Submitted:
16 April 2026
Posted:
20 April 2026
You are already at the latest version
Abstract
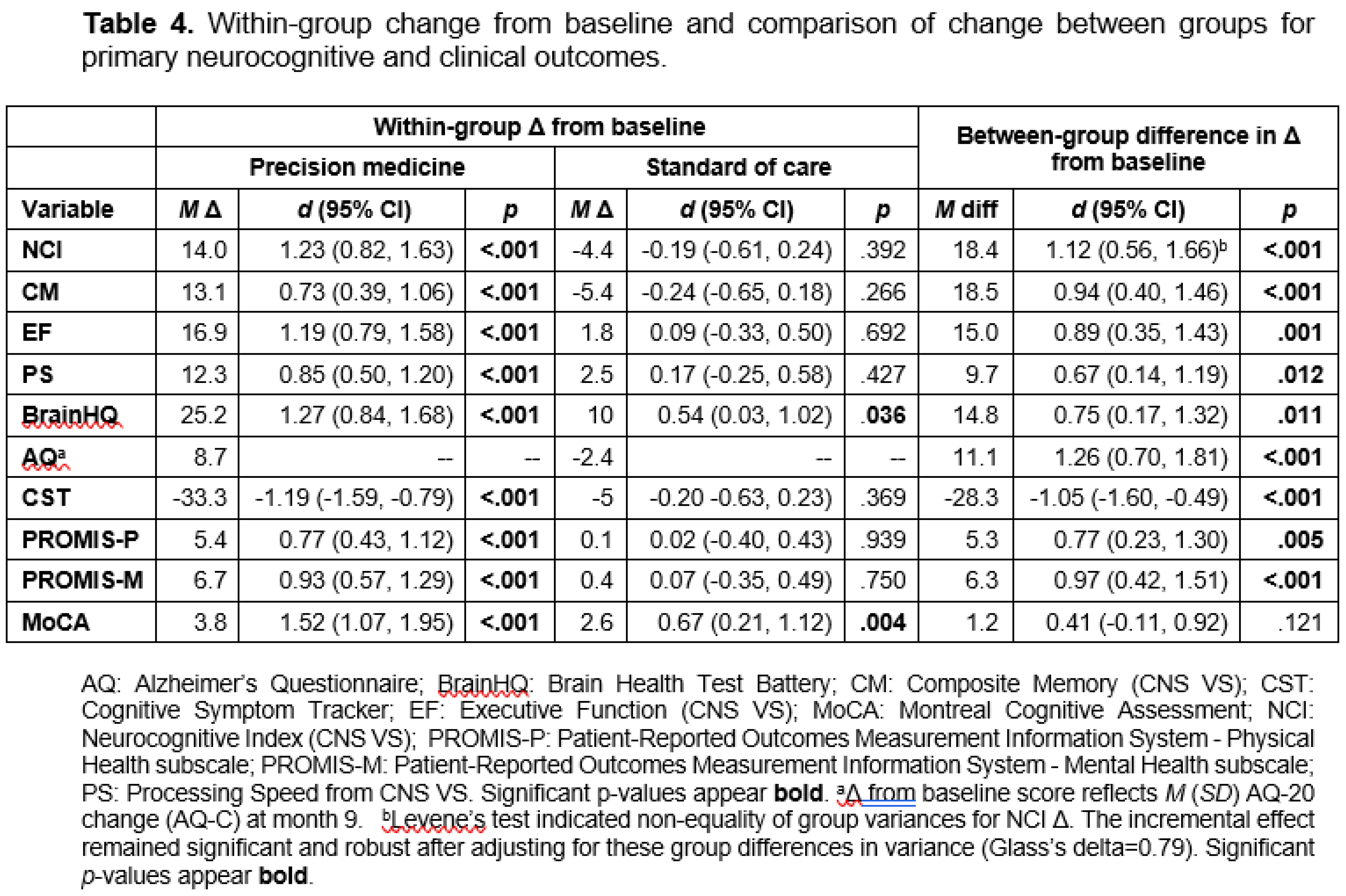
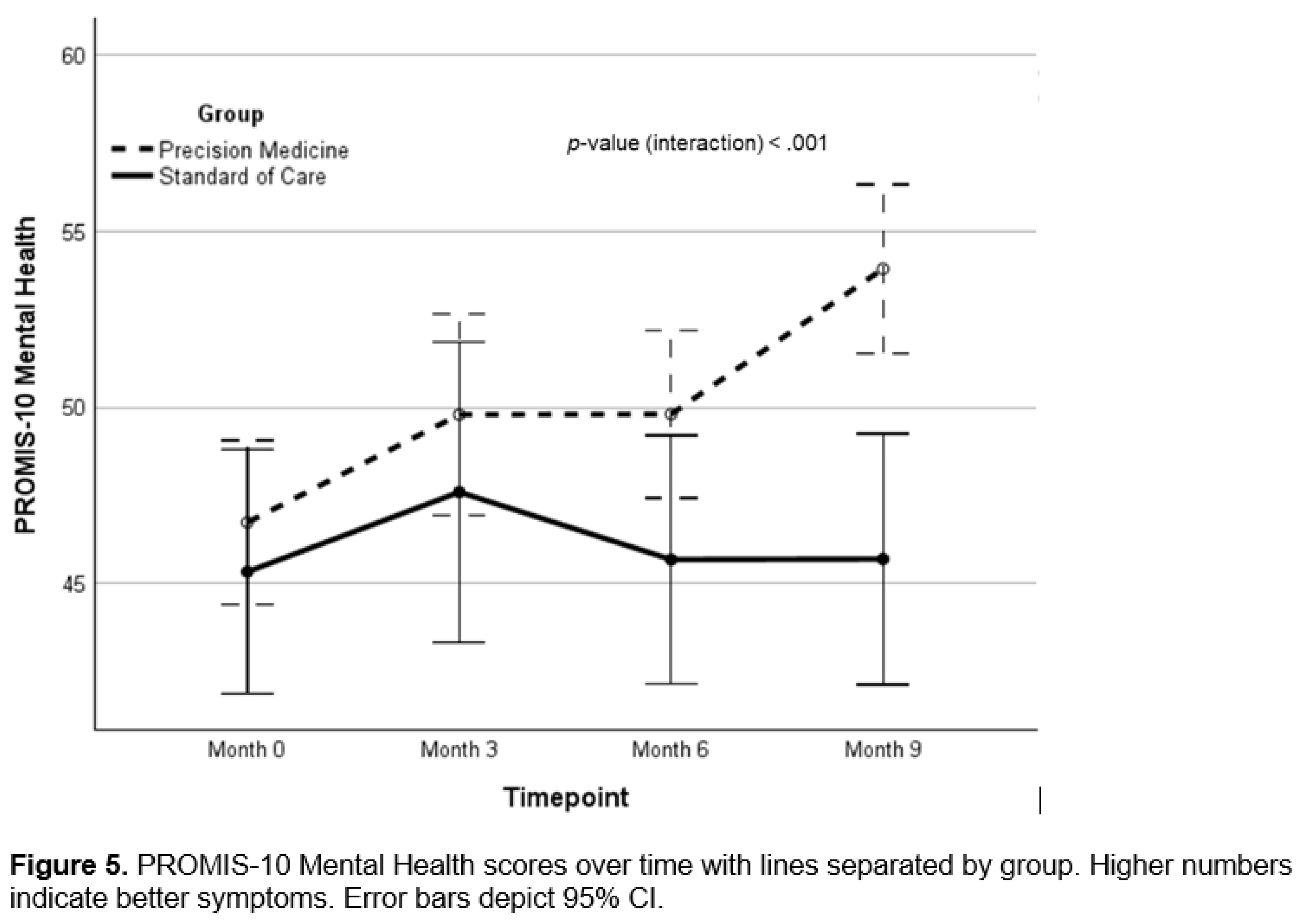
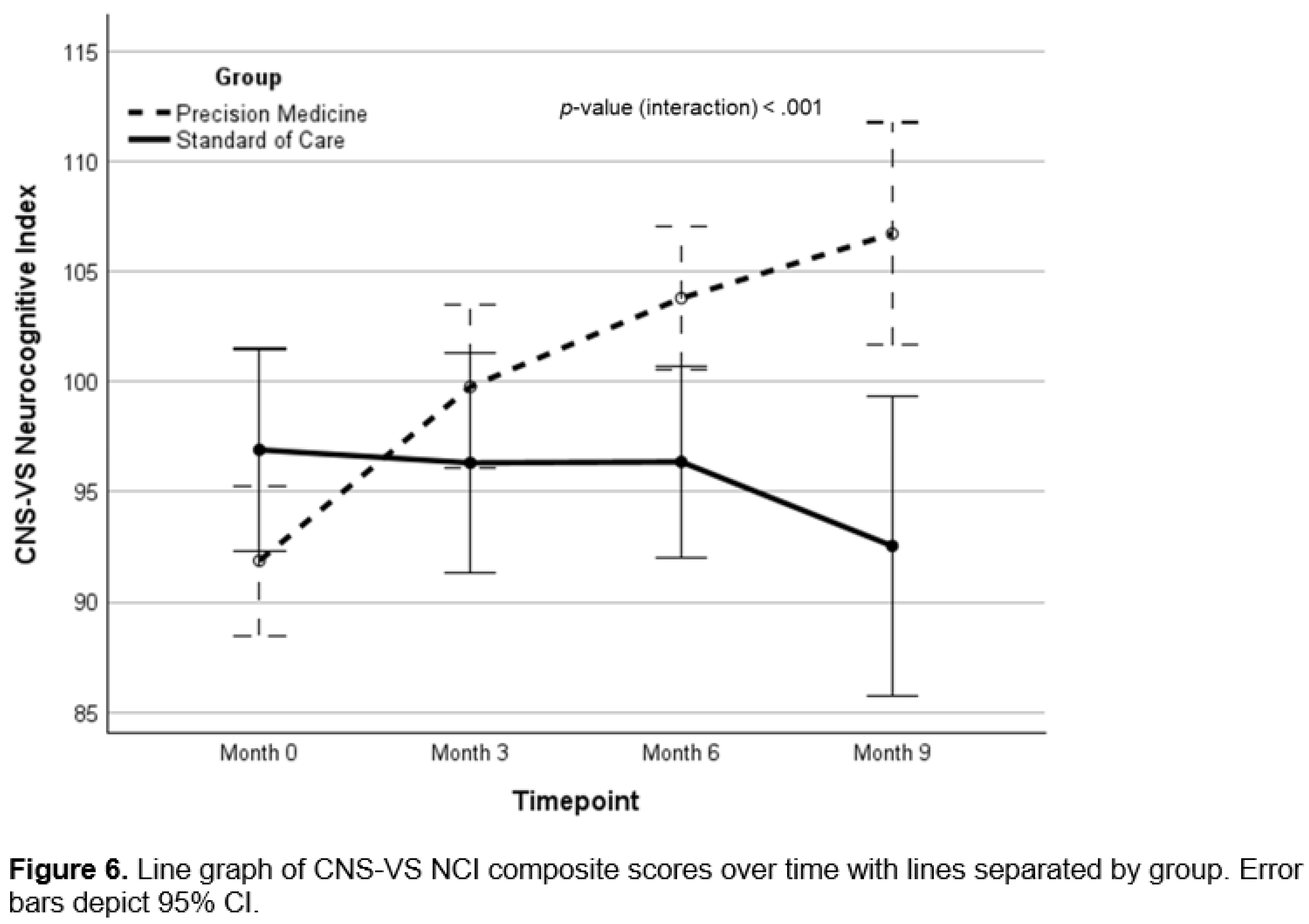
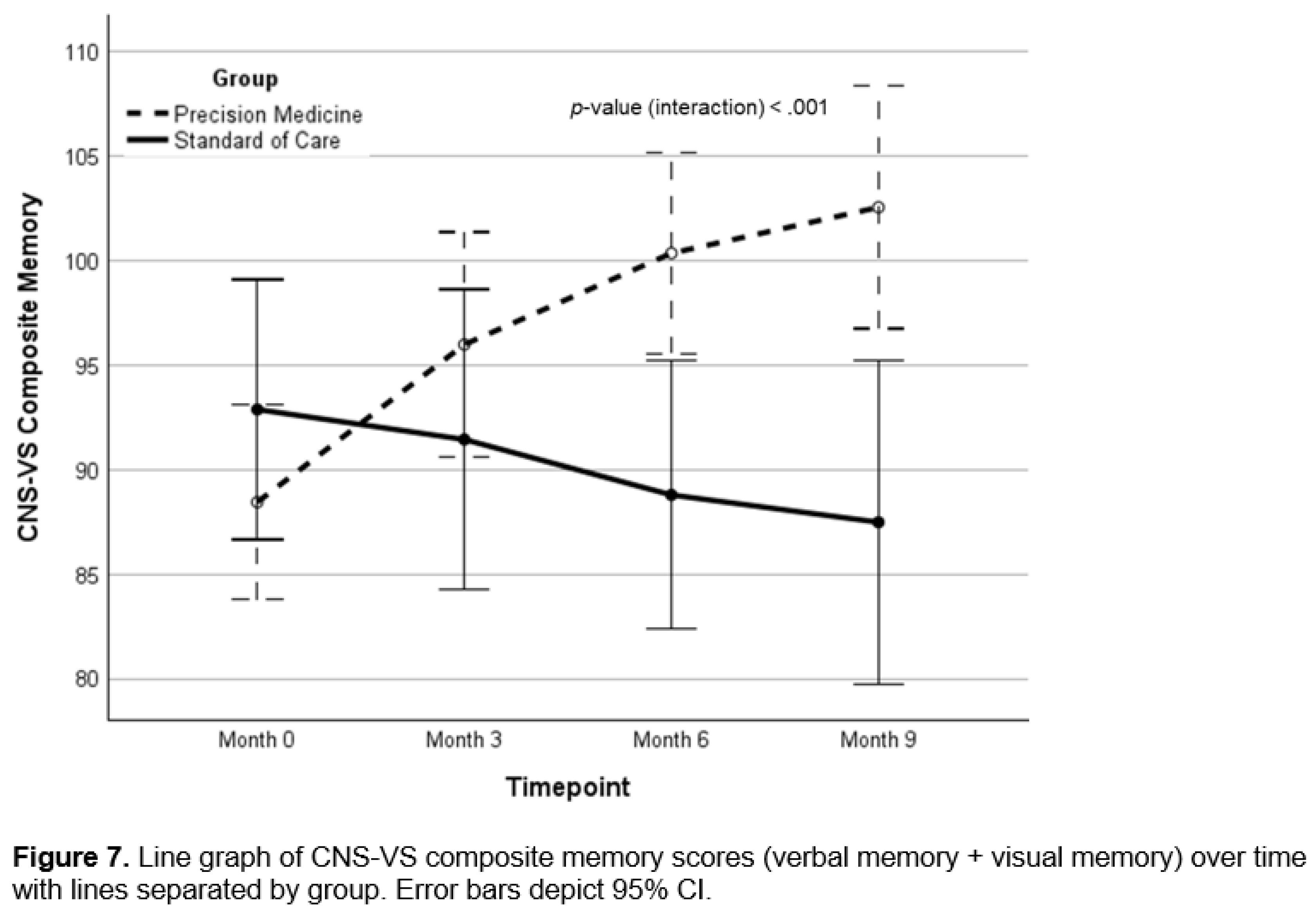
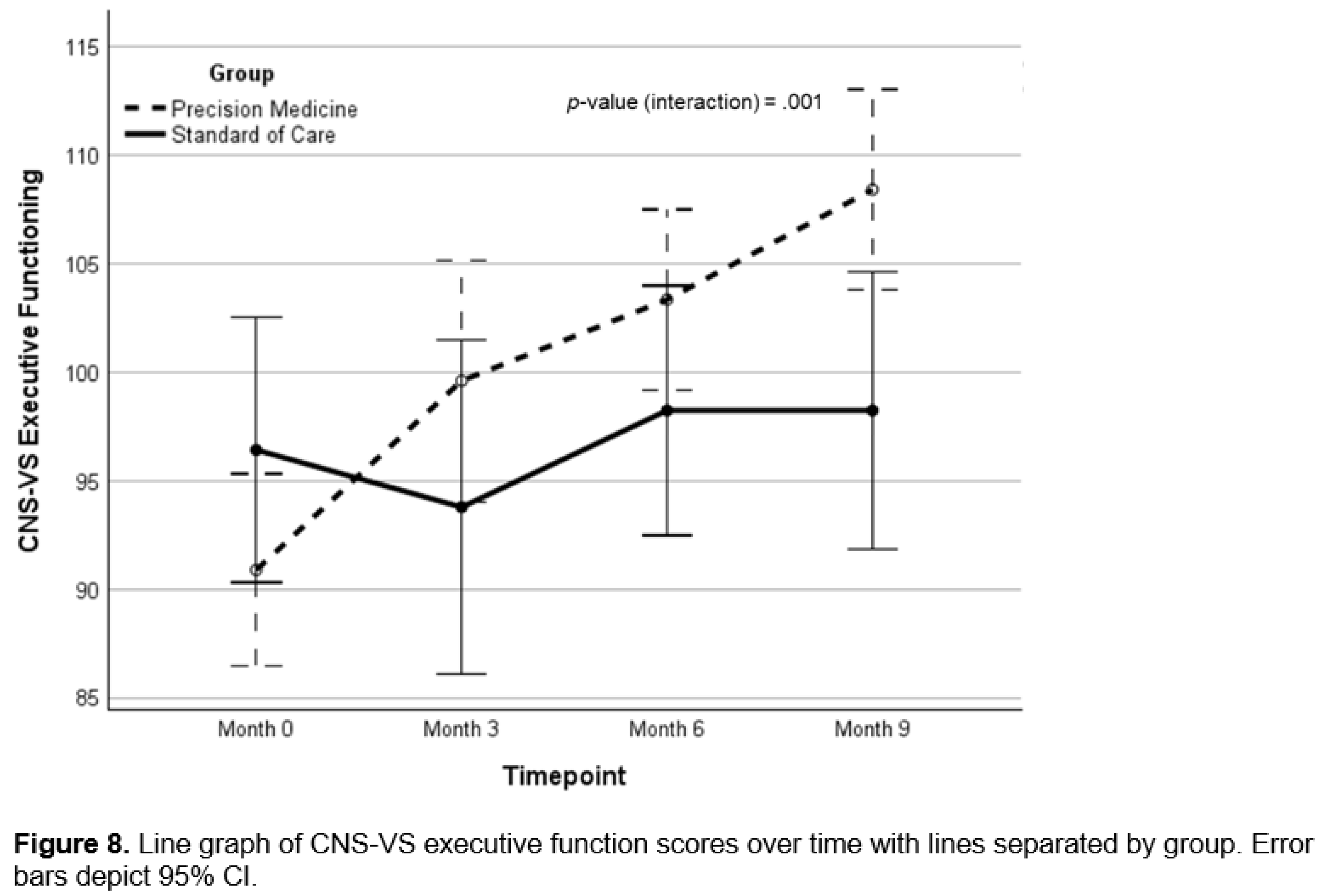
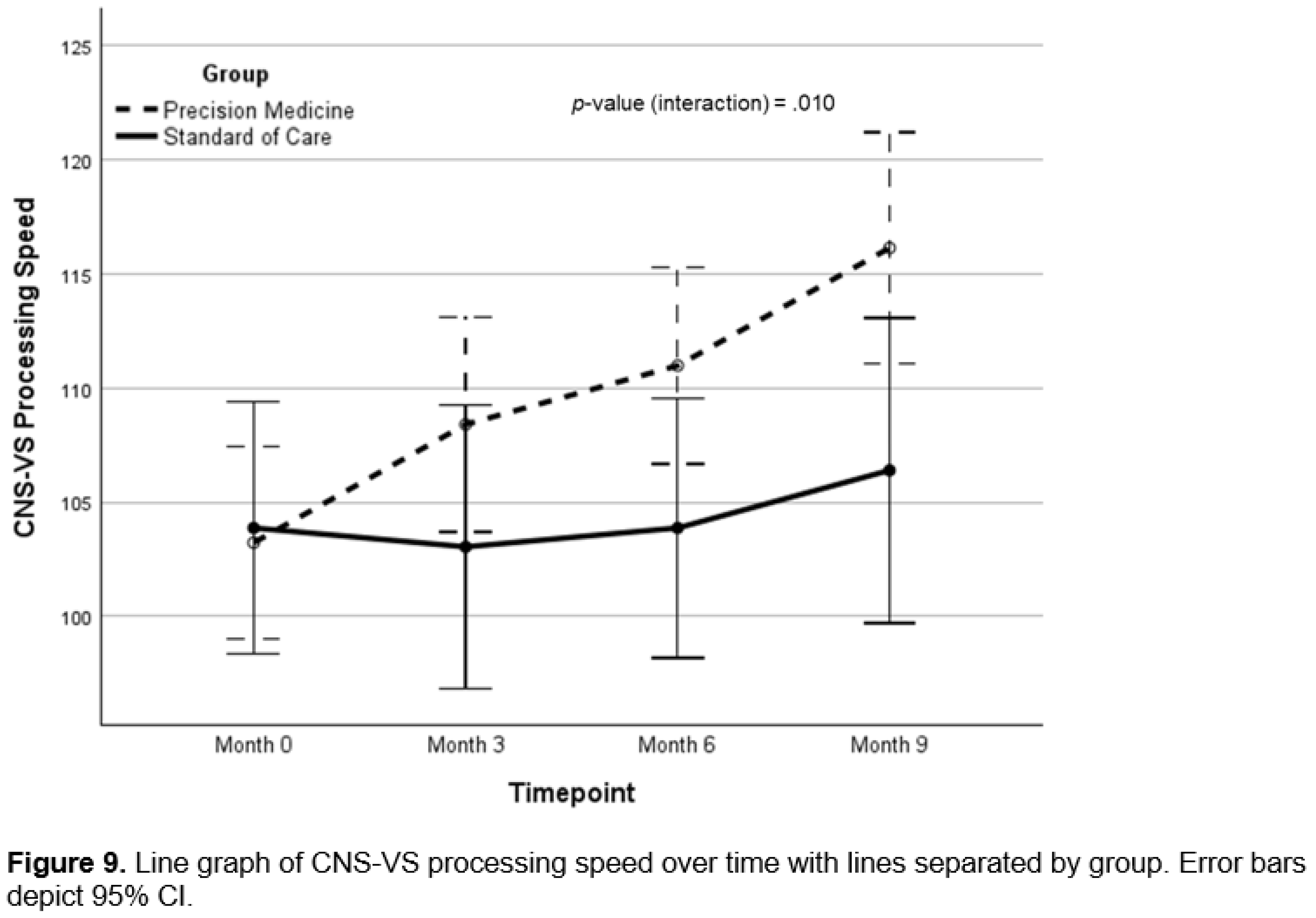
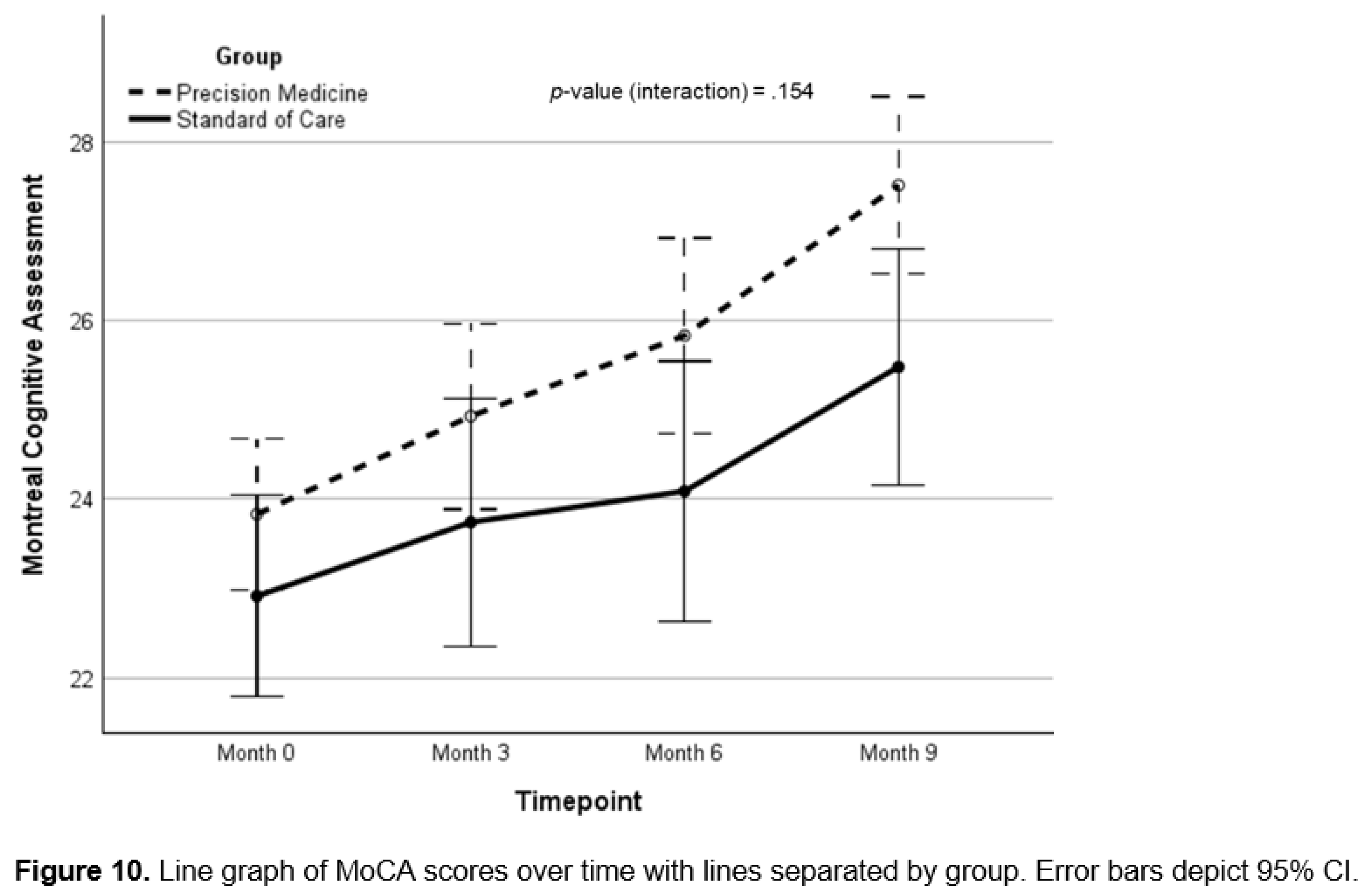
Background: There is a critical need for effective therapeutics for Alzheimer’s. Personalized, precision medicine approaches represent a potentially effective strategy, and proof-of-concept trials have provided supportive data. Objective: To determine whether a precision medicine approach to Alzheimer’s at the mild cognitive impairment or early dementia stage is effective in a randomized controlled