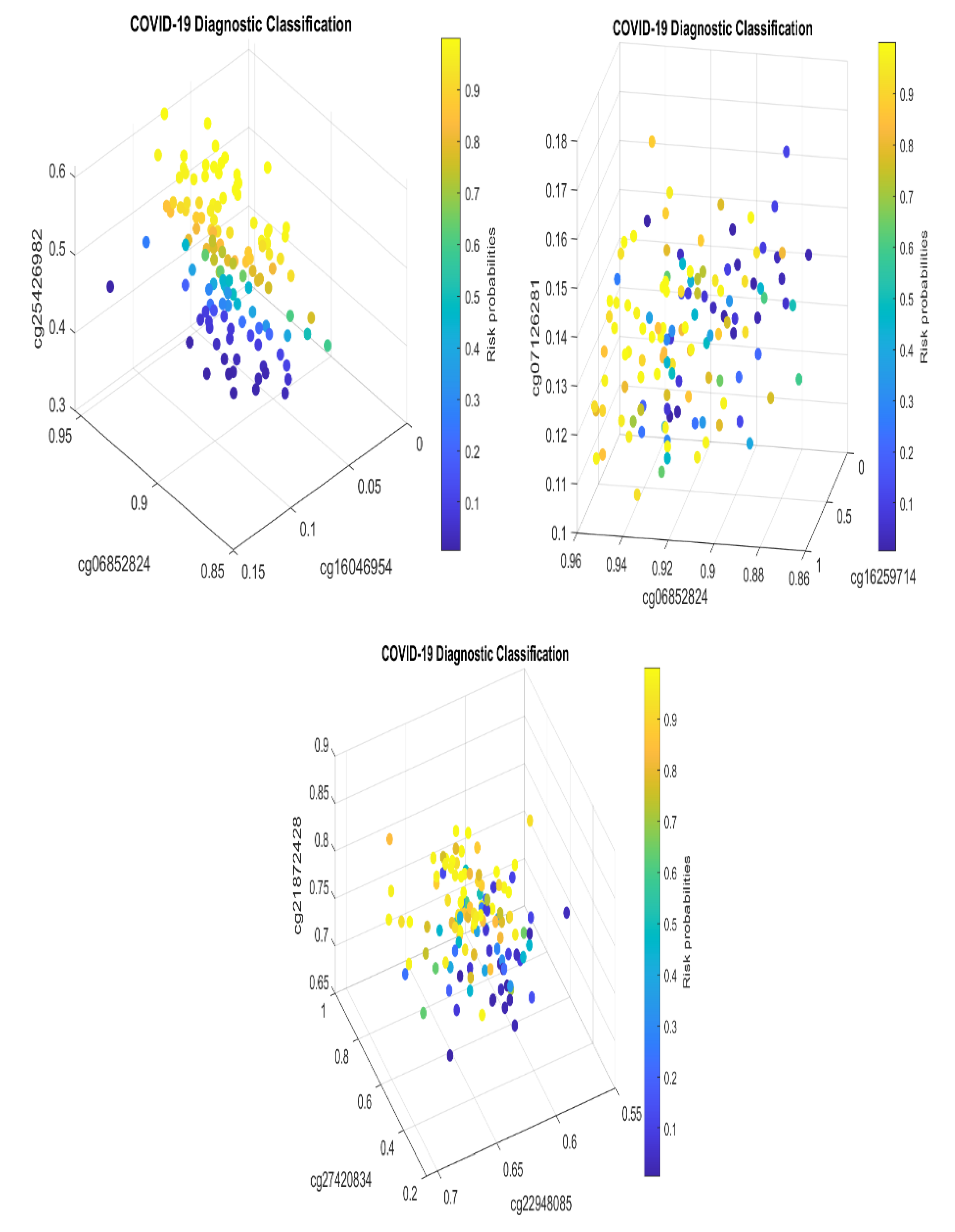
The Analysis
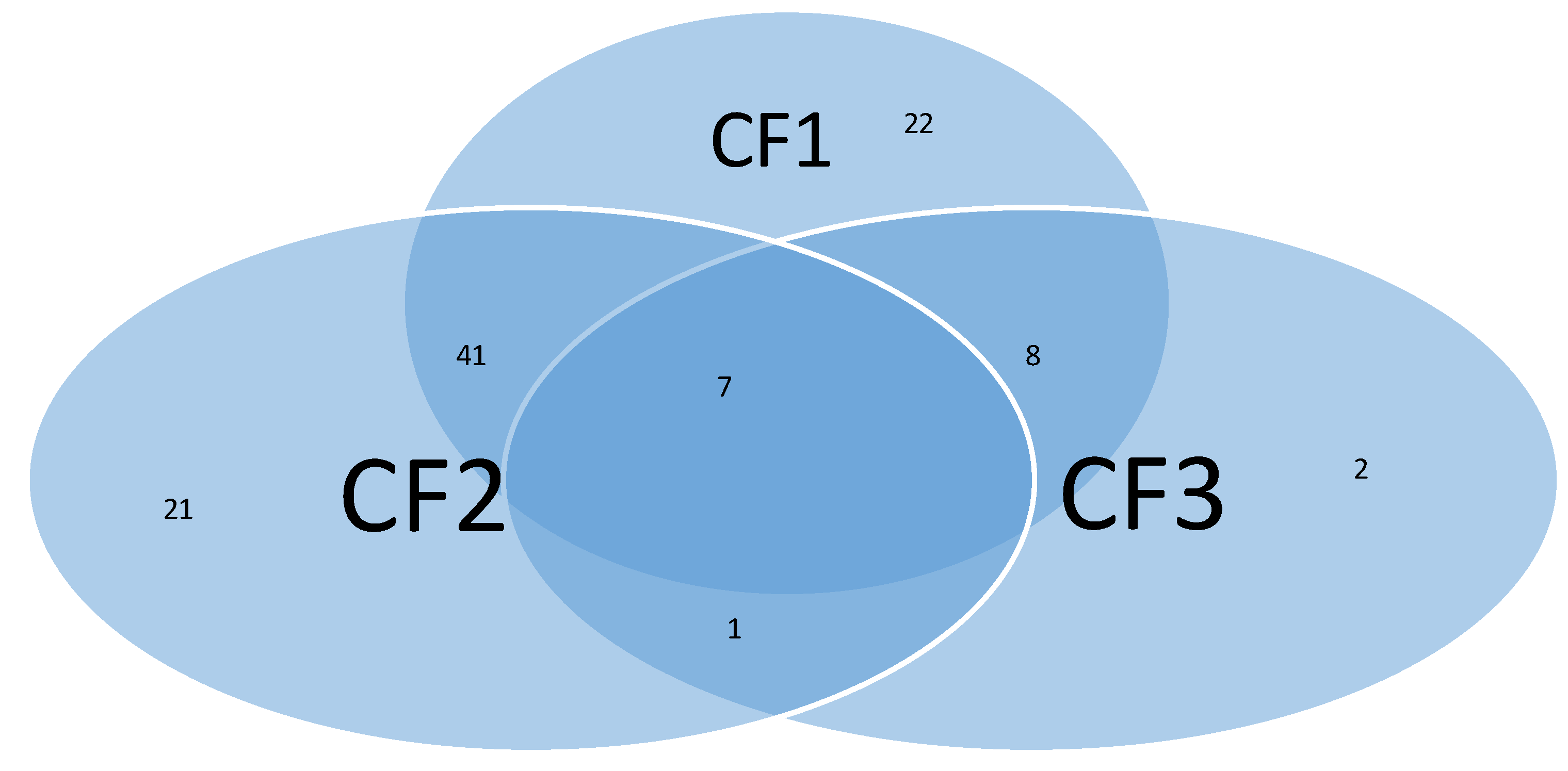
| CF1: | -27.7252 -92.8578*MFSD11 +20.3366*CAB39 +35.1956*SERPINB8 |
| CF2: | -78.6488 +55.57*SDK1 +42.3906*CAB39 -60.8459*SAMM50 |
