Submitted:
06 January 2026
Posted:
08 January 2026
You are already at the latest version
Abstract
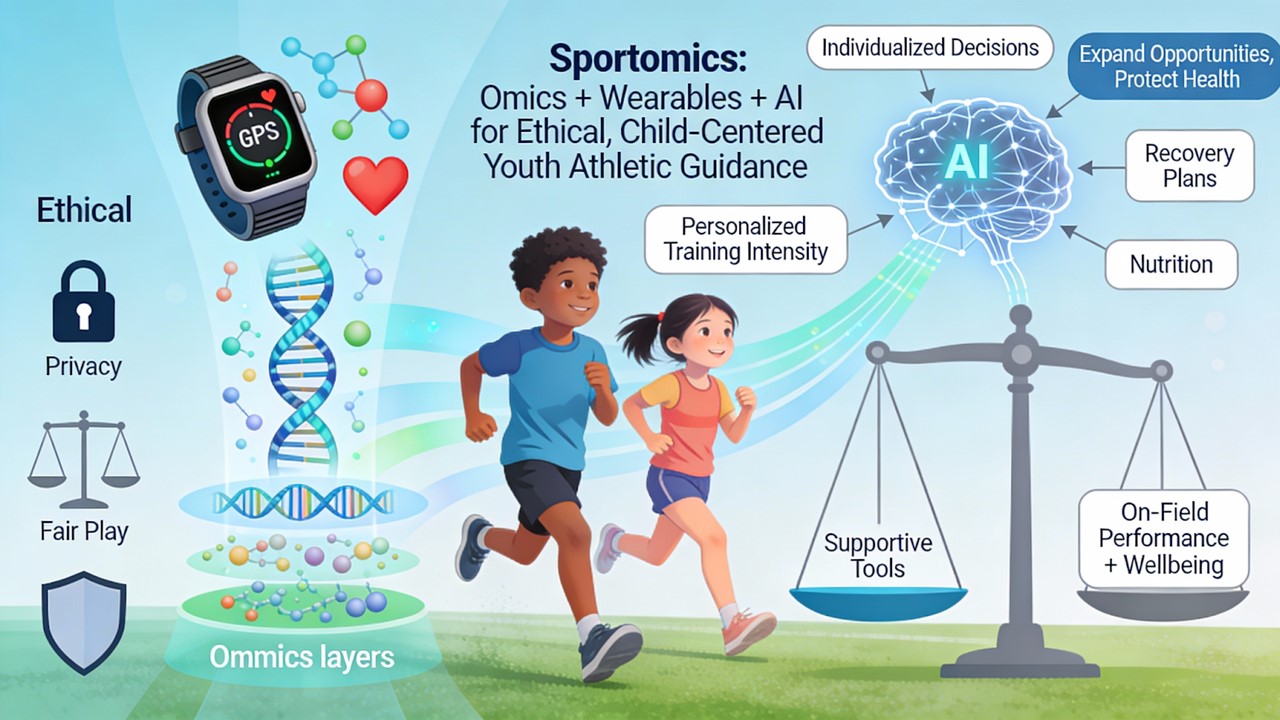

Keywords:
1. Introduction
1.2. Overview of Recent Advancements in Genomics
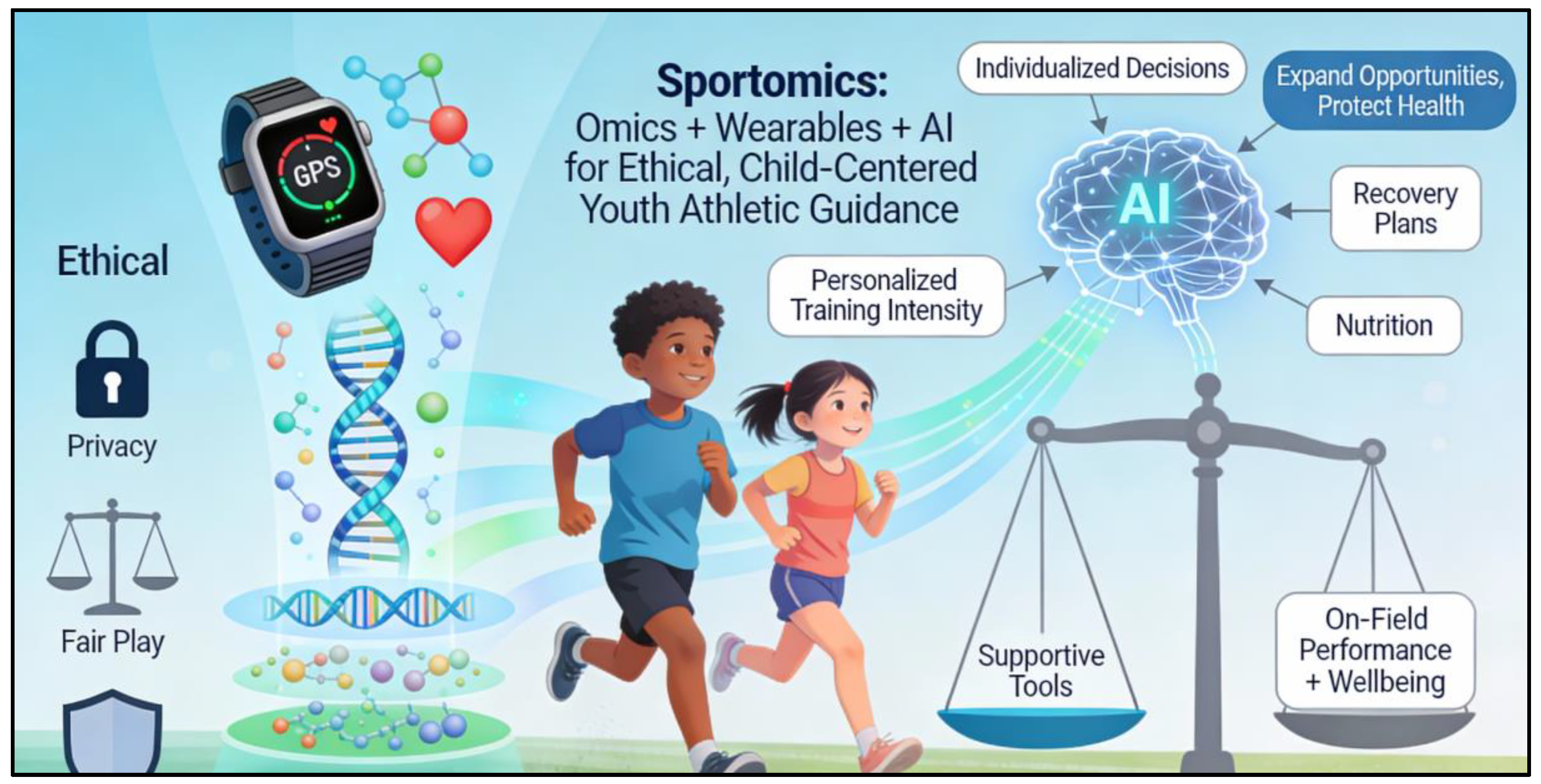
1.3. Relation Between Sportomics and Digital Monitoring Devices
1.4. Ethical Guidelines Responsible for Athletic Empowerment
2. Key Molecular Techniques and Their Relevance
2.1. Genomics: Mapping the Blueprint of Potential
- ACE (Angiotensin-Converting Enzyme) I/D polymorphism affects cardiovascular efficiency and endurance capacity (Ahmetov & Rogozkin, 2020; Rodríguez-Larrad et al., 2020).
- ACTN3 R577X variant influences the balance between fast- and slow-twitch muscle fibers, shaping sprint versus endurance predispositions (El Ouali et al., 2024; Ahmetov et al., 2021).
- COL1A1 and COL5A1 variants relate to collagen structure and ligament strength, indicating susceptibility to tendon or ligament injury (Rodríguez-Romo & Peñalvo, 2022; Kwasniewski et al., 2024).
