Submitted:
14 March 2024
Posted:
15 March 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. COVID-19-Related Viruses
1.2. Peculiar Relationships between SARS-CoV-2, Omicron, and Pangolin-CoVs
2. Materials and Methods
2.1. Computational Biology: SDMs and Protein Intrinsic Disorder
2.2. Experimental Biology: Cells and Viruses
2.3. Experimental Biology: Viral One-Step Growth Curve
2.4. Experimental Biology: Cytopathic Effect Analysis (CPE) and Plaque Assay
3. Results
3.1. The Shell Disorder Models (SDMs)
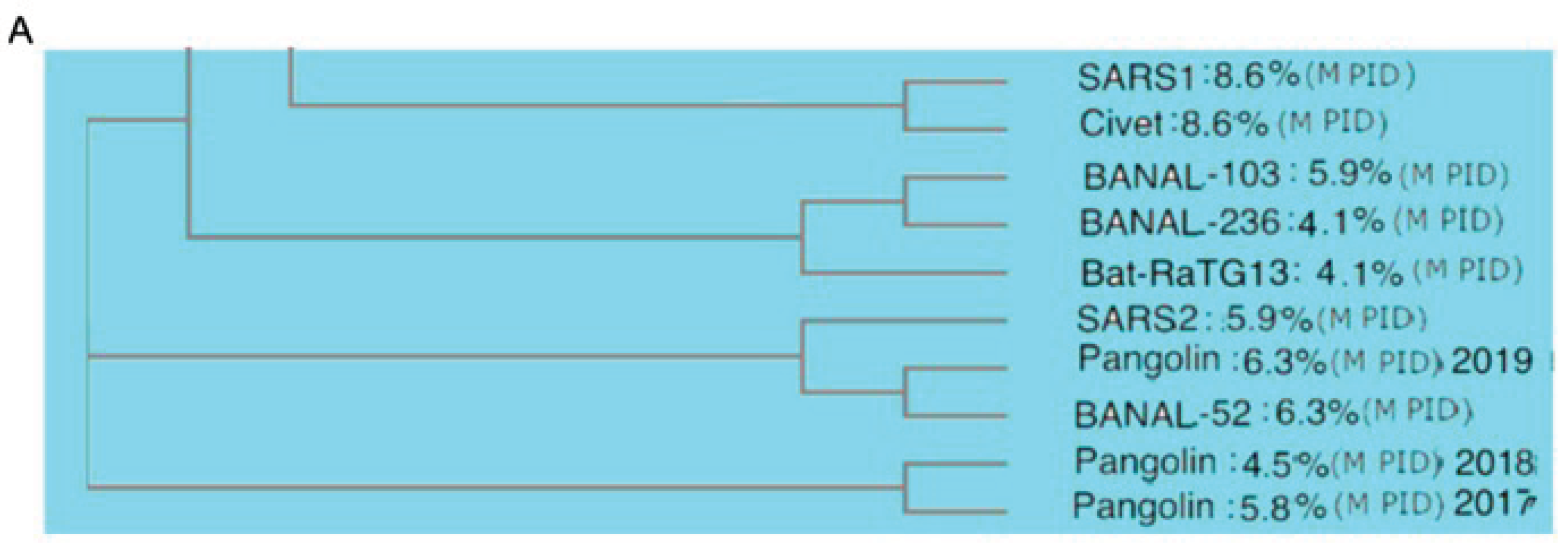
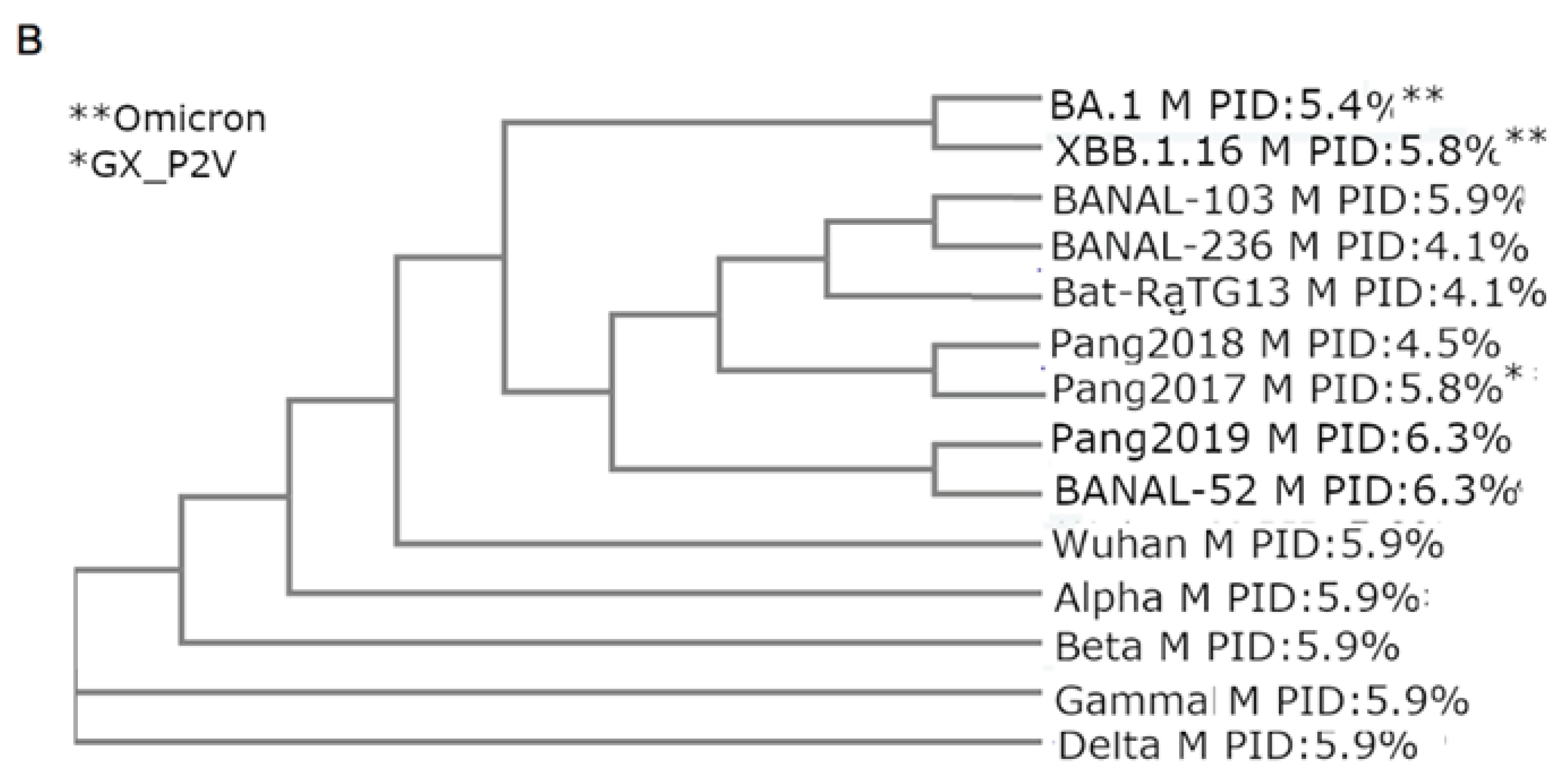
3.2. Phylogenetic Study Using M Reveals Intimate Relationship Between Pangolin-CoV and SARS-CoV-2/Omicron
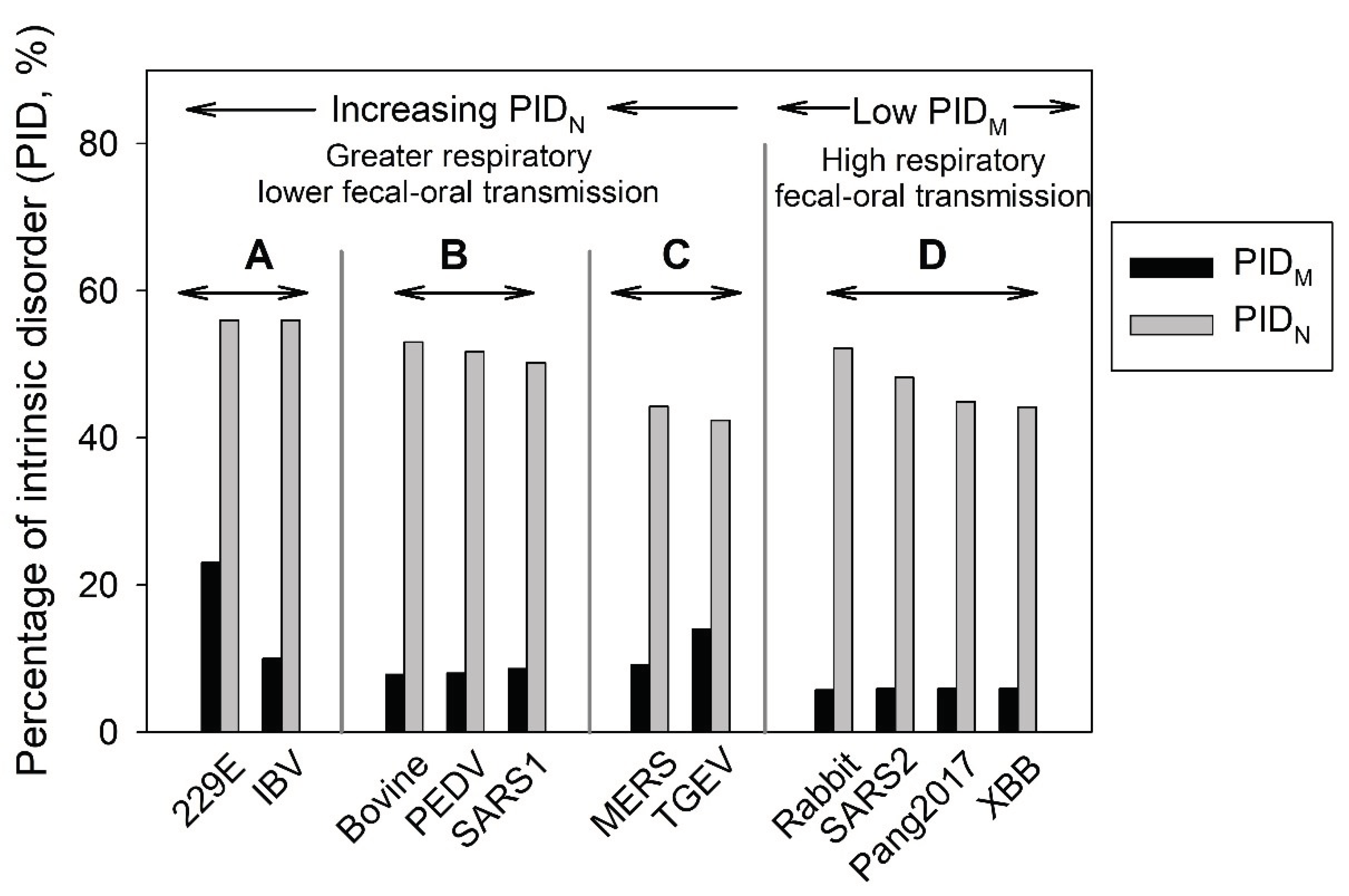