| CF3: | -20.8835 -101.445* KCNAB1 -24.2856* ZNF280D +116.8975* RANP1 |
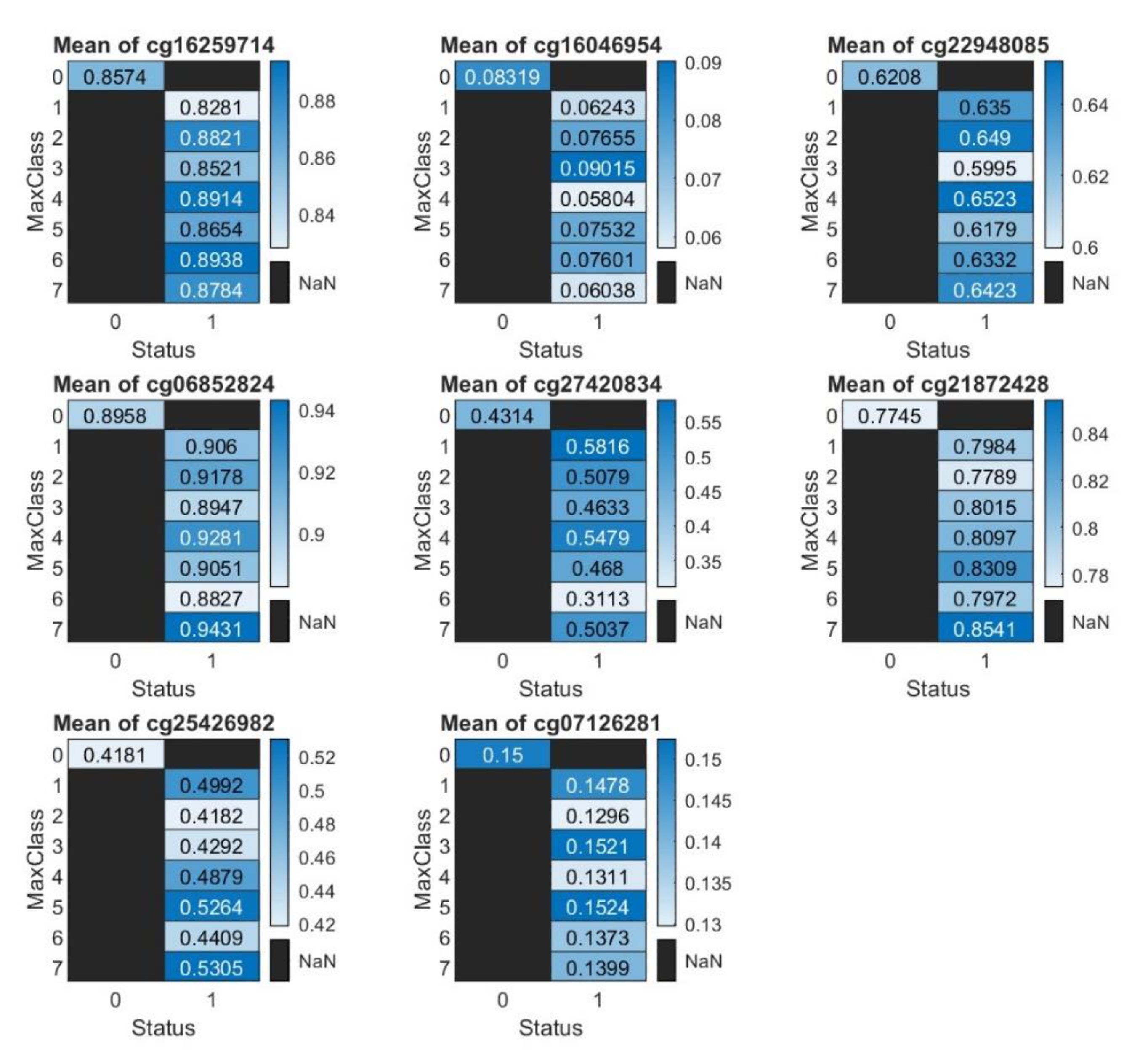
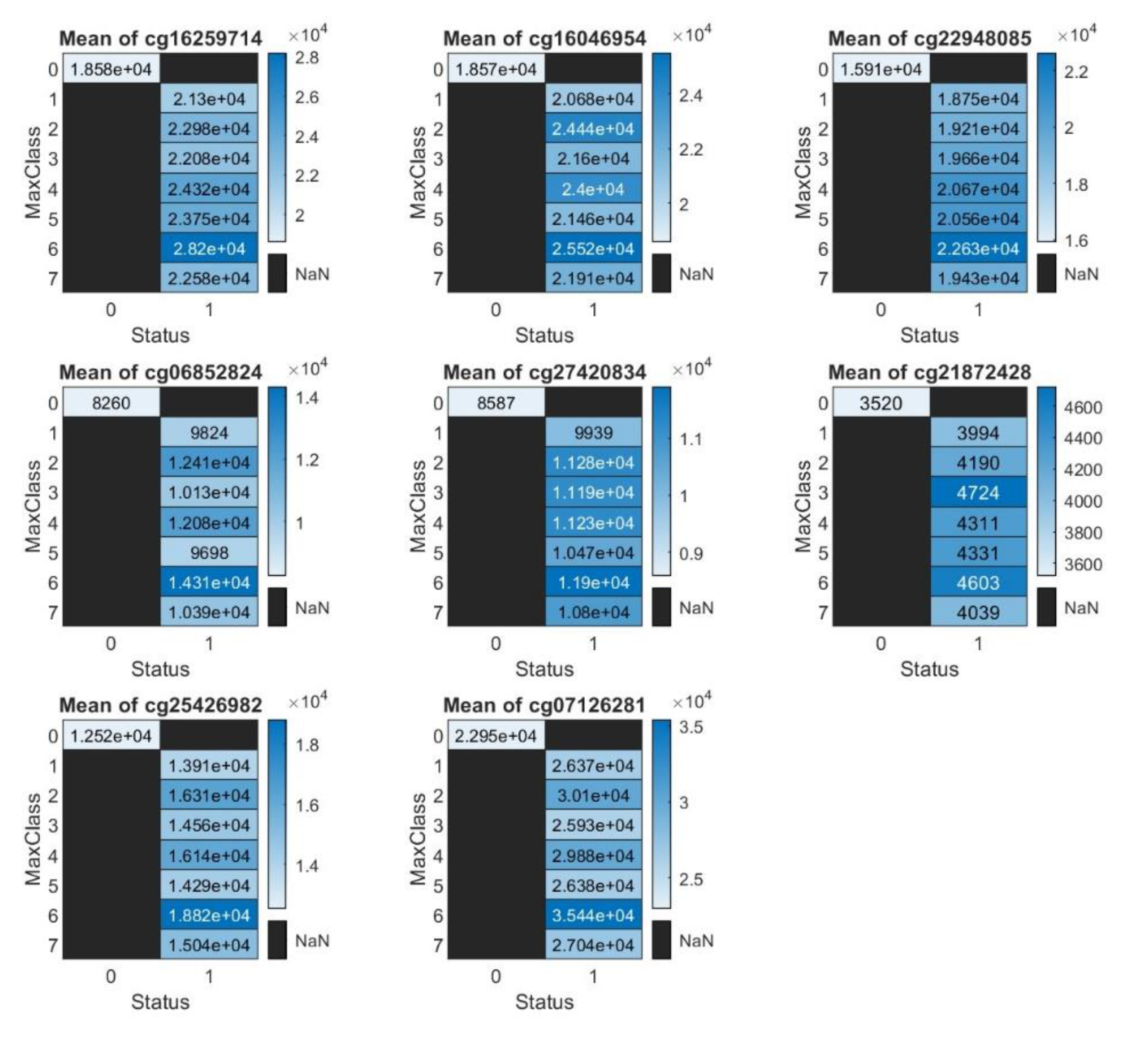
Heatmap Illustration
Cohort-to-Cohort Cross-Validation
Discussions
Conclusions
Data Availability
Competing Interests
Statement of ethics
Limitation statements
Author Contributions
References
- Callaway, E. The quest to find genes that drive severe covid. Nature 2021, 595, 346–348. [CrossRef]
- COVID-19 Host Genetics Initiative. Mapping the human genetic architecture of COVID-19. Nature, pages 474-477, 2021. [CrossRef]
- Davalos V, García-Prieto CA, Ferrer G, Aguilera-Albesa S et al. Epigenetic profiling linked to multisystem inflammatory syndrome in children (MIS-C): A multicenter, retrospective study. EClinicalMedicine 2022 Aug;50:101515. PMID: 35770252.