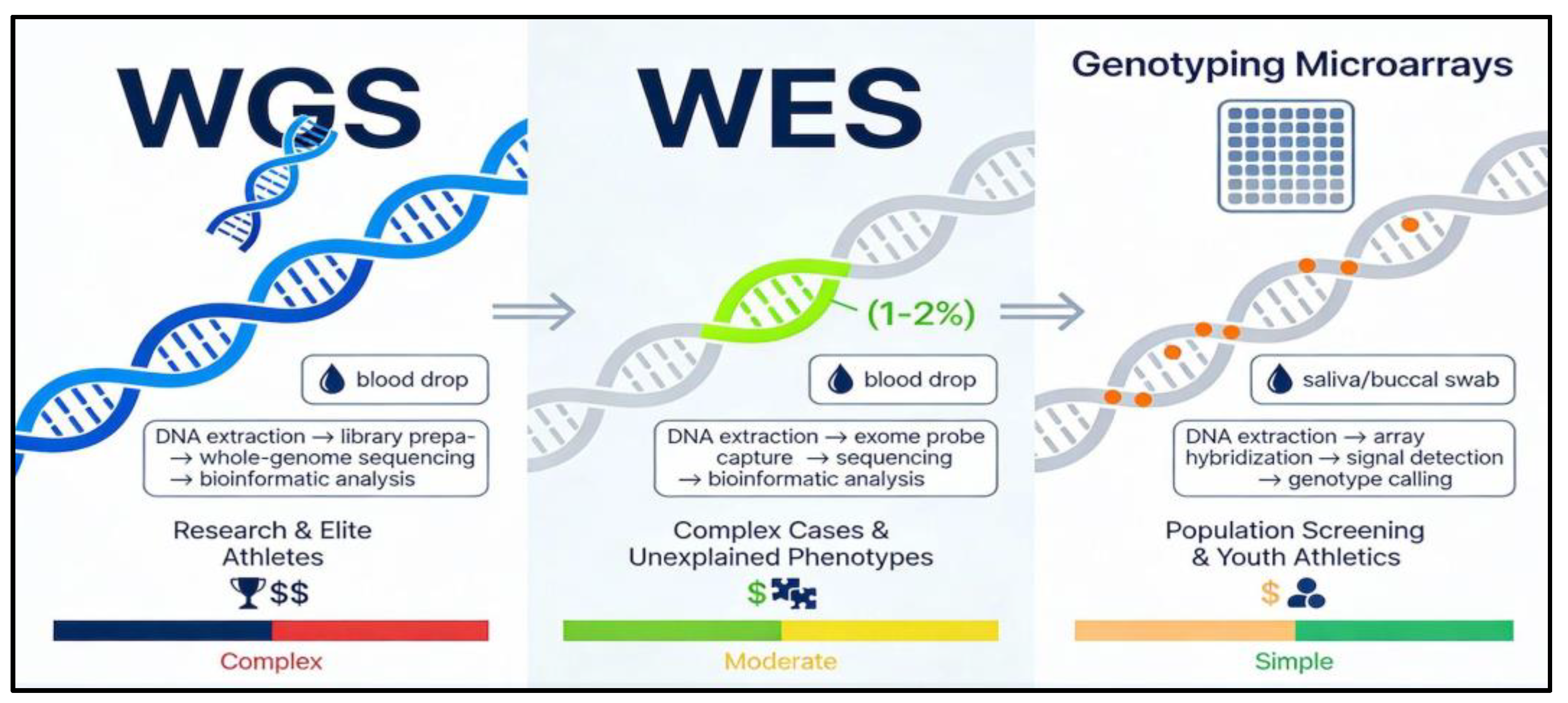
- Whole Exome Sequencing (WES)
- 2.
- Whole Genome Sequencing (WGS)
- 3.