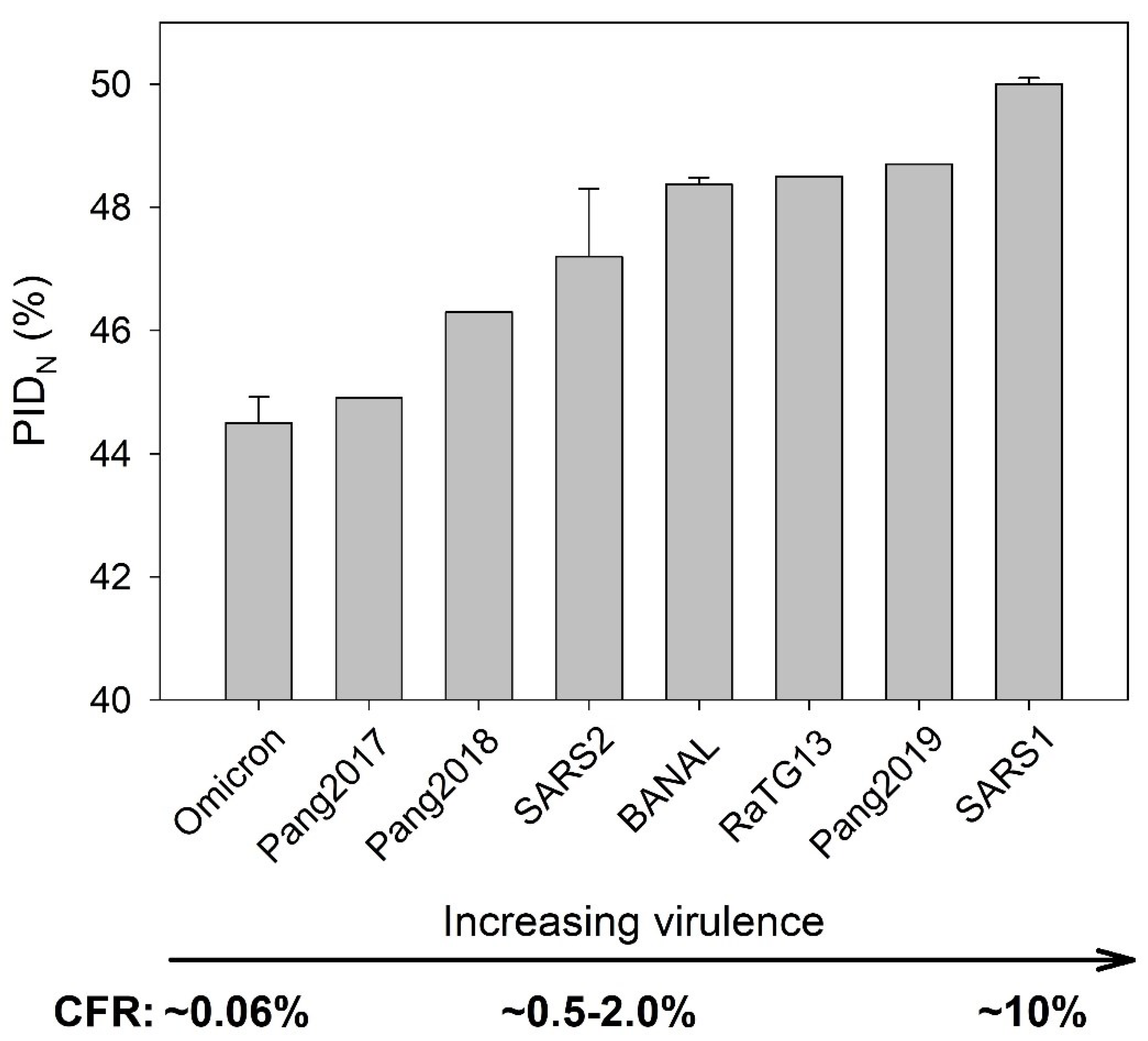
3.3. Omicron and Pang2017: Low PIDN and Attenuation
3.4. Omicron Has a Lower PIDN Similar to Pango2017 but has a lower PIDM: Attenuation and Faster Spread
3.5. The Role of N in CoV-Transmission SDM and Virulence-Inner Shell Disorder Model
3.6. All Known COVID-19 Viruses have Hard Outer Shell: Evolutionary Association with Pangolins
3.7. Correlation Between Viral Growth and N Disorder of COVID-19-Related Viruses
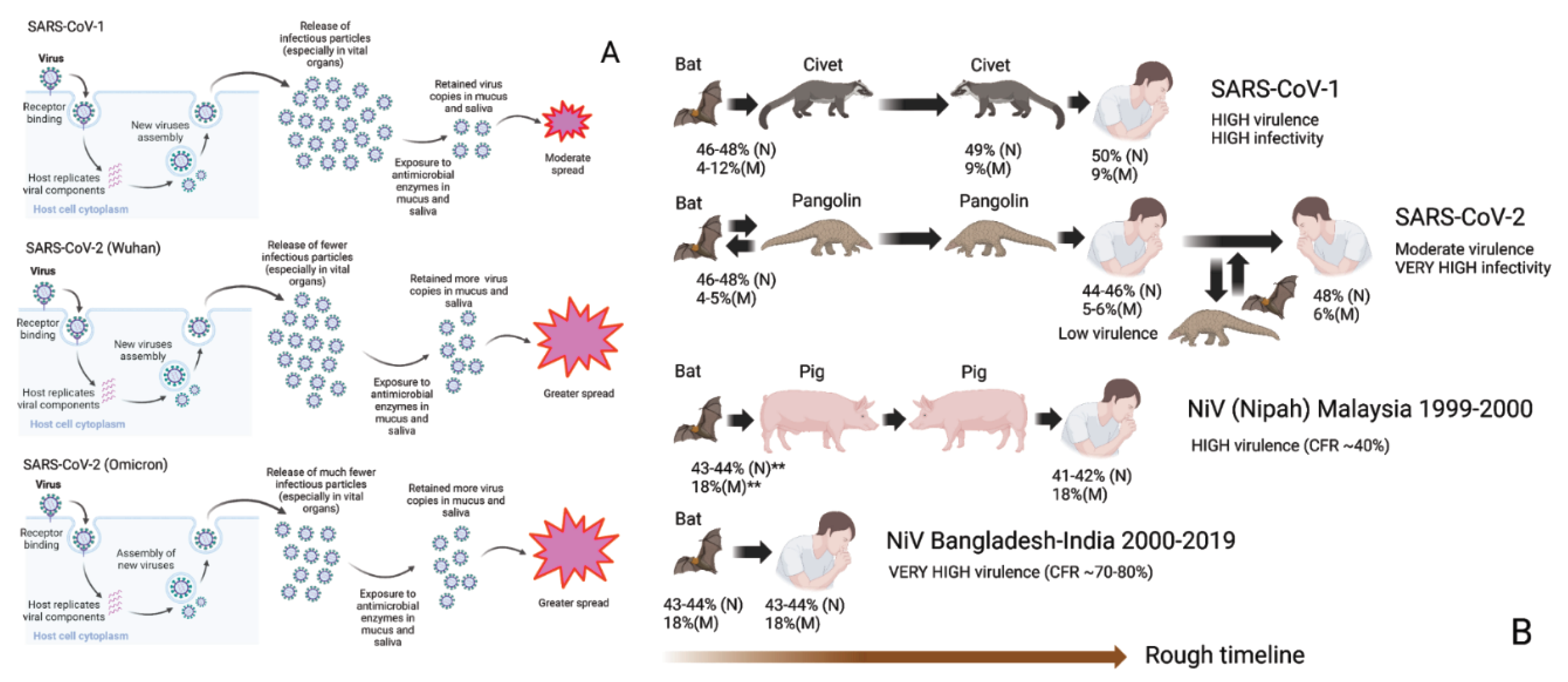
3.8. Molecular Analysis SARS-CoV-2's Evolution within Animals Affects its Virulence and Human Spread
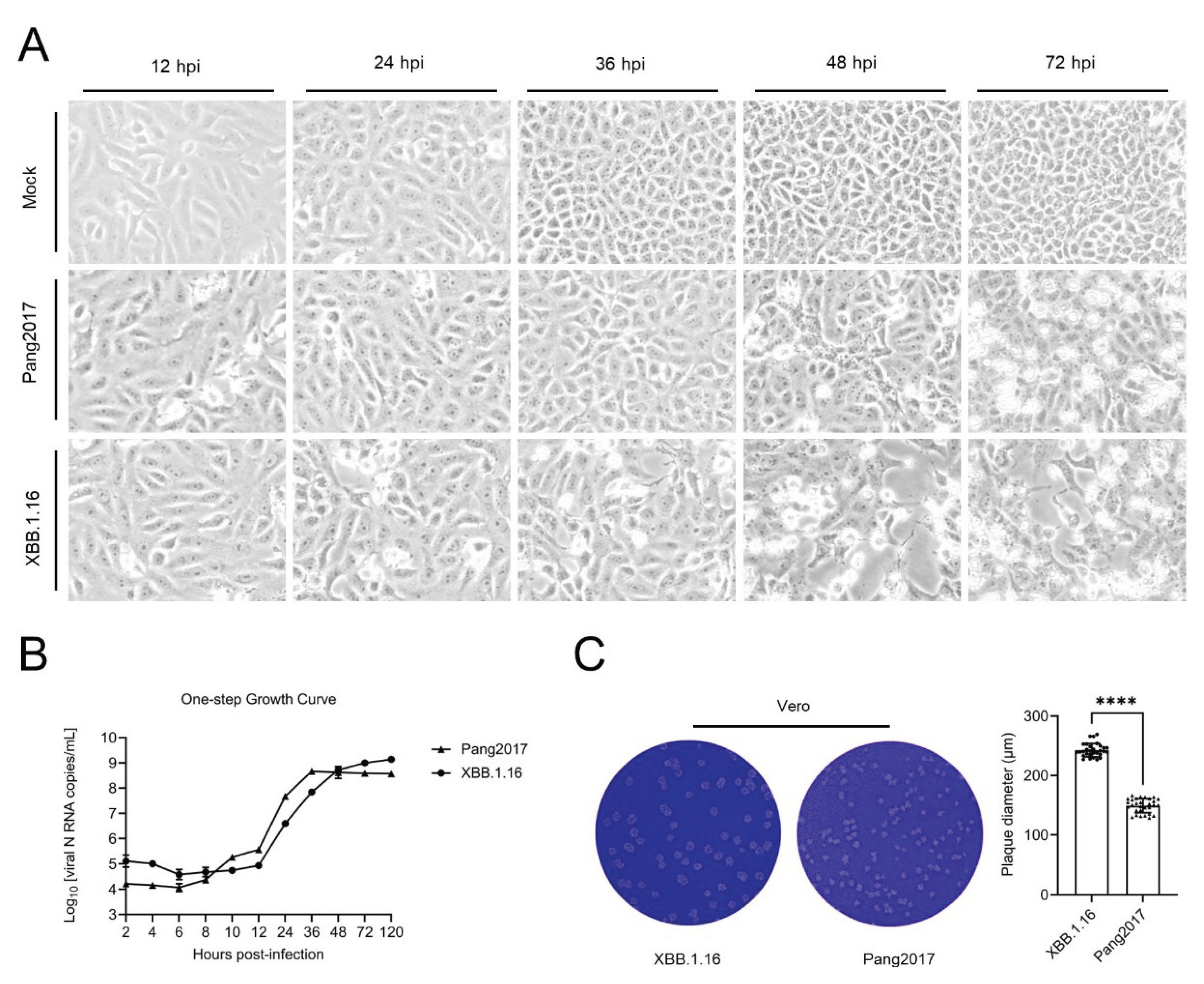
3.9. Comparison of Cytopathic Effects, One-Step Growth Curve, and Plaque Size of Pang2017 and SARS-CoV-2 XBB.1.16 in Vero Cells