clinical trial. Methods: Seventy-three patients with mild cognitive impairment or early dementia were evaluated for biochemical, microbiological, genetic, epigenetic, and imaging parameters associated with cognitive decline, then assigned randomly to a precision medicine approach or standard of care treatment. Results: Statistically significant effects of the precision medicine approach were observed for overall neurocognitive functioning (d=1.12; 95% CI, 0.56-1.66; p<0.001), memory (d=0.94; 95% CI, 0.40-1.46; p<0.001), executive function (d=0.89; 95% CI, 0.35-1.43; p=0.001), processing speed (d=0.67; 95% CI, 0.14-1.19; p=0.012), self-reported cognitive symptom severity (d=-1.05; 95% CI, -1.60, -0.49, p<0.001), and partner-reported cognitive symptom severity (d=1.26; 95% CI, 0.70-1.81; p<0.001), with MoCA scores showing a trend to improvement (p=0.154). Furthermore, overall health was enhanced, with improvements in blood pressure, body mass index, glycemic index, lipid profiles, and methylation status. Treatment effect size on overall cognitive function exceeded previous trials, being 2-3 times larger than effects of lifestyle interventions and 4-7-times larger than those of anti-amyloid therapies. Conclusion: A personalized, precision medicine approach represents an effective treatment for patients with mild cognitive impairment or early-stage dementia. This treatment improves cognition and overall health rather than simply retarding decline, without significant negative side effects such as brain edema, microhemorrhage, or atrophy.