- Genotyping Microarrays
- Endurance vs power tendency: Variants in genes such as ACE and ACTN3 do not decide which sport a child “must” play, but they can indicate whether the body is naturally more efficient at long-duration, oxygen-based work or at short, explosive efforts. This helps in deciding how much of the weekly training load should be endurance-focused versus power-focused, not in excluding any sport (Konopka MJ et al.2023).
- Soft-tissue and ligament vulnerability: Variants in collagen-related genes (for example COL1A1, COL5A1) may signal that an athlete’s tendons and ligaments need more gradual loading, longer warm-ups, and a stronger focus on technique and neuromuscular control to prevent injuries ( Bojarczuk A; 2024).
- Recovery profile: Markers linked to inflammation and oxidative stress regulation (such as IL6 or TNF-α polymorphisms) can help explain why some children “bounce back” quickly after hard sessions, while others feel sore and fatigued longer. Coaches can respond by adjusting recovery days, sleep education, and nutrition strategies for each child (Malsagova et al.,2024).
- The child’s best interest must come first (health and enjoyment before performance).
- Results should always be explained in language that avoids labels such as “talented” or “untalented”, which can harm self-confidence.
- Data must be stored securely, with parental consent and the possibility for the young person to withdraw their data when they are older.
2.2. Proteomics: Understanding Muscle Recovery and Fatigue
- How much muscle damage did yesterday’s session cause?
- Is the athlete’s inflammation at a normal adaptive level or trending towards overuse?
- Are stress hormones balanced, or is the child showing signs of chronic overload?
2.3. Metabolomics: Tracking Energy and Nutrition Dynamics
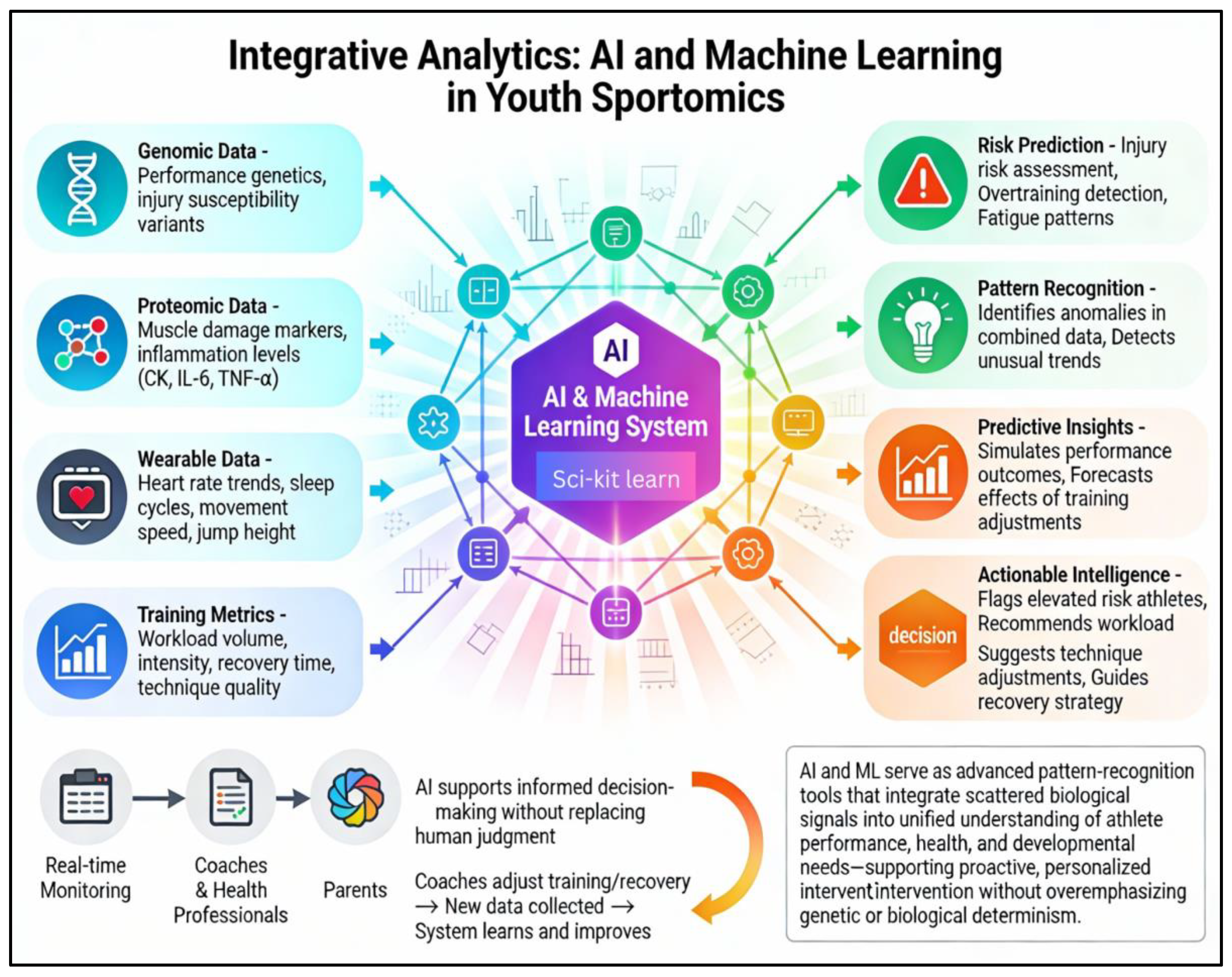
3. Integrative Analytics: The Role of AI and Machine Learning
3.1. AI-Enabled Innovations in Training Optimization and Injury Prevention
4. Building Trust: Making AI a Reliable Partner in Sports
5. Guarding Privacy: Ethics First in Sportomics
6. Precision Matters: Ensuring Data Accuracy and Reliability
- Health protection and injury prevention over selection or deselection
- Non-directive counselling, where results are explained neutrally and options are kept open
- Re-consent when the athlete reaches adulthood, including the possibility to delete their data.
7. The Future of Molecular Insights in Sport
8. Discussion
9. Conclusion
Authors Declaration
Funding
Ethical approval
Consent to participate
Consent to publication
Acknowledgments
Conflicts of interest
Abbreviations
| ACE– | Angiotensin-Converting Enzyme |
| ACTN3– | Alpha-Actinin-3 |
| AI– | Artificial Intelligence |