4. Discussion
4.1. COVID-19 Special Relationship with Pangolin-CoVs: Can Be Found in the Abnormally Hard M: Burrowing Animal
4.2. Evidence of an Even Closer Relationship Between Omicron and Pangolin-CoVs
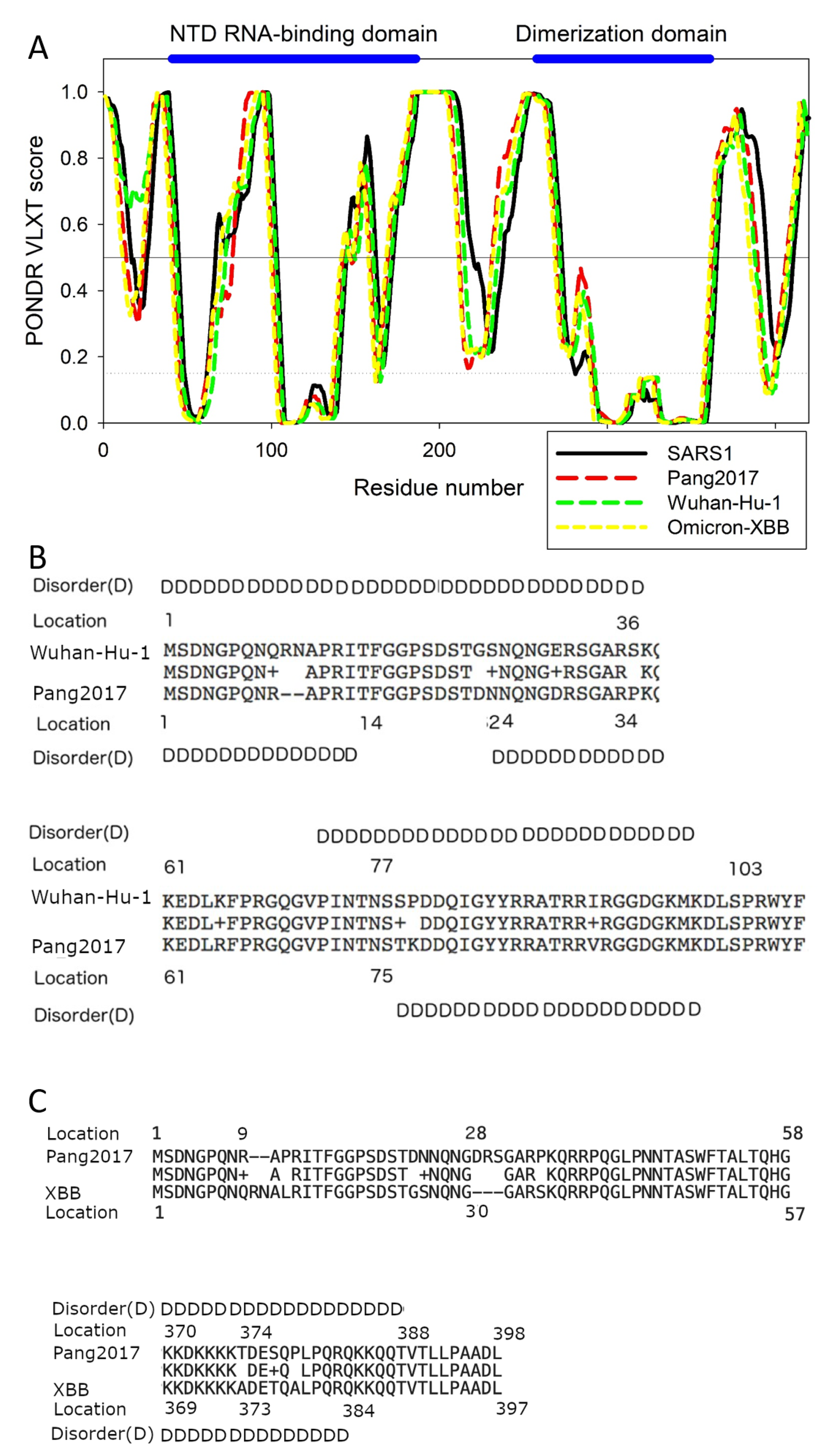
4.3. Range of SARS-CoV-2 N Disorder Matches that Pangolin-CoVs2017, Not Bat-CoVs
4.4. Differences in Pang2017 and XBB.1.16 N Disorder Patterns Can Explain Subtle Discrepancy in Experimental Results for the Two Viruses
5. Summary and Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Worobey, M.; Levy, J.I.; Malpica Serrano, L.; Crits-Christoph, A.; Pekar, J.E.; Goldstein, S.A.; Rasmussen, A.L.; Kraemer, M.U.G.; Newman, C.; Koopmans, M.P.G.; et al. The Huanan Seafood Wholesale Market in Wuhan was the early epicenter of the COVID-19 pandemic. Science 2022, 377, 951–959. [Google Scholar] [CrossRef]
- Pekar, J.E.; Magee, A.; Parker, E.; Moshiri, N.; Izhikevich, K.; Havens, J.L.; Gangavarapu, K.; Malpica Serrano, L.M.; Crits-Christoph, A.; Matteson, N.L.; et al. The molecular epidemiology of multiple zoonotic origins of SARS-CoV-2. Science 2022, 377, 960–966. [Google Scholar] [CrossRef]
- Hu, B.; Guo, H.; Zhou, P.; Shi, Z.L. Characteristics of SARS-CoV-2 and COVID-19. Nat Rev Microbiol 2021, 19, 141–154. [Google Scholar] [CrossRef] [PubMed]
- WHO. Coronavirus disease (COVID-19) pandemic. Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019 (accessed on).