Keywords:
Introduction
Methods
Trial Design
Participants
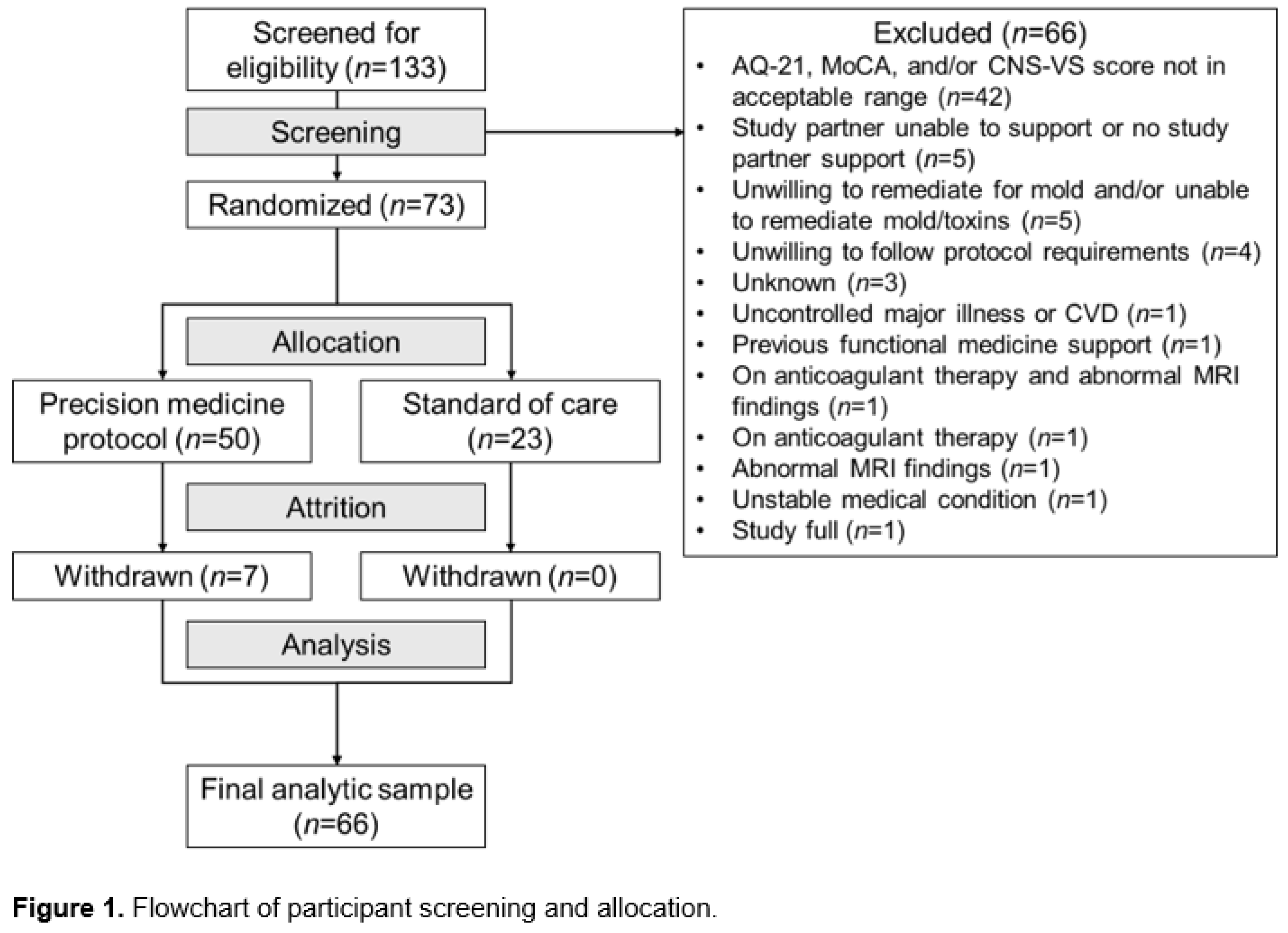
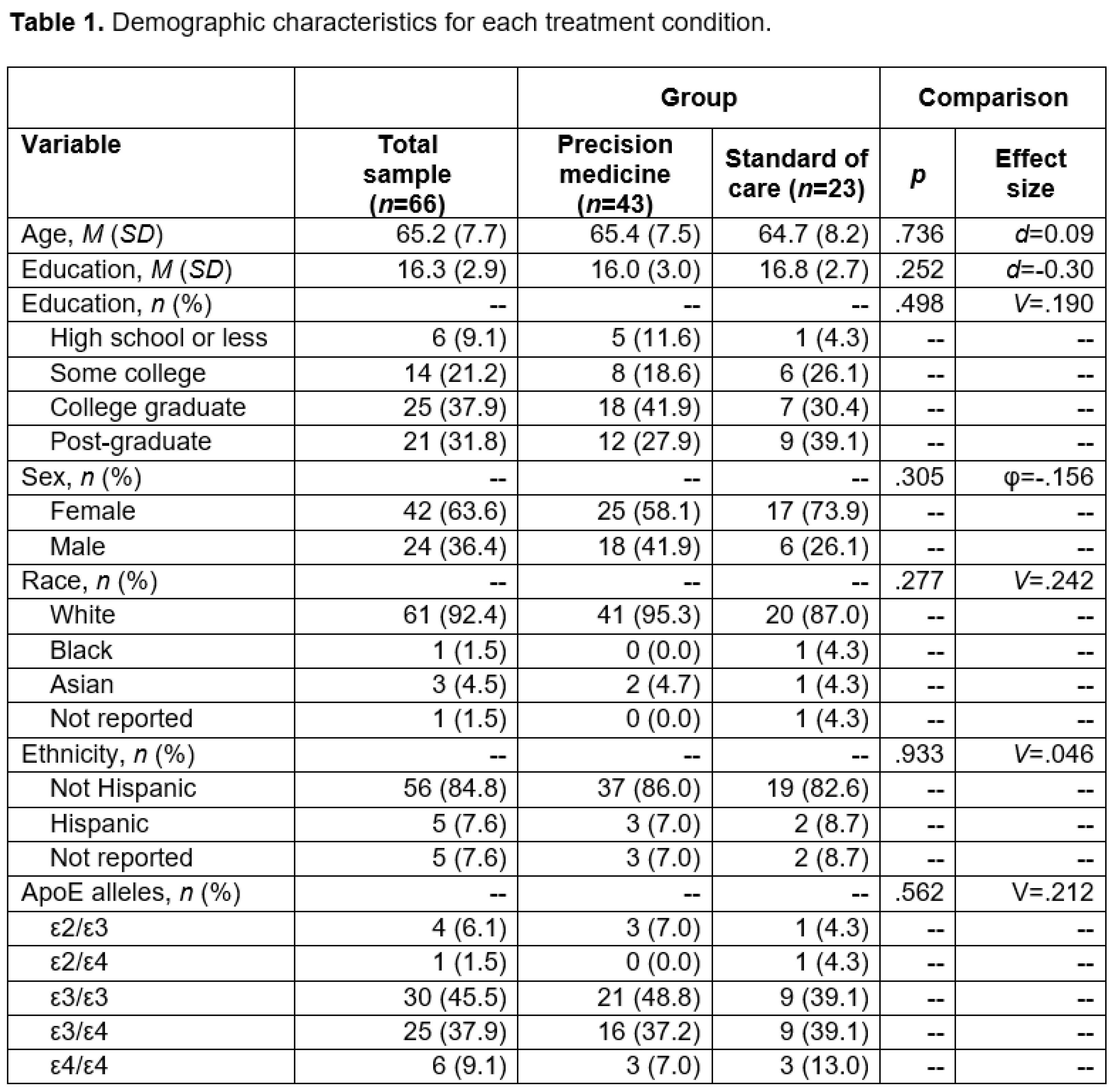
Measures
Treatment Procedures
Statistical Analysis for Neurocognitive, Clinical, and Biomarker Outcomes
Trial Safety
Results
Metabolic Effects
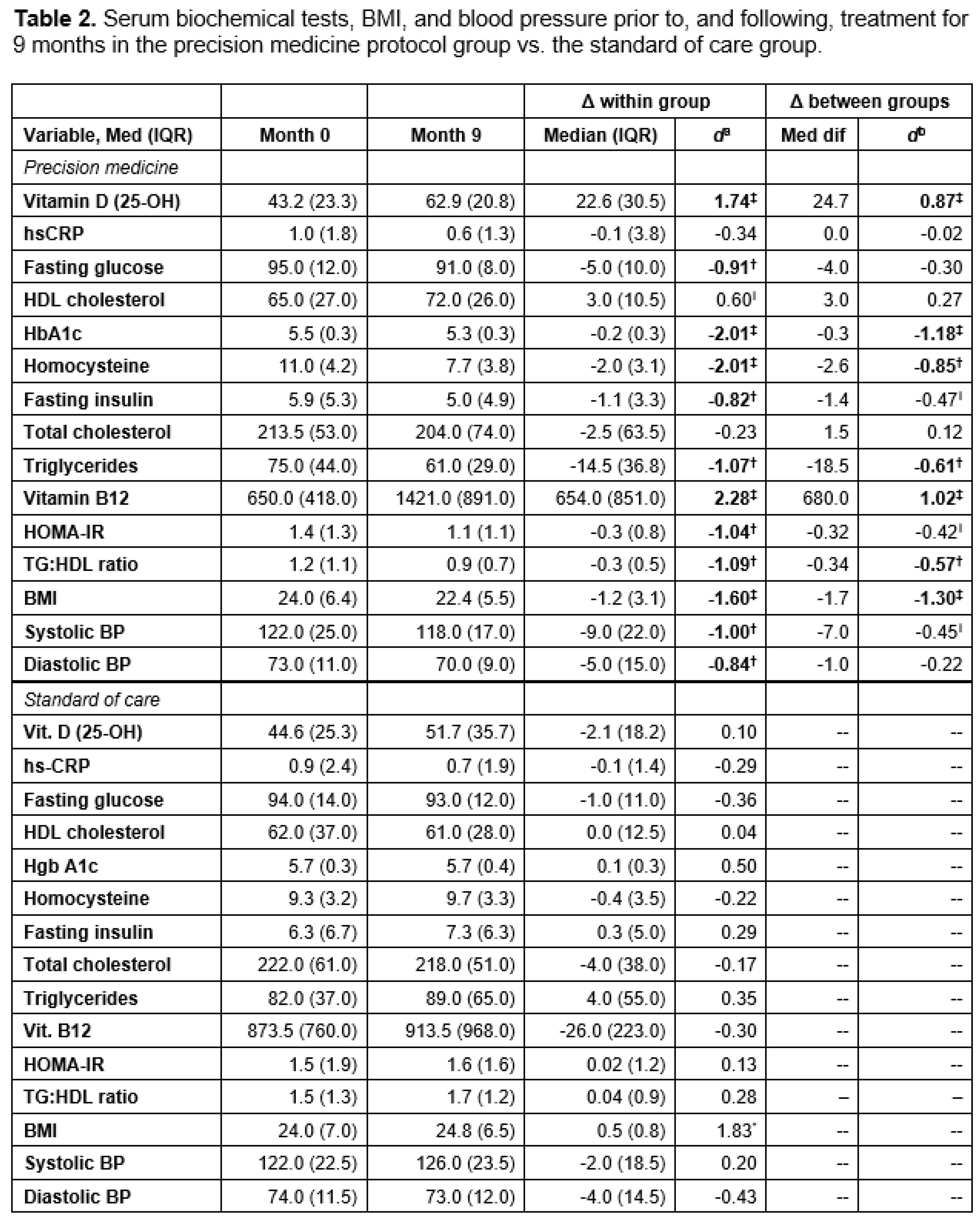
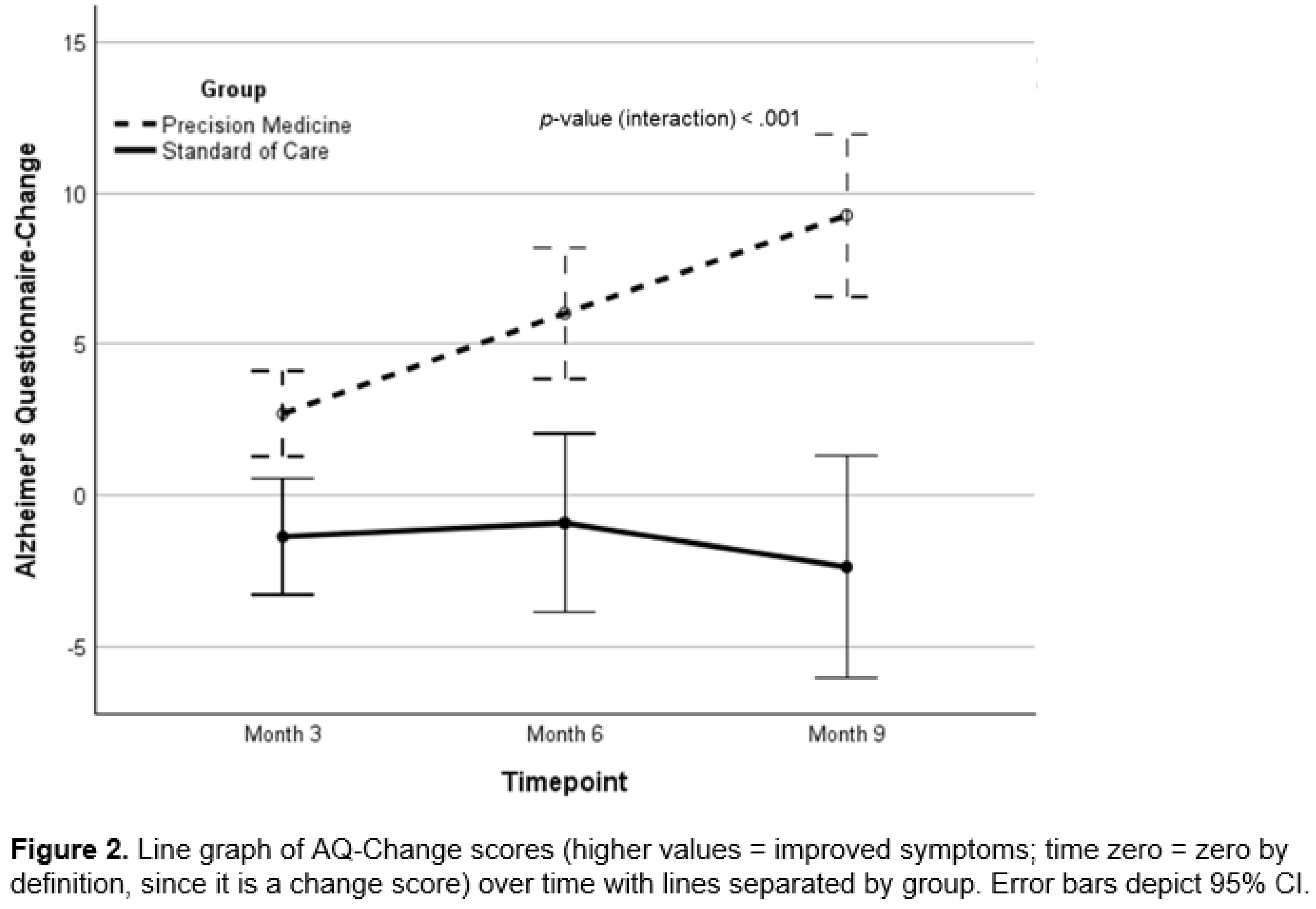
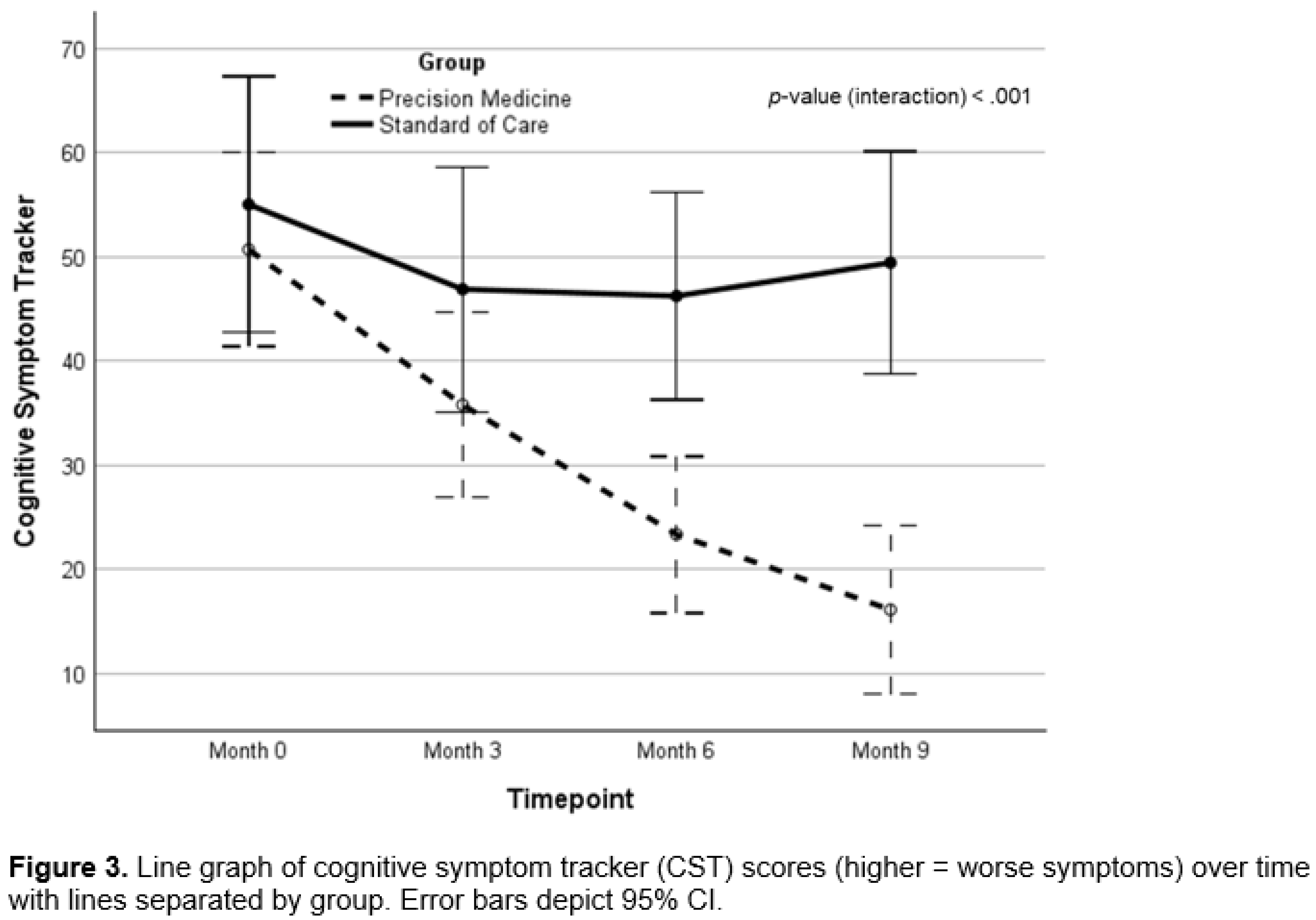
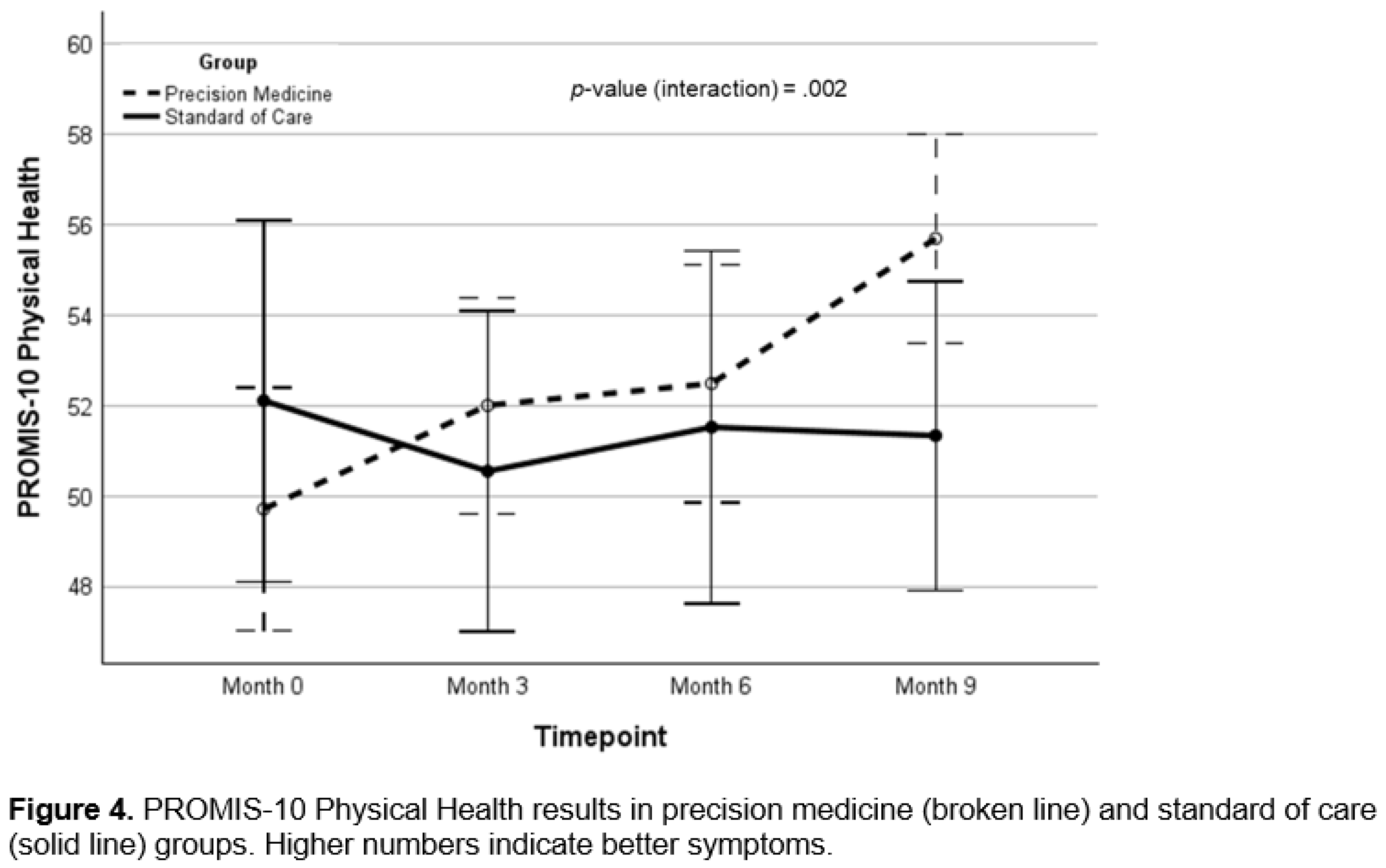
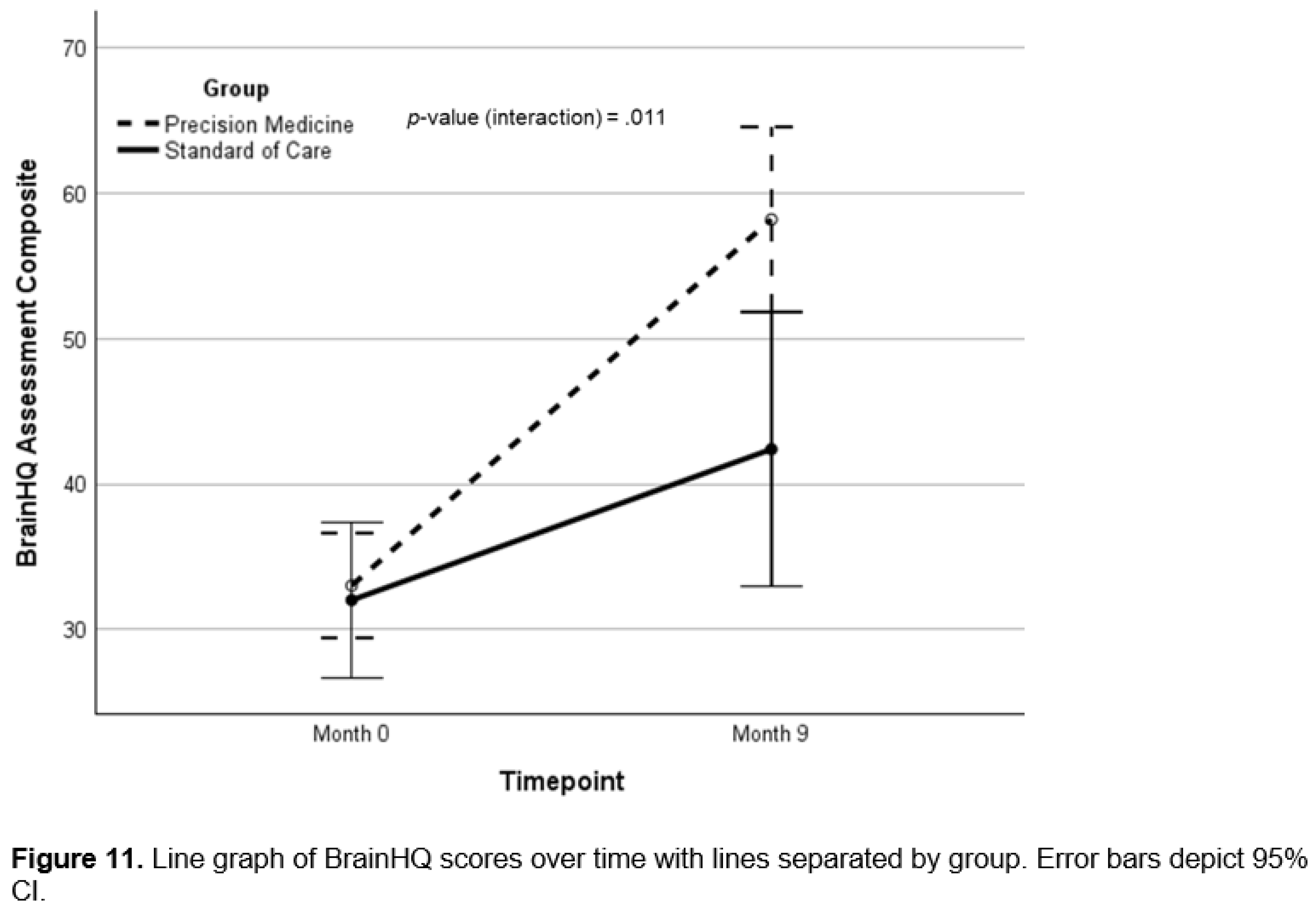
Cognitive Function and Clinical Symptom Outcomes
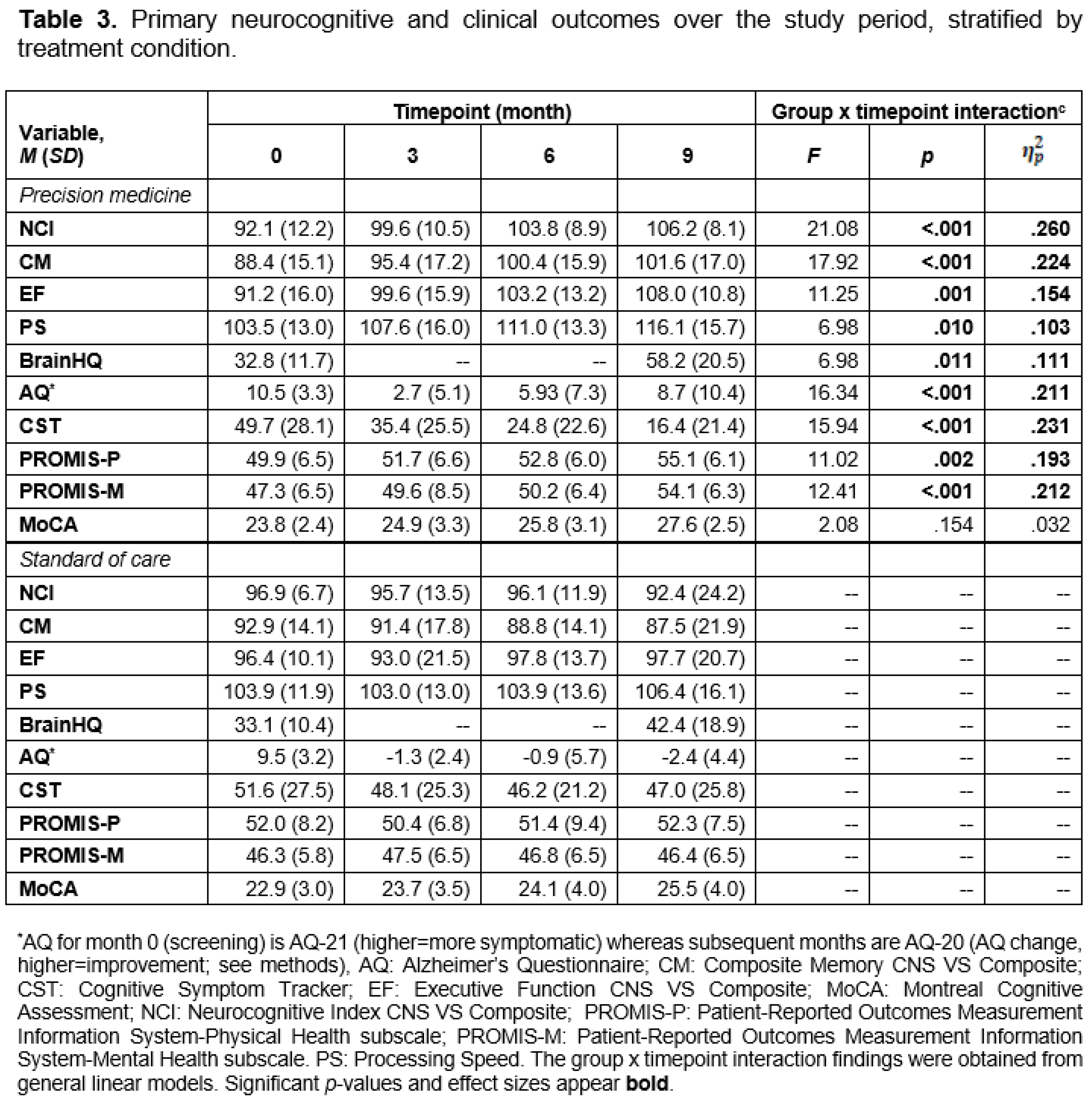
Brain Training
Brain MRI with Volumetric Quantification
Epigenetics
Biomarkers of AD
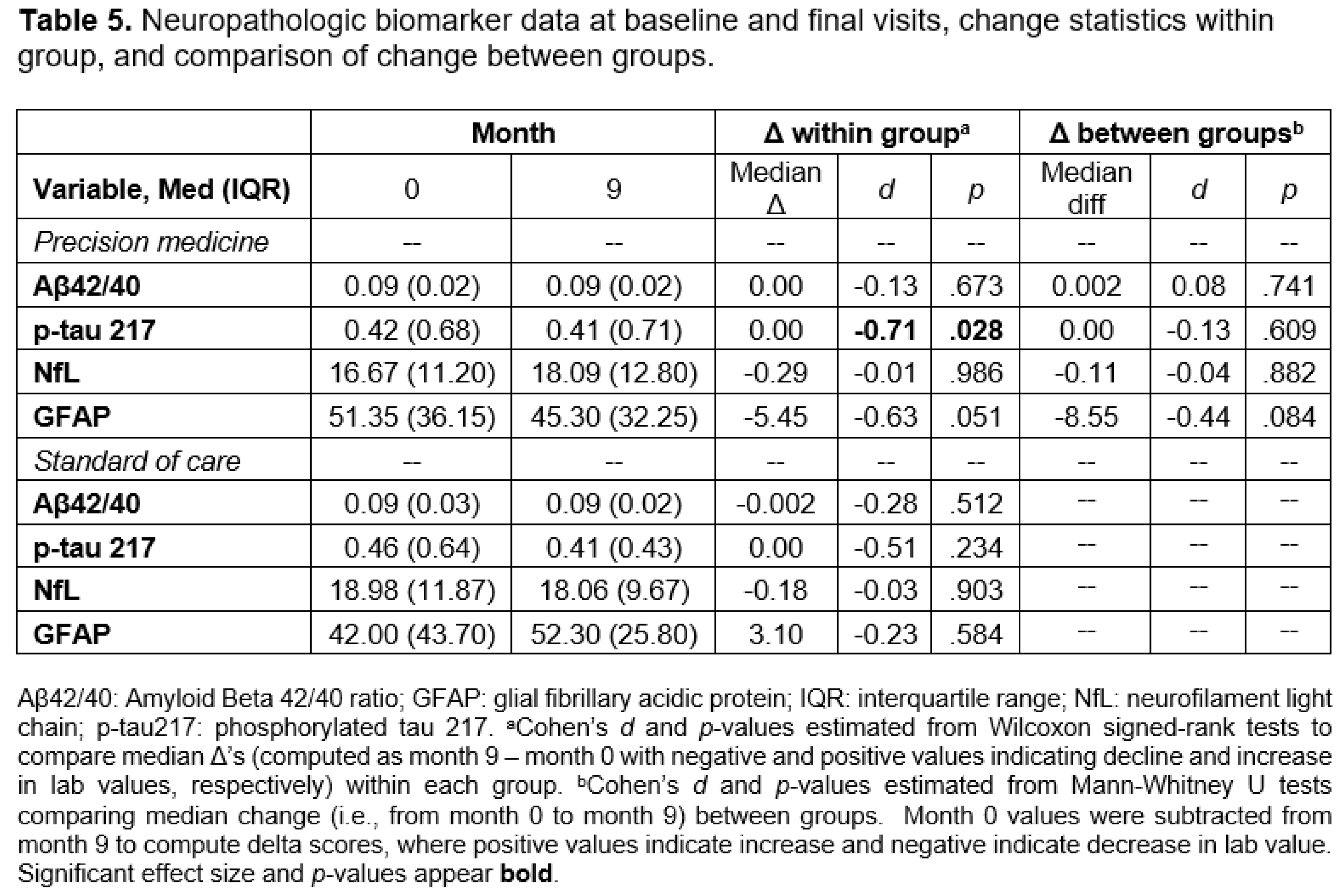
Safety
Discussion
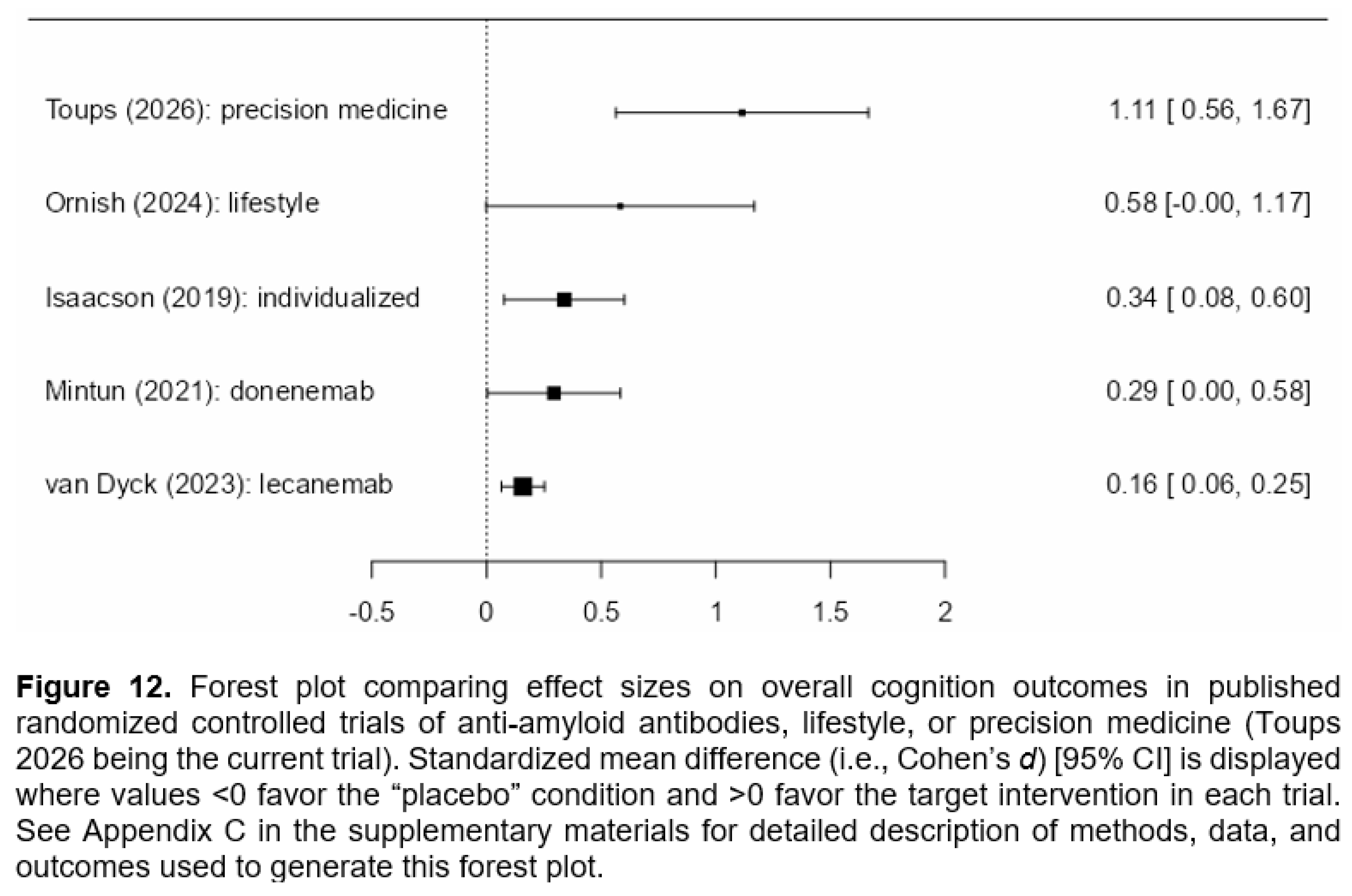
Limitations of the Study
Supplementary Materials
Funding
Acknowledgments
Declaration of conflicting interests
References
- James BD, Leurgans SE, Hebert LE, et al. Contribution of Alzheimer disease to mortality in the United States. Neurology 2014; 82: 1045–1050.
- Mintun MA, Lo AC, Evans CD, et al. Donanemab in Early Alzheimer’s Disease. New England Journal of Medicine 2021; 384: 1691–1704.
- Kato S, Kim KH, Lim HJ, et al. Real-world data from a molecular tumor board demonstrates improved outcomes with a precision N-of-One strategy. Nat Commun 2020; 11: 4965.
- Toups K, Hathaway A, Gordon D, et al. Precision Medicine Approach to Alzheimer’s Disease: Successful Pilot Project. Journal of Alzheimer’s Disease 2022; 88: 1411–1421.
- Sandison H, Callan NGL, Rao R V., et al. Observed Improvement in Cognition during a Personalized Lifestyle Intervention in People with Cognitive Decline. Journal of Alzheimer’s Disease 2023; 94: 993–1004.