| COL5A1– | Collagen Type V Alpha 1 |
| DNA– | Deoxyribonucleic Acid |
| EMG– | Electromyography (implied in biomechanical assessments) |
| GPS– | Global Positioning System (used in wearables for athlete tracking) |
| HRV– | Heart Rate Variability (component of fatigue monitoring) |
| ML– | Machine Learning |
| MRI– | Magnetic Resonance Imaging (clinical imaging context) |
| NGS– | Next-Generation Sequencing (used in genomics context) |
| RTP– | Return to Play |
| VO₂max– | Maximal Oxygen Uptake (implied in endurance assessment) |
| OMICs Terminology: | |
| Genomics– | Study of genes and genetic variation |
| Transcriptomics– | Study of RNA expression |
| Proteomics– | Study of protein structure and function |
| Metabolomics– | Study of metabolic pathways and metabolites |
| Microbiomics / Microbiome Profiling– | Study of gut microbial communities |
References
- Ahmetov, I. I.; Fedotovskaya, O. N. Current progress in sports genomics. Exercise and Sport Sciences Reviews 2020, 48(4), 191–199. [Google Scholar] [CrossRef]
- Ahmetov, I. I.; Kulemin, N. A.; Popov, D. V. Genomic predictors of athlete status and response to training. European Journal of Sport Science 2021, 21(7), 1–12. [Google Scholar] [CrossRef]
- Ahmetov, I. I.; Rogozkin, V. Genetic profile of endurance athletes and strength athletes. Journal of Sports Sciences 2020, 38(8), 987–997. [Google Scholar] [CrossRef]
- Baca, A.; Kornfeind, P. Motion analysis in sports: A critical review. Sensors 2021, 21(7), 2121. [Google Scholar] [CrossRef]
- Babu, M.; et al. Wearable devices: Implications for precision medicine and population health. Annual Review of Medicine 2024, 75, 1–20. [Google Scholar] [CrossRef]
- Bassini, A.; Bragazzi, N. L.; Cameron-Smith, D. Sportomics method to assess acute-phase proteins and metabolome changes in athletes. Scientific Reports 2022, 12, 23300. [Google Scholar] [CrossRef]
- Bergeron, M. F. Youth athletic development: Managing training load in children and adolescents. Journal of Strength and Conditioning Research 2018, 32(7), 2001–2011. [Google Scholar]
- Bragazzi, N. L.; Bassini, A. Sportomics overview: Metabolic signatures of exercise. European Review 2020, 28(3), 1–12. [Google Scholar] [CrossRef]
- Buchheit, M. Monitoring training responses in elite athletes using new technologies. Sports Medicine 2020, 50(6), 1025–1042. [Google Scholar]
- Cameron-Smith, D.; et al. Protein biomarkers to monitor training adaptation and overreaching. Proteomics Clinical Applications 2021, 15(6), e2000030. [Google Scholar] [CrossRef]
- Cavalli, G.; Meijer, A. H.; Reinders, M. J. T. Ethical considerations in sports genomics. Nature Genetics 2023, 55(2), 141–150. [Google Scholar] [CrossRef]