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. Addendum: A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 588, E6. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zai, J.; Zhao, Q.; Nie, Q.; Li, Y.; Foley, B.T.; Chaillon, A. Evolutionary history, potential intermediate animal host, and cross-species analyses of SARS-CoV-2. J Med Virol 2020, 92, 602–611. [Google Scholar] [CrossRef] [PubMed]
- Low, Z.Y.; Yip, A.J.W.; Sharma, A.; Lal, S.K. SARS coronavirus outbreaks past and present-a comparative analysis of SARS-CoV-2 and its predecessors. Virus Genes 2021, 57, 307–317. [Google Scholar] [CrossRef] [PubMed]
- Xiao, K.; Zhai, J.; Feng, Y.; Zhou, N.; Zhang, X.; Zou, J.J.; Li, N.; Guo, Y.; Li, X.; Shen, X.; et al. Isolation of SARS-CoV-2-related coronavirus from Malayan pangolins. Nature 2020, 583, 286–289. [Google Scholar] [CrossRef]
- Fan, H.H.; Wang, L.Q.; Liu, W.L.; An, X.P.; Liu, Z.D.; He, X.Q.; Song, L.H.; Tong, Y.G. Repurposing of clinically approved drugs for treatment of coronavirus disease 2019 in a 2019-novel coronavirus-related coronavirus model. Chin Med J (Engl) 2020, 133, 1051–1056. [Google Scholar] [CrossRef]
- Lam, T.T.; Jia, N.; Zhang, Y.W.; Shum, M.H.; Jiang, J.F.; Zhu, H.C.; Tong, Y.G.; Shi, Y.X.; Ni, X.B.; Liao, Y.S.; et al. Identifying SARS-CoV-2-related coronaviruses in Malayan pangolins. Nature 2020, 583, 282–285. [Google Scholar] [CrossRef] [PubMed]
- Temmam, S.; Vongphayloth, K.; Baquero, E.; Munier, S.; Bonomi, M.; Regnault, B.; Douangboubpha, B.; Karami, Y.; Chretien, D.; Sanamxay, D.; et al. Bat coronaviruses related to SARS-CoV-2 and infectious for human cells. Nature 2022, 604, 330–336. [Google Scholar] [CrossRef]
- Nga, N.T.T.; Latinne, A.; Thuy, H.B.; Long, N.V.; Ngoc, P.T.B.; Anh, N.T.L.; Thai, N.V.; Phuong, T.Q.; Thai, H.V.; Hai, L.K.; et al. Evidence of SARS-CoV-2 Related Coronaviruses Circulating in Sunda pangolins (Manis javanica) Confiscated From the Illegal Wildlife Trade in Viet Nam. Front Public Health 2022, 10, 826116. [Google Scholar] [CrossRef]
- Garner, E.; Romero, P.; Dunker, A.K.; Brown, C.; Obradovic, Z. Predicting Binding Regions within Disordered Proteins. Genome Inform Ser Workshop Genome Inform 1999, 10, 41–50. [Google Scholar] [PubMed]
- Li, X.; Romero, P.; Rani, M.; Dunker, A.K.; Obradovic, Z. Predicting Protein Disorder for N-, C-, and Internal Regions. Genome Inform Ser Workshop Genome Inform 1999, 10, 30–40. [Google Scholar] [PubMed]
- Romero, P.; Obradovic, Z.; Li, X.; Garner, E.C.; Brown, C.J.; Dunker, A.K. Sequence complexity of disordered protein. Proteins 2001, 42, 38–48. [Google Scholar] [CrossRef] [PubMed]
- Goh, G.K.; Dunker, A.K.; Uversky, V.N. Protein intrinsic disorder toolbox for comparative analysis of viral proteins. BMC Genomics 2008, 9 (Suppl. S2), S4. [Google Scholar] [CrossRef] [PubMed]
- Goh, G.K. Viral shapeshifters: strange behaviors of HIV and other viruses; Simplicity Research Institute: 2017.