- Kandimalla R, Thirumala V, Reddy PH. Is Alzheimer’s disease a Type 3 Diabetes? A critical appraisal. Biochim Biophys Acta Mol Basis Dis 2017; 1863: 1078–1089.
- Itzhaki RF, Lathe R, Balin BJ, et al. Microbes and Alzheimer’s Disease. J Alzheimers Dis 2016; 51: 979–984.
- Karran E, De Strooper B. The amyloid cascade hypothesis: are we poised for success or failure? J Neurochem 2016; 139: 237–252.
- Kametani F, Hasegawa M. Reconsideration of amyloid hypothesis and tau hypothesis in Alzheimer’s disease. Front Neurosci 2018; 12: 328460.
- Aoyagi A, Condello C, Stöhr J, et al. Aβ and tau prion-like activities decline with longevity in the Alzheimer’s disease human brain. Sci Transl Med; 11. Epub ahead of print 2019. DOI: 10.1126/SCITRANSLMED.AAT8462.
- Heneka MT, Carson MJ, Khoury J El, et al. Neuroinflammation in Alzheimer’s disease. Lancet Neurol 2015; 14: 388–405.
- Craft S. Insulin Resistance and Alzheimers Disease Pathogenesis: Potential Mechanisms and Implications for Treatment. Curr Alzheimer Res 2007; 4: 147–152.
- Ye X, Tai W, Zhang D. The early events of Alzheimer’s disease pathology: From mitochondrial dysfunction to BDNF axonal transport deficits. Neurobiol Aging 2012; 33: 1122.e1-1122.e10.
- Bredesen DE. Reversal of cognitive decline: a novel therapeutic program. Aging 2014; 6: 707–717.
- Bredesen DE, Amos EC, Canick J, et al. Reversal of cognitive decline in Alzheimer’s disease. Aging 2016; 8: 1250–1258.
- Bredesen DE, Sharlin K, Jenkins D, et al. Reversal of Cognitive Decline: 100 Patients. J Alzheimers Dis Parkinsonism 2018; 8: 1–6.