- Clarke, A.; et al. Data governance, privacy and ethics in youth sports genomics. Journal of Medical Ethics 2022, 48(9), 600–607. [Google Scholar] [CrossRef]
- Claudino, J. G.; et al. Injury risk assessment and performance prediction using AI. Sports Medicine – Open 2020, 6, 61. [Google Scholar] [CrossRef]
- Claudino, J. G.; et al. Machine learning in sports biomechanics. British Journal of Sports Medicine 2021, 55(19), 1099–1108. [Google Scholar] [CrossRef]
- Claudino, J. G.; et al. Technologies for monitoring athlete workload. Sports Medicine 2022, 52(8), 1715–1734. [Google Scholar] [CrossRef]
- de la Rosa, A.; et al. Multi-omics integration for sports injury biomarkers. OMICS 2022, 26(5), 274–289. [Google Scholar] [CrossRef]
- El Ouali, E. M.; et al. ACTN3 R577X genotype and athletic status. Sports Medicine – Open 2024, 10, 24. [Google Scholar] [CrossRef]
- El-Sayed, M. S.; Bouhlel, E. Metabolomic assessment of training load. Journal of Applied Physiology 2020, 128(4), 765–775. [Google Scholar] [CrossRef]
- Fernández-García, M.; et al. Predictive modelling of injury risk with wearables. Sensors 2023, 23(4), 1789. [Google Scholar] [CrossRef]
- Freitas, V. H.; Nakamura, F. Y.; Milos, N. Integrating molecular biomarkers with training data. Sports Medicine 2021, 51(4), 677–694. [Google Scholar]
- Fuller, J. T.; Bellenger, C. R.; Buckley, J. D. Wearable sensors in youth sports. Frontiers in Sports and Active Living 4 2022, 879123. [Google Scholar] [CrossRef]
- Ghosh, I.; et al. Sports analytics review. WIREs Data Mining and Knowledge Discovery 2023, 13, e1496. [Google Scholar] [CrossRef]
- Gibson, E.; Adams, M.; Chin, J. Protecting minors in genomic research. American Journal of Bioethics 2021, 21(3), 40–52. [Google Scholar] [CrossRef]
- Ginevičienė, V.; et al. Perspectives in sports genomics. Biomedicines 2022, 10(2), 298. [Google Scholar] [CrossRef]
- Goes, F. R.; et al. Big data in tactical soccer analysis. European Journal of Sport Science 2021, 21(4), 481–496. [Google Scholar] [CrossRef]
- Halperin, I. Psychological readiness in training prescription. Sports Medicine 2018, 48(11), 2551–2558. [Google Scholar]
- Hamilton, B.; Hopkins, W. Statistical models for athlete development. Medicine & Science in Sports & Exercise 2021, 53(12), 2547–2558. [Google Scholar] [CrossRef]
- Horvath, S.; Raj, K. DNA methylation biomarkers in athletic performance. Sports Medicine 2020, 50(1), 131–145. [Google Scholar] [CrossRef]
- Huygens, W.; et al. Precision performance frameworks. International Journal of Sports Medicine 2022, 43(10), 809–818. [Google Scholar] [CrossRef]
- Jia, Y.; et al. Deep learning in sports analytics. Frontiers in Computer Science 2025, 3, 123456. [Google Scholar] [CrossRef]
- Khosravi, P.; et al. Explainable AI in sports. IEEE Access 2022, 10, 45712–45727. [Google Scholar] [CrossRef]