- Goh, G.K.; Dunker, A.K.; Foster, J.A.; Uversky, V.N. Shell Disorder Analysis Suggests That Pangolins Offered a Window for a Silent Spread of an Attenuated SARS-CoV-2 Precursor among Humans. J Proteome Res 2020, 19, 4543–4552. [Google Scholar] [CrossRef] [PubMed]
- Goh, G.K.; Dunker, A.K.; Foster, J.A.; Uversky, V.N. Shell disorder analysis predicts greater resilience of the SARS-CoV-2 (COVID-19) outside the body and in body fluids. Microb Pathog 2020, 144, 104177. [Google Scholar] [CrossRef]
- Goh, G.K.; Dunker, A.K.; Foster, J.A.; Uversky, V.N. Rigidity of the Outer Shell Predicted by a Protein Intrinsic Disorder Model Sheds Light on the COVID-19 (Wuhan-2019-nCoV) Infectivity. Biomolecules 2020, 10. [Google Scholar] [CrossRef]
- Goh, G.K.; Dunker, A.K.; Uversky, V.N. A comparative analysis of viral matrix proteins using disorder predictors. Virol J 2008, 5, 126. [Google Scholar] [CrossRef]
- Goh, G.K.; Dunker, A.K.; Foster, J.A.; Uversky, V.N. Zika and Flavivirus Shell Disorder: Virulence and Fetal Morbidity. Biomolecules 2019, 9. [Google Scholar] [CrossRef]
- Goh, G.K.; Dunker, A.K.; Foster, J.A.; Uversky, V.N. HIV Vaccine Mystery and Viral Shell Disorder. Biomolecules 2019, 9. [Google Scholar] [CrossRef]
- Goh, G.K.; Dunker, A.K.; Uversky, V.N. Correlating Flavivirus virulence and levels of intrinsic disorder in shell proteins: protective roles vs. immune evasion. Mol Biosyst 2016, 12, 1881–1891. [Google Scholar] [CrossRef]
- Goh, G.K.; Dunker, A.K.; Foster, J.A.; Uversky, V.N. Computational, Experimental, and Clinical Evidence of a Specific but Peculiar Evolutionary Nature of (COVID-19) SARS-CoV-2. J Proteome Res 2022, 21, 874–890. [Google Scholar] [CrossRef]
- Goh, G.K.; Dunker, A.K.; Foster, J.A.; Uversky, V.N. Shell Disorder Models Detect That Omicron Has Harder Shells with Attenuation but Is Not a Descendant of the Wuhan-Hu-1 SARS-CoV-2. Biomolecules 2022, 12. [Google Scholar] [CrossRef] [PubMed]
- Goh, G.K.; Dunker, A.K.; Uversky, V.N. Detection of links between Ebola nucleocapsid and virulence using disorder analysis. Mol Biosyst 2015, 11, 2337–2344. [Google Scholar] [CrossRef] [PubMed]
- Guo, Z.; Zhang, C.; Zhang, C.; Cui, H.; Chen, Z.; Jiang, X.; Wang, T.; Li, Y.; Liu, J.; Wan, Z.; et al. SARS-CoV-2-related pangolin coronavirus exhibits similar infection characteristics to SARS-CoV-2 and direct contact transmissibility in hamsters. iScience 2022, 25, 104350. [Google Scholar] [CrossRef]
- Lu, S.; Luo, S.; Liu, C.; Li, M.; An, X.; Li, M.; Hou, J.; Fan, H.; Mao, P.; Tong, Y.; et al. Induction of significant neutralizing antibodies against SARS-CoV-2 by a highly attenuated pangolin coronavirus variant with a 104nt deletion at the 3'-UTR. Emerg Microbes Infect 2023, 12, 2151383. [Google Scholar] [CrossRef] [PubMed]
- Goh, G.K.; Dunker, A.K.; Foster, J.A.; Uversky, V.N. A Study on the Nature of SARS-CoV-2 Using the Shell Disorder Models: Reproducibility, Evolution, Spread, and Attenuation. Biomolecules 2022, 12. [Google Scholar] [CrossRef]
- Hui, K.P.Y.; Ho, J.C.W.; Cheung, M.C.; Ng, K.C.; Ching, R.H.H.; Lai, K.L.; Kam, T.T.; Gu, H.; Sit, K.Y.; Hsin, M.K.Y.; et al. SARS-CoV-2 Omicron variant replication in human bronchus and lung ex vivo. Nature 2022, 603, 715–720. [Google Scholar] [CrossRef]
- Wolter, N.; Jassat, W.; Walaza, S.; Welch, R.; Moultrie, H.; Groome, M.; Amoako, D.G.; Everatt, J.; Bhiman, J.N.; Scheepers, C.; et al. Early assessment of the clinical severity of the SARS-CoV-2 omicron variant in South Africa: a data linkage study. Lancet 2022, 399, 437–446. [Google Scholar] [CrossRef]
- Sigal, A. Milder disease with Omicron: is it the virus or the pre-existing immunity? Nat Rev Immunol 2022, 22, 69–71. [Google Scholar] [CrossRef]
- Abdullah, F.; Myers, J.; Basu, D.; Tintinger, G.; Ueckermann, V.; Mathebula, M.; Ramlall, R.; Spoor, S.; de Villiers, T.; Van der Walt, Z.; et al. Decreased severity of disease during the first global omicron variant covid-19 outbreak in a large hospital in tshwane, south africa. Int J Infect Dis 2022, 116, 38–42. [Google Scholar] [CrossRef]
- Hou, Y.J.; Chiba, S.; Leist, S.R.; Meganck, R.M.; Martinez, D.R.; Schafer, A.; Catanzaro, N.J.; Sontake, V.; West, A.; Edwards, C.E.; et al. Host range, transmissibility and antigenicity of a pangolin coronavirus. Nat Microbiol 2023, 8, 1820–1833. [Google Scholar] [CrossRef]
- European_CDC. European Center for Disease Prevention and Control, Weekly epidemilogical update: Omicron variant of concern (VOC). Available online: https://www.ecdc.europa.eu/en/news-events/weekly-epidemiological-update-omicron-variant-concern-voc-week-2-data-20-january-2022 (accessed on Augusr=t 14).
- DeGrasse, D.C.; Black, S.D. The Rise of SARS-CoV-2 (COVID-19) Omicron Subvariant Pathogenicity. Cureus 2023, 15, e40148. [Google Scholar] [CrossRef]
- Kupferschmidt, K. Where did 'weird' Omicron come from? Science 2021, 374, 1179. [Google Scholar] [CrossRef] [PubMed]
- Bondos, S.E.; Dunker, A.K.; Uversky, V.N. Intrinsically disordered proteins play diverse roles in cell signaling. Cell Commun Signal 2022, 20, 20. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Oldfield, C.J.; Meng, J.; Romero, P.; Uversky, V.N.; Dunker, A.K. Mining alpha-helix-forming molecular recognition features with cross species sequence alignments. Biochemistry 2007, 46, 13468–13477. [Google Scholar] [CrossRef] [PubMed]
- Dunker, A.K.; Brown, C.J.; Lawson, C.J.D.; Iakoucheva-Sebat, L.M.; Vucetic, S.; Obradovic, Z. The Protein Trinity: Structure/Function Relationships That Include Intrinsic Disorder. ScientificWorldJournal 2002, 2, 49–50. [Google Scholar] [CrossRef] [PubMed]
- Dunker, A.K.; Brown, C.J.; Lawson, J.D.; Iakoucheva, L.M.; Obradovic, Z. Intrinsic disorder and protein function. Biochemistry 2002, 41, 6573–6582. [Google Scholar] [CrossRef] [PubMed]
- Dunker, A.K.; Lawson, J.D.; Brown, C.J.; Williams, R.M.; Romero, P.; Oh, J.S.; Oldfield, C.J.; Campen, A.M.; Ratliff, C.M.; Hipps, K.W.; et al. Intrinsically disordered protein. J Mol Graph Model 2001, 19, 26–59. [Google Scholar] [CrossRef] [PubMed]
- Mohan, A.; Oldfield, C.J.; Radivojac, P.; Vacic, V.; Cortese, M.S.; Dunker, A.K.; Uversky, V.N. Analysis of molecular recognition features (MoRFs). J Mol Biol 2006, 362, 1043–1059. [Google Scholar] [CrossRef]