- Bredesen DE, Toups K, Hathaway A, et al. Precision Medicine Approach to Alzheimer’s Disease: Rationale and Implications. J Alzheimers Dis 2023; 96: 429–437.
- Teipel S, Gustafson D, Ossenkoppele R, et al. Alzheimer Disease: Standard of Diagnosis, Treatment, Care, and Prevention. Journal of Nuclear Medicine 2022; 63: 981.
- Nasreddine ZS, Phillips NA, Bédirian V, et al. The Montreal Cognitive Assessment, MoCA: A brief screening tool for mild cognitive impairment. J Am Geriatr Soc. Epub ahead of print 2005. DOI: 10.1111/j.1532-5415.2005.53221.x.
- Gualtieri CT, Johnson LG. Reliability and validity of a computerized neurocognitive test battery, CNS Vital Signs. Arch Clin Neuropsychol 2006; 21: 623–643.
- Bojar I, Wójcik-Fatla A, Owoc A, et al. Polymorphisms of apolipoprotein E gene and cognitive functions of postmenopausal women, measured by battery of computer tests - central nervous system vital signs. Activitas Nervosa Superior Rediviva; 54.
- Malek-Ahmadi M, Sabbagh MN. Development and Validation of the Alzheimer’s Questionnaire (AQ). J Nat Sci 2015; 1: e104.
- Hays RD, Bjorner JB, Revicki DA, et al. Development of physical and mental health summary scores from the patient-reported outcomes measurement information system (PROMIS) global items. Qual Life Res 2009; 18: 873–880.
- Attarha M, Mahncke H, Merzenich M. The Real-World Usability, Feasibility, and Performance Distributions of Deploying a Digital Toolbox of Computerized Assessments to Remotely Evaluate Brain Health: Development and Usability Study. JMIR Form Res 2024; 8: e53623.
- Attarha M, Carolina de Figueiredo Pelegrino A, Ouellet L, et al. Bringing Executive Function Testing Online: Assessment Validation Study. JMIR Form Res 2025; 9: e75687.