- Dite GS, Murphy NM, and Allman R. Development and validation of a clinical and genetic model for predicting risk of severe COVID-19. Epidemiology and Infection, 149:e162, 2021. [CrossRef]
- Konigsberg IR, Barnes B, Campbell M, Davidson E, Zhen Y, Pallisard O, Boorgula MP, Cox C, Nandy D, Barnes KC, et al. Host methylation predicts SARS-CoV-2 infection and clinical outcome. Commun Med (Lond). 2021;1(1):42. [CrossRef]
- Melms, J.C., Biermann, J., Huang, H. et al. A molecular single-cell lung atlas of lethal COVID-19. Nature 595, 114–119 (2021). [CrossRef]
- Pairo-Castineira E, Clohisey S, Klaric L, et al. Genetic mechanisms of critical illness in COVID-19. Nature, 591:92-98, 2021. [CrossRef]
- Zhang Z. Five critical genes related to seven COVID-19 subtypes: A data science discovery. Journal of Data Science, 19(1):142-150, 2021. [CrossRef]
- Zhang Z. The existence of at least three genomic signature patterns and at least seven subtypes of COVID-19 and the end of the disease. Vaccines, 10, 761, 2022. [CrossRef]
- Zhang Z. Genomic Biomarker Heterogeneities Between SARS-CoV-2 and COVID-19. Vaccines 2022, 10(10), 1657; [CrossRef]
- Zhang Z, Genomic Transcriptome Benefits and Potential Harms of COVID-19 Vaccines Indicated from Optimized Genomic Biomarkers. Vaccines 2022, 10(11), 1774. [CrossRef]
- Zhang, Z. Omicron’s Intrinsic Gene-Gene Interactions Jumped Away from Earlier SARS-CoV-2 Variants and Gene Homologs between Humans and Animals. Advances in Biomarker Sciences and Technology 2023. [CrossRef]
- Zhang, Z. The Initial COVID-19 Reliable Interactive DNA Methylation Markers and Biological Implications. Biology. 2024; 13(4):245. [CrossRef]
- Zhang Y, Guo X, Li C, Kou Z et al. Transcriptome Analysis of Peripheral Blood Mononuclear Cells in SARS-CoV-2 Naïve and Recovered Individuals Vaccinated with Inactivated Vaccine. Front Cell Infect Microbiol 2021;11:821828.
- Zhang Z. Lift the veil of breast cancers using 4 or fewer critical genes. Cancer Informatics, 21:1-11, 2022. [CrossRef]
- Phillips, T. (2008) The role of methylation in gene expression. Nature Education 1(1):116.
- Zhang Z. Functional effects of four or fewer critical genes linked to lung cancers and new sub-types detected by a new machine learning classifier. Journal of Clinical Trials, 11:S14:100001, 2021. https://www.longdom.org/open-access/functional-effects-of-four-or-fewer-critical-genes-linked-to-lung-cancers-and-new-subtypes-detected-by-a-new-machine-learning-clas-88321.html.
- Liu, Y., Xu, Y., Li, X., Chen, M., Wang, X., Zhang, N., Zhang, H., Zhang, Z. (2023), Towards Precision Oncology Discovery: Four Less Known Genes and Their Unknown Interactions as Highest-Performed Biomarkers for Colorectal Cancer. npj Precision Oncology. https://www.nature.com/articles/s41698-024-00512-1.
- Liu Y, Zhang H, Xu Y, Liu, Y-Z, Al-Adra DP, Yeh MM, et al. (2023), Five Critical Gene-based Biomarkers with Optimal Performance for Hepatocellular Carcinoma, Cancer Informatics, 22. [CrossRef]
- Zhang, Z. Quotient correlation: A sample based alternative to Pearson’s correlation. Ann. Stat. 2008, 36, 1007–1030 . [CrossRef]
- Carapito R, Li R, Helms J, Carapito C et al. Identification of driver genes for critical forms of COVID-19 in a deeply phenotyped young patient cohort. Sci Transl Med 2022 Jan 19;14(628):eabj7521.
- Balnis J, Madrid A, Hogan KJ, Drake LA et al. Blood DNA methylation and COVID-19 outcomes. Clin Epigenetics 2021 May 25;13(1):118. PMID: 34034806.
- Du, P., Zhang, X., Huang, CC. et al. Comparison of Beta-value and M-value methods for quantifying methylation levels by microarray analysis. BMC Bioinformatics 11, 587 (2010). [CrossRef]
- Van Zandt KE, Greer MT, Gelhaus HC. Glanders: an overview of infection in humans. Orphanet J Rare Dis. 2013 Sep 3;8:131. [CrossRef]
- Wright G. What is quantum interference? https://www.techtarget.com/whatis/definition/quantum-interference.