- Oldfield, C.J.; Cheng, Y.; Cortese, M.S.; Romero, P.; Uversky, V.N.; Dunker, A.K. Coupled folding and binding with alpha-helix-forming molecular recognition elements. Biochemistry 2005, 44, 12454–12470. [Google Scholar] [CrossRef]
- Oldfield, C.J.; Dunker, A.K. Intrinsically disordered proteins and intrinsically disordered protein regions. Annu Rev Biochem 2014, 83, 553–584. [Google Scholar] [CrossRef]
- Tompa, P. Intrinsically unstructured proteins. Trends Biochem Sci 2002, 27, 527–533. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N. Natively unfolded proteins: a point where biology waits for physics. Protein Sci 2002, 11, 739–756. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N. What does it mean to be natively unfolded? Eur J Biochem 2002, 269, 2–12. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N. Unusual biophysics of intrinsically disordered proteins. Biochim Biophys Acta 2013, 1834, 932–951. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N.; Dunker, A.K. Understanding protein non-folding. Biochim Biophys Acta 2010, 1804, 1231–1264. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N.; Gillespie, J.R.; Fink, A.L. Why are "natively unfolded" proteins unstructured under physiologic conditions? Proteins 2000, 41, 415–427. [Google Scholar] [CrossRef]
- Vacic, V.; Oldfield, C.J.; Mohan, A.; Radivojac, P.; Cortese, M.S.; Uversky, V.N.; Dunker, A.K. Characterization of molecular recognition features, MoRFs, and their binding partners. J Proteome Res 2007, 6, 2351–2366. [Google Scholar] [CrossRef] [PubMed]
- van der Lee, R.; Buljan, M.; Lang, B.; Weatheritt, R.J.; Daughdrill, G.W.; Dunker, A.K.; Fuxreiter, M.; Gough, J.; Gsponer, J.; Jones, D.T.; et al. Classification of intrinsically disordered regions and proteins. Chem Rev 2014, 114, 6589–6631. [Google Scholar] [CrossRef]
- Wright, P.E.; Dyson, H.J. Intrinsically unstructured proteins: re-assessing the protein structure-function paradigm. J Mol Biol 1999, 293, 321–331. [Google Scholar] [CrossRef] [PubMed]
- Goh, G.K.; Dunker, A.K.; Uversky, V.N. Protein intrinsic disorder and influenza virulence: the 1918 H1N1 and H5N1 viruses. Virol J 2009, 6, 69. [Google Scholar] [CrossRef]
- Xue, B.; Williams, R.W.; Oldfield, C.J.; Goh, G.K.; Dunker, A.K.; Uversky, V.N. Viral disorder or disordered viruses: do viral proteins possess unique features? Protein Pept Lett 2010, 17, 932–951. [Google Scholar] [CrossRef]
- Goh, G.K.; Dunker, A.K.; Uversky, V.N. Understanding Viral Transmission Behavior via Protein Intrinsic Disorder Prediction: Coronaviruses. J Pathog 2012, 2012, 738590. [Google Scholar] [CrossRef] [PubMed]
- Goh, G.K.; Dunker, A.K.; Uversky, V. Prediction of Intrinsic Disorder in MERS-CoV/HCoV-EMC Supports a High Oral-Fecal Transmission. PLoS Curr 2013, 5. [Google Scholar] [CrossRef] [PubMed]
- Goh, G.K.; Dunker, A.K.; Foster, J.A.; Uversky, V.N. A Novel Strategy for the Development of Vaccines for SARS-CoV-2 (COVID-19) and Other Viruses Using AI and Viral Shell Disorder. J Proteome Res 2020, 19, 4355–4363. [Google Scholar] [CrossRef]
- Goh, G.K.; Dunker, A.K.; Foster, J.A.; Uversky, V.N. Feasibility of the vaccine development for SARS-CoV-2 and other viruses using the shell disorder analysis. Pac Symp Biocomput 2021, 26, 143–153. [Google Scholar] [PubMed]
- UniProt_Consortium. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res 2021, 49, D480–D489. [Google Scholar] [CrossRef]
- MySQL. Available online: https://www.mysql.com (accessed on August 14).
- JAVA: JAVA Programming Language. Available online: https://www.java.com (accessed on August, 14).
- Python: Python Programming Language. Available online: https://www.python.org (accessed on 22 December ).
- TREX: CLUSTALW. Available online: http://www.trex.uqam.ca/index.php?action=align&project=trex (accessed on August, 14).
- OpenOffice. Available online: https://www.openoffice.org/ (accessed on August, 14).
- GIMP: GNU Image Manipulator Program. Available online: https://www.gimp.org (accessed on August, 14).
- R_Core_Team. R: A language and environment for statistical computing. 2013.
- NCBI-BLASTP. Available online: https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE=Proteins (accessed on August, 14). Available online: https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE=Proteins (accessed on August, 14).
- Lu, S.; Luo, S.; Bai, B.; Fan, Z.; Hong, B.; Xu, W.; Ma, H.; Chen, W.; Fan, H.; Tong, Y.; et al. Omicron breakthrough infections in wild-type SARS-CoV-2 vaccinees elicit high levels of neutralizing antibodies against pangolin coronavirus GX_P2V. J Med Virol 2023, 95, e29031. [Google Scholar] [CrossRef]
- Goh, G.K.; Dunker, A.K.; Foster, J.A.; Uversky, V.N. Nipah shell disorder, modes of infection, and virulence. Microb Pathog 2020, 141, 103976. [Google Scholar] [CrossRef] [PubMed]
- Longhi, S.; Bloyet, L.M.; Gianni, S.; Gerlier, D. How order and disorder within paramyxoviral nucleoproteins and phosphoproteins orchestrate the molecular interplay of transcription and replication. Cell Mol Life Sci 2017, 74, 3091–3118. [Google Scholar] [CrossRef] [PubMed]
- Malamud, D.; Abrams, W.R.; Barber, C.A.; Weissman, D.; Rehtanz, M.; Golub, E. Antiviral activities in human saliva. Adv Dent Res 2011, 23, 34–37. [Google Scholar] [CrossRef] [PubMed]
- Ganz, T. Antimicrobial polypeptides in host defense of the respiratory tract. J Clin Invest 2002, 109, 693–697. [Google Scholar] [CrossRef] [PubMed]
- Wolfel, R.; Corman, V.M.; Guggemos, W.; Seilmaier, M.; Zange, S.; Muller, M.A.; Niemeyer, D.; Jones, T.C.; Vollmar, P.; Rothe, C.; et al. Virological assessment of hospitalized patients with COVID-2019. Nature 2020, 581, 465–469. [Google Scholar] [CrossRef] [PubMed]
- Fahy, J.V.; Dickey, B.F. Airway mucus function and dysfunction. N Engl J Med 2010, 363, 2233–2247. [Google Scholar] [CrossRef] [PubMed]
- Ward, H.E.; Nicholas, T.E. Alveolar type I and type II cells. Aust N Z J Med 1984, 14, 731–734. [Google Scholar] [CrossRef] [PubMed]
- Abrahams, P., (Ed.) How the Body Works: A Comprehensive Illustrated Encyclopedia of Anatomy. Amber Books Limited: 2016.