- Attarha M, De Figueiredo Pelegrino A, Ouellet L, et al. Association of a Brief Computerized Cognitive Assessment With Cholinergic Neurotransmission: Assessment Validation Study. JMIR Form Res 2025; 9: e68374.
- Ahdidan J, Raji CA, DeYoe EA, et al. Quantitative neuroimaging software for clinical assessment of hippocampal volumes on MR imaging. Journal of Alzheimer’s Disease 2015; 49: 723–732.
- Cunnane SC, Sieber CC, Swerdlow RH, et al. Mild cognitive impairment: when nutrition helps brain energy rescue-a report from the EuGMS 2020 Congress. Eur Geriatr Med 2021; 12: 1285–1292.
- H. Field L. L, Edwards SD, Edwards DJ, et al. Influence of HeartMath Training Programme on Physiological and Psychological Variables. Glob J Health Sci 2018; 10: p126.
- Shah TM, Weinborn M, Verdile G, et al. Enhancing Cognitive Functioning in Healthly Older Adults: a Systematic Review of the Clinical Significance of Commercially Available Computerized Cognitive Training in Preventing Cognitive Decline. Neuropsychol Rev 2017; 27: 62–80.
- Attarha M, Pelegrino A de F, Ouellet L, et al. Effects of Computerized Cognitive Training on Vesicular Acetylcholine Transporter Levels using [18F]Fluoroethoxybenzovesamicol Positron Emission Tomography in Healthy Older Adults: Results from the Improving Neurological Health in Aging via Neuroplastici…. JMIR Serious Games 2025;13:e75161 https://games.jmir.org/2025/1/e75161 2025; 13: e75161.
- Smith GE, Housen P, Yaffe K, et al. A cognitive training program based on principles of brain plasticity: results from the Improvement in Memory with Plasticity-based Adaptive Cognitive Training (IMPACT) study. J Am Geriatr Soc 2009; 57: 594–603.