- Kobayashi, H.; Koga, H.; Yokoyama, K. Advances in MRI for musculoskeletal injuries. British Journal of Sports Medicine 2021, 55(9), 468–474. [Google Scholar]
- Kwasniewski, M.; et al. Combining genomics & biomechanics. Journal of Biomechanics 2024, 156, 111111. [Google Scholar] [CrossRef]
- Lemmink, K. A. P. M.; et al. Longitudinal monitoring in youth athletes. Journal of Sports Sciences 2021, 39(9), 1012–1024. [Google Scholar] [CrossRef]
- Li, R.; et al. ML for youth talent identification. International Journal of Sports Science & Coaching 2020, 15(4), 623–642. [Google Scholar] [CrossRef]
- Li, W.; Zhou, D.; Zhang, X. Deep learning for athlete tracking. Computer Vision in Sports 2023, 12(3), 101–120. [Google Scholar] [CrossRef]
- Li, X.; et al. ML for sports injury prediction. IEEE TNSRE 2021, 29, 1–14. [Google Scholar] [CrossRef]
- Lloyd, R. S.; Oliver, J. L. Youth physical development model. Strength & Conditioning Journal 2019, 41(3), 2–17. [Google Scholar] [CrossRef]
- Martin, D.; et al. ML pipelines for sports genomics. Bioinformatics 2024, 40(12), 3450–3462. [Google Scholar] [CrossRef]
- Mehta, K.; Kulkarni, A.; Raj, A. Sleep quality & adolescent athletes. Sports Medicine – Open 2023, 9(1), 12. [Google Scholar] [CrossRef]
- Mittelstadt, B. D. Principles of data ethics. Big Data & Society 2019, 6(2). [Google Scholar] [CrossRef]
- Muniz-Santos, R.; et al. Metabolomics in elite athlete monitoring. Journal of Sports Medicine and Physical Fitness 2021, 61(9), 1243–1254. [Google Scholar] [CrossRef]
- Muniz-Santos, R.; et al. Metabolic responses in Olympic athletes. Nutrients 2024, 17(11), 1924. [Google Scholar] [CrossRef]
- Muro-De-La-Herran, A.; García-Zapirain, B.; Méndez-Zorrilla, A. Gait analysis for performance. Applied Sciences 2022, 12(6), 3204. [Google Scholar] [CrossRef]
- Nasb, M.; et al. Precision exercise. Precision Exercise Medicine 2024, 1(1), 45–65. [Google Scholar] [CrossRef]
- Nashef, R.; Smith, J. Microbiome & exercise. Trends in Microbiology 2023, 31(2), 122–136. [Google Scholar] [CrossRef]
- Nascimento, D. C.; et al. Biomarkers of training load. International Journal of Sports Physiology and Performance 2022, 17(6), 825–836. [Google Scholar] [CrossRef]
- Nassis, G. P.; et al. Big data in sport. Journal of Sports Sciences 2020, 38(23), 1–14. [Google Scholar] [CrossRef]
- O’Connell, J.; et al. Longitudinal multimodal monitoring. Pediatric Exercise Science 2021, 33(2), 103–118. [Google Scholar] [CrossRef]
- Peake, J. M.; et al. Integrative omics in sport. Journal of Proteome Research 2020, 19(5), 2020–2036. [Google Scholar] [CrossRef]
- Pereira, T.; Ribeiro, J.; Oliveira, J. Wearable technology in sports. Journal of Sports Sciences 2020, 38(17), 2018–2027. [Google Scholar] [CrossRef]