- Barreto EA, Cruz AS, Veras FP, Martins R, Bernardelli RS, Paiva IM, et al. COVID-19-related hyperglycemia is associated with infection of hepatocytes and stimulation of gluconeogenesis. Proc Natl Acad Sci U S A. 2023 May 23;120(21):e2217119120 . [CrossRef]
- Guarnieri JW, Dybas JM, Fazelinia H, Kim MS, Frere J, Zhang Y, Soto Albrecht Y, Murdock DG, Angelin A, Singh LN, Weiss SL, Best SM, Lott MT, Zhang S, Cope H, Zaksas V, Saravia-Butler A, Meydan C, Foox J, Mozsary C, Bram Y, Kidane Y, Priebe W, Emmett MR, Meller R, Demharter S, Stentoft-Hansen V, Salvatore M, Galeano D, Enguita FJ, Grabham P, Trovao NS, Singh U, Haltom J, Heise MT, Moorman NJ, Baxter VK, Madden EA, Taft-Benz SA, Anderson EJ, Sanders WA, Dickmander RJ, Baylin SB, Wurtele ES, Moraes-Vieira PM, Taylor D, Mason CE, Schisler JC, Schwartz RE, Beheshti A, Wallace DC. Core mitochondrial genes are down-regulated during SARS-CoV-2 infection of rodent and human hosts. Sci Transl Med. 2023 Aug 9;15(708):eabq1533. [CrossRef]
- Parreno V, Loubiere V, Schuettengruber B, Fritsch L, Rawal CC, Erokhin M, Győrffy B, Normanno D, Di Stefano M, Moreaux J, Butova NL, Chiolo I, Chetverina D, Martinez AM, Cavalli G. Transient loss of Polycomb components induces an epigenetic cancer fate. Nature. 2024 Apr 24. [CrossRef]
- Cavaillon JM, Singer M, Skirecki T. Sepsis therapies: learning from 30 years of failure of translational research to propose new leads. EMBO Mol Med. 2020 Apr 7;12(4):e10128. [CrossRef]
- Liu, W.J., Liu, P., Lei, W. et al. Surveillance of SARS-CoV-2 at the Huanan Seafood Market. Nature 631, 402–408 (2024). [CrossRef]
- Crits-Christoph A, Levy JI, Pekar JE, Goldstein SA, Singh R, Hensel Z, Gangavarapu K, Rogers MB, Moshiri N, Garry RF, Holmes EC, Koopmans MPG, Lemey P, Popescu S, Rambaut A, Robertson DL, Suchard MA, Wertheim JO, Rasmussen AL, Andersen KG, Worobey M, Débarre F. Genetic tracing of market wildlife and viruses at the epicenter of the COVID-19 pandemic. Cell. 2024 Sep 19;187(19):5468-5482.e11. [CrossRef]
| sites | gene | CF1 | CF2 | CF3 | CFmax |
| Intercept | -27.7252 | -78.6488 | -20.8835 | ||
| cg16259714 | SDK1 | 55.57 | |||
| cg16046954 | MFSD11 | -92.8578 | |||
| cg06852824 | CAB39 | 20.3366 | 42.3906 | ||
| cg25426982 | SERPINB8 | 35.1956 | |||
| cg07126281 | SAMM50 | -60.8459 | |||
| cg22948085 | KCNAB1 | -101.445 | |||
| cg27420834 | ZNF280D | -24.2856 | |||
| cg21872428 | RANP1 | 116.8975 | |||
| Accuracy | % | 81.25 | 75 | 34.38 | 100 |
| Sensitivity | % | 76.47 | 68.63 | 17.65 | 100 |
| Specificity | % | 100 | 100 | 100 | 100 |
| sites | gene | CF1 | CF2 | CFmax |
| Intercept | 3.1477 | 8.8827 | ||
| cg16046954 | MFSD11 | 73.5632 | ||
| cg25426982 | SERPINB8 | -19.5833 | ||
| cg07126281 | SAMM50 | -115.236 | -110.706 | |
| cg27420834 | ZNF280D | 7.9757 | 18.9041 | |
| Accuracy | % | 88.19 | 85.83 | 87.40 |
| Sensitivity | % | 73.33 | 33.33 | 86.67 |
| Specificity | % | 90.18 | 92.86 | 87.50 |
| sites | gene | CF1 | CF2 | CFmax |
| Intercept | 35.2131 | 15.351 | ||
| cg12654612 | ZNF71 | 53.162 | ||
| cg19770550 | MMS19 | 218.9845 | ||
| cg03841686 | MRPS35 | 101.8423 | ||
| cg25042073 | SLC10A7 | -54.9763 | ||
| cg06852824 | CAB39 | -23.5245 | ||
| cg07126281 | SAMM50 | -73.7777 | ||
| Accuracy | % | 95.28 | 92.13 | 98.43 |
| Sensitivity | % | 73.33 | 33.33 | 100.00 |
| Specificity | % | 98.21 | 100.00 | 98.21 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