- Rebok GW, Ball K, Guey LT, et al. Ten-year effects of the advanced cognitive training for independent and vital elderly cognitive training trial on cognition and everyday functioning in older adults. J Am Geriatr Soc 2014; 62: 16–24.
- Wharton W, Gleason CE, Lorenze KR, et al. Potential role of estrogen in the pathobiology and prevention of Alzheimer’s disease. Am J Transl Res 2009; 1: 131.
- Brown N, Panksepp J. Low-dose naltrexone for disease prevention and quality of life. Med Hypotheses 2009; 72: 333–337.
- Miklossy J. Alzheimer’s disease - a neurospirochetosis. Analysis of the evidence following Koch’s and Hill’s criteria. J Neuroinflammation; 8. Epub ahead of print 4 August 2011. DOI: 10.1186/1742-2094-8-90.
- Feng J, Leone J, Schweig S, et al. Evaluation of Natural and Botanical Medicines for Activity Against Growing and Non-growing Forms of B. burgdorferi. Front Med (Lausanne) 2020; 7: 492971.
- Ramanishankar A, Ankul Singh S, Begum RF, et al. Unleashing light’s healing power: an overview of photobiomodulation for Alzheimer’s treatment. Future Sci OA 2024; 10: FSO922.
- Ferguson CJ. An Effect Size Primer: A Guide for Clinicians and Researchers. Prof Psychol Res Pr 2009; 40: 532–538.
- Cohen J. Statistical power analysis for the behavioural sciences. 2nd ed. Lawrence Erlbaum Associates, 1988.
- Resources for Researchers: Common Terminology Criteria for Adverse Events (CTCAE) - National Cancer Institute, https://www.cancer.gov/research/resources/resource/210 (accessed 7 March 2026).
- Dyck CH van, Swanson CJ, Aisen P, et al. Lecanemab in Early Alzheimer’s Disease. New England Journal of Medicine 2023; 22: 142–143.
- Ornish D, Madison C, Kivipelto M, et al. Effects of intensive lifestyle changes on the progression of mild cognitive impairment or early dementia due to Alzheimer’s disease: a randomized, controlled clinical trial. Alzheimer’s Research & Therapy 2024 16:1 2024; 16: 122-.
- Isaacson RS, Hristov H, Saif N, et al. Individualized clinical management of patients at risk for Alzheimer’s dementia. Alzheimers Dement 2019; 15: 1588–1602.
- Mohammadi S, Rahmani F, Dolatshahi M, et al. Effects of obesity on plasma biomarker and amyloid PET trajectories in Alzheimer’s disease. Alzheimers Dement (Amst); 17. Epub ahead of print 1 July 2025. DOI: 10.1002/DAD2.70143.
- Hardy J, Allsop D. Amyloid deposition as the central event in the aetiology of Alzheimer’s disease. Trends Pharmacol Sci 1991; 12: 383–388.
- Ngandu T, Lehtisalo J, Solomon A, et al. A 2 year multidomain intervention of diet, exercise, cognitive training, and vascular risk monitoring versus control to prevent cognitive decline in at-risk elderly people (FINGER): A randomised controlled trial. The Lancet 2015; 385: 2255–2263.
- Cooley SA, Heaps JM, Bolzenius JD, et al. Longitudinal change in performance on the Montreal Cognitive Assessment in older adults. Clin Neuropsychol 2015; 29: 824.
- Jutten RJ, Grandoit E, Foldi NS, et al. Lower practice effects as a marker of cognitive performance and dementia risk: A literature review. Alzheimer’s & Dementia: Diagnosis, Assessment & Disease Monitoring 2020; 12: e12055.
- Elston DM. Participation bias, self-selection bias, and response bias. J Am Acad Dermatol; 0. Epub ahead of print 2021. DOI: 10.1016/j.jaad.2021.06.025.
References
- Aryee MJ, Jaffe AE, Corrada-Bravo H, et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 2014; 30: 1363–1369.
- Xu Z, Niu L, Li L, et al. ENmix: a novel background correction method for Illumina HumanMethylation450 BeadChip. Nucleic Acids Res; 44. Epub ahead of print 18 February 2016. [CrossRef]
- Triche TJ, Weisenberger DJ, Van Den Berg D, et al. Low-level processing of Illumina Infinium DNA Methylation BeadArrays. Nucleic Acids Res; 41. Epub ahead of print April 2013. [CrossRef]
- Fortin JP, Triche TJ, Hansen KD. Preprocessing, normalization and integration of the Illumina HumanMethylationEPIC array with minfi. Bioinformatics 2016; 33: 558.
- Du P, Zhang X, Huang CC, et al. Comparison of Beta-value and M-value methods for quantifying methylation levels by microarray analysis. BMC Bioinformatics; 11. Epub ahead of print 30 November 2010. [CrossRef]
- Horvath S. DNA methylation age of human tissues and cell types. Genome Biol; 14. Epub ahead of print 21 October 2013. [CrossRef]
- Hannum G, Guinney J, Zhao L, et al. Genome-wide Methylation Profiles Reveal Quantitative Views of Human Aging Rates. Mol Cell 2013; 49: 359–367.
- Levine ME, Lu AT, Quach A, et al. An epigenetic biomarker of aging for lifespan and healthspan. Aging (Albany NY) 2018; 10: 573.
- Lu AT, Quach A, Wilson JG, et al. DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging (Albany NY) 2019; 11: 303.
- Lu AT, Binder AM, Zhang J, et al. DNA methylation GrimAge version 2. Aging 2022; 14: 9484–9549.
- Pelegi-Siso D, De Prado P, Ronkainen J, et al. methylclock: a Bioconductor package to estimate DNA methylation age. Bioinformatics 2021; 37: 1759–1760.
- de Lima Camillo LP. pyaging: a Python-based compendium of GPU-optimized aging clocks. Bioinformatics; 40. Epub ahead of print 29 March 2024. [CrossRef]
- Ying K, Paulson S, Eames A, et al. A unified framework for systematic curation and evaluation of aging biomarkers. Nature Aging 2025 5:11 2025; 5: 2323–2339.
- Belsky DW, Caspi A, Corcoran DL, et al. DunedinPACE, A DNA methylation biomarker of the Pace of Aging. Elife; 11. Epub ahead of print 1 January 2022. [CrossRef]
- Higgins-Chen AT, Thrush KL, Wang Y, et al. A computational solution for bolstering reliability of epigenetic clocks: Implications for clinical trials and longitudinal tracking. Nat Aging 2022; 2: 644.
- Shireby GL, Davies JP, Francis PT, et al. Recalibrating the epigenetic clock: implications for assessing biological age in the human cortex. Brain 2020; 143: 3763–3775.
- Luo Q, Dwaraka VB, Chen Q, et al. A meta-analysis of immune-cell fractions at high resolution reveals novel associations with common phenotypes and health outcomes. Genome Medicine 2023 15:1 2023; 15: 59-.
- Teschendorff AE, Breeze CE, Zheng SC, et al. A comparison of reference-based algorithms for correcting cell-type heterogeneity in Epigenome-Wide Association Studies. BMC Bioinformatics 2017 18:1 2017; 18: 105-.
- Gadd DA, Hillary RF, McCartney DL, et al. Epigenetic scores for the circulating proteome as tools for disease prediction. Elife; 11. Epub ahead of print 1 January 2022. [CrossRef]
- Chen Q, Dwaraka VB, Carreras-Gallo N, et al. OMICmAge: An integrative multi-omics approach to quantify biological age with electronic medical records. bioRxiv 2023; 2023.10.16.562114.
- Carreras-Gallo N, Chen Q, Balagué-Dobón L, et al. Leveraging DNA methylation to create Epigenetic Biomarker Proxies that inform clinical care: A new framework for Precision Medicine. medRxiv 2024; 2024.12.06.24318612.
- Benjamini Y, Hochberg Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J R Stat Soc Series B Stat Methodol 1995; 57: 289–300.
- Du P, Zhang X, Huang CC, et al. Comparison of Beta-value and M-value methods for quantifying methylation levels by microarray analysis. BMC Bioinformatics; 11. Epub ahead of print 30 November 2010. [CrossRef]
- Smyth GK. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol; 3. Epub ahead of print 2004. [CrossRef]
- Ritchie ME, Phipson B, Wu D, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 2015; 43: e47.
- Peters TJ, Buckley MJ, Statham AL, et al. De novo identification of differentially methylated regions in the human genome. Epigenetics Chromatin 2015; 8: 6.
- GitHub - jokergoo/IlluminaHumanMethylationEPICv2anno.20a1.hg38, https://github.com/jokergoo/IlluminaHumanMethylationEPICv2anno.20a1.hg38 (accessed 10 January 2026).
- Yu G, Wang LG, He QY. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 2015; 31: 2382–2383.
- Yu G, Wang LG, Han Y, et al. clusterProfiler: an R Package for Comparing Biological Themes Among Gene Clusters. OMICS 2012; 16: 284.
- Ren X, Kuan PF. methylGSA: a Bioconductor package and Shiny app for DNA methylation data length bias adjustment in gene set testing. Bioinformatics 2019; 35: 1958–1959.
- Liberzon A, Birger C, Thorvaldsdóttir H, et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst 2015; 1: 417.
- Maksimovic J, Oshlack A, Phipson B. Gene set enrichment analysis for genome-wide DNA methylation data. Genome Biol 2021; 22: 173.
- Gu Z, Hübschmann D. rGREAT: an R/bioconductor package for functional enrichment on genomic regions. Bioinformatics; 39. Epub ahead of print 1 January 2023. [CrossRef]
- Mintun MA, Lo AC, Evans CD, et al. Donanemab in Early Alzheimer’s Disease. New England Journal of Medicine 2021; 384: 1691–1704.
- Borenstein M, Hedges L, Higgins J, et al. Comprehensive Meta-Analysis, https://www.meta-analysis.com/ (2022, accessed 26 June 2023).
- Dyck CH van, Swanson CJ, Aisen P, et al. Lecanemab in Early Alzheimer’s Disease. New England Journal of Medicine 2023; 22: 142–143.
- Ornish D, Madison C, Kivipelto M, et al. Effects of intensive lifestyle changes on the progression of mild cognitive impairment or early dementia due to Alzheimer’s disease: a randomized, controlled clinical trial. Alzheimer’s Research & Therapy 2024 16:1 2024; 16: 122-.
- Isaacson RS, Hristov H, Saif N, et al. Individualized clinical management of patients at risk for Alzheimer’s dementia. Alzheimers Dement 2019; 15: 1588–1602.
- The jamovi project. jamovi, Retrieved from https://www.jamovi.org (2025, accessed 27 February 2026).
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