- Mire, C.E.; Satterfield, B.A.; Geisbert, J.B.; Agans, K.N.; Borisevich, V.; Yan, L.; Chan, Y.P.; Cross, R.W.; Fenton, K.A.; Broder, C.C.; et al. Pathogenic Differences between Nipah Virus Bangladesh and Malaysia Strains in Primates: Implications for Antibody Therapy. Sci Rep 2016, 6, 30916. [Google Scholar] [CrossRef]
- Santaniello, A.; Perruolo, G.; Cristiano, S.; Agognon, A.L.; Cabaro, S.; Amato, A.; Dipineto, L.; Borrelli, L.; Formisano, P.; Fioretti, A.; et al. SARS-CoV-2 Affects Both Humans and Animals: What Is the Potential Transmission Risk? A Literature Review. Microorganisms 2023, 11. [Google Scholar] [CrossRef]
- Uversky, V.N.; Elrashdy, F.; Aljadawi, A.; Redwan, E.M. Household pets and SARS-CoV2 transmissibility in the light of the ACE2 intrinsic disorder status. J Biomol Struct Dyn 2022, 40, 1441–1444. [Google Scholar] [CrossRef]
- Lehmann, D.; Halbwax, M.L.; Makaga, L.; Whytock, R.; Ndindiwe Malata, L.L.; Bombenda Mouele, W.; Momboua, B.R.; Koumba Pambo, A.F.; White, L.J.T. Pangolins and bats living together in underground burrows in Lope National Park, Gabon. Afr J Ecol 2020, 58, 540–542. [Google Scholar] [CrossRef] [PubMed]
- Posada, D. How does recombination affect phylogeny estimation? Trends Ecol Evol 2000, 15, 489–490. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Wu, Q.; Zhang, Z. Probable Pangolin Origin of SARS-CoV-2 Associated with the COVID-19 Outbreak. Curr Biol 2020, 30, 1346–1351. [Google Scholar] [CrossRef] [PubMed]
- Ogando, N.S.; Dalebout, T.J.; Zevenhoven-Dobbe, J.C.; Limpens, R.; van der Meer, Y.; Caly, L.; Druce, J.; de Vries, J.J.C.; Kikkert, M.; Barcena, M.; et al. SARS-coronavirus-2 replication in Vero E6 cells: replication kinetics, rapid adaptation and cytopathology. J Gen Virol 2020, 101, 925–940. [Google Scholar] [CrossRef]
- McBride, R.; van Zyl, M.; Fielding, B.C. The coronavirus nucleocapsid is a multifunctional protein. Viruses 2014, 6, 2991–3018. [Google Scholar] [CrossRef]
- Riddell, S.; Goldie, S.; Hill, A.; Eagles, D.; Drew, T.W. The effect of temperature on persistence of SARS-CoV-2 on common surfaces. Virol J 2020, 17, 145. [Google Scholar] [CrossRef]
- Wei, C.; Shan, K.J.; Wang, W.; Zhang, S.; Huan, Q.; Qian, W. Evidence for a mouse origin of the SARS-CoV-2 Omicron variant. J Genet Genomics 2021, 48, 1111–1121. [Google Scholar] [CrossRef] [PubMed]
- Schmid-Holmes, S.; Drickamer, L.C.; Robinson, A.S.; Gillie, L.L. Burrows and burrow-cleaning behavior of house mice (Mus musculus domesticus). The American Midland Naturalist 2001, 146, 53–62. [Google Scholar] [CrossRef]
| Year of First Publication | Shell Disorder Model | Details |
|---|---|---|
| 2008 | Parent Viral Shape-shifter Model | Abnormally huge levels of disorder were found at the outer shell of many HIV-1 variants and may sexually transmitted viruses such as HSV and HCV. This could account for the lack of an effective HIV vaccine. |
| 2012 | CoV Transmission SDM | Levels of fecal-oral and respiratory CoV transmission is predicted by levels of shell disorder. |
| 2015 | Virulence-inner Shell Disorder Model | High correlations between the inner shell and virulence of a variety of viruses have been detected. |
| Coronavirus | Se-quence Similari-ty M (%) |
PIDM (%) |
Accession: UniProt(U) GenBank(G) |
Sequence Similarity N (%) |
PIDN (%) |
Accession UniProt(U) GenBank(G |
|---|---|---|---|---|---|---|
|
SARS-CoV-1 Civet-SARS-CoV |
90.5 90.1 |
8.6 8.6 |
P59596(U) Q3ZTE9(U) |
90.5 90.01 |
50.2 49.1 |
P59595(U) Q3ZTE4(U) |
| COVID-Related Bat-CoV RaTG13 Laotian Bat-CoV [Banal-52] [Banal-103] [Banal-236] |
99.6 98,7 98.7 99.1 |
6.0+0.2 4.1 6.3 5.9 4.1 |
QHR63303(G) UAY13220(G) UAY13232(G) UAY13256(G) |
99.1 99.3 99.1 99.3 |
48.3+0.2 48.2 48.5 48.5 |
QHR63308(G) UAY13225.1 UAY13257.1 UAY1326.1 |
| Pangolin-CoV 2019 2018 2017*** |
98.2 97.7 98.2 |
5.6+0.9a 6.3 4.5 5.9 |
QIG55948(G) QIQ54051(G) QIA48617(G) |
98 93.8 94 93.32 |
46.6+1.6a48.7 46.3 44.8 46.5 |
QIG55953(G) QIQ54056(G) QIA48630(G) QIA48656(G) |
| SARS-CoV-2 [Wuhan] [Delta1] [Delta2[ [Omicron]** |
100 99.1 99.1 98.7 |
5.9 5.9 5.9 5.7+0.4 |
YP009724393(G) QUX81285(G) QUX81285(G) |
100 99.3 99.1 98.6 |
48.2 46.8 47.5 44.5+0.4 |
YP009724397(G) QYM89997(G) QYM89845(G) |
| Omicron BA.1.44 XBB.1.16 |
98.7 99.1 |
5.4 5.9 |
UFO69282(G) WIL50320(G) |
98.6 98.3 |
44.7 44.2 |
UFO69287(G) WIL50325(G) |
| Bat-CoV RATG13 Bat 512 HKU3 HKU4 HKU5 |
99.6 35.5 91 42.7 44.7 |
11+15a 4.1 15.3 7.7 16.4 11.8 |
QHR63303(G) Q0Q463(U) Q3LZX9(U) A3EXA0(U) A3EXD6(U) |
99.1 29.4 89.6 51.1 47.9 |
47.7+0.9a 48.5 46.5 48 48.5 47.1 |
QHR63308(G) Q0Q462(U) Q3LZX4(U) A3EXA1(U) A3EXD7(U) |
| Virus/Isolate | PIDN | Non-Attenuation/Aggressiveness |
| SARS-CoV-1 | 50.00% | +++ |
| BANAL-236 | 48.5% | ++ |
| Pang2019 | 48.5 | ++ |
| Wuhan-Hu-1 | 48.20% | + |
| XBB.1.16 | 44.5 | - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