- Philip, A.; Bradley, J.; Wilson, F. Biomechanical risk factors in youth athletes. Sports Medicine 2023, 53(2), 215–230. [Google Scholar]
- Pietraszewski, P.; et al. AI in sports analytics. Applied Sciences 2025, 15, 7254. [Google Scholar] [CrossRef]
- Piette, M.; et al. Wearables & AI for youth athletes. Frontiers in Sports and Active Living 2022, 4, 800512. [Google Scholar] [CrossRef]
- Powers, S. K.; Jackson, M. J. Exercise-induced oxidative stress. Free Radical Biology & Medicine 2020, 146, 1–13. [Google Scholar] [CrossRef]
- Roberts, T.; et al. AI for rehabilitation. npj Digital Medicine 2023, 6, 94. [Google Scholar] [CrossRef]
- Rodríguez-Larrad, A.; et al. Genetic testing for sports performance. British Journal of Sports Medicine 2020, 54(12), 694–700. [Google Scholar] [CrossRef]
- Rodríguez-Matoso, D.; et al. Muscle damage and recovery. Frontiers in Physiology 2021, 12, 657430. [Google Scholar] [CrossRef]
- Rodríguez-Romo, G.; Peñalvo, J. L. Precision nutrition & omics. Nutrition Reviews 2022, 80(6), 1234–1249. [Google Scholar] [CrossRef]
- Saxon, L.; Hayes, M.; Williams, S. Digital privacy in youth athletes. Journal of Athletic Training 2021, 56(5), 532–540. [Google Scholar] [CrossRef]
- Sellami, M.; Bragazzi, N. L.; Chamari, K.; Franchini, E. Molecular big data in sports sciences. Frontiers in Molecular Biosciences 2021, 8, 815410. [Google Scholar] [CrossRef]
- Sellami, M.; Bragazzi, N. L. Gene–environment interactions in sportomics. Genes 2021, 12(3), 389. [Google Scholar] [CrossRef]
- Sellami, M.; et al. Multi-omics approaches in sport. International Journal of Sports Physiology and Performance 2020, 15(9), 1252–1266. [Google Scholar] [CrossRef]
- Stokes, K. A.; Jones, B.; Green, B. S. Human interpretation in data analytics. BMJ Open Sport & Exercise Medicine 2020, 6(1), e000775. [Google Scholar] [CrossRef]
- Taddeo, M.; Floridi, L. Ethics of explainable AI. Philosophy & Technology 2018, 31(4), 611–627. [Google Scholar] [CrossRef]
- Thorpe, R. T.; Atkinson, G.; Drust, B. AI in youth sports decision-making. International Journal of Sports Science & Coaching 2022, 17(4), 874–885. [Google Scholar]
- Williamson, P.; Keller, J. AI ethics in sports. Ethics and Information Technology 2023, 25, 1–18. [Google Scholar] [CrossRef]
- Zand, A.; et al. Precision sports medicine. Journal of Personalized Medicine 2024, 14(3), 412. [Google Scholar] [CrossRef]
- Ziemann, E.; et al. Epigenetic markers & youth athlete development. European Journal of Applied Physiology 2020, 120(11), 2501–2515. [Google Scholar] [CrossRef]
- Musat, C. L.; Mereuta, C.; Nechita, A.; Tutunaru, D.; Voipan, A. E.; Voipan, D.; Mereuta, E.; Gurau, T. V.; Gurău, G.; Nechita, L. C. Diagnostic applications of AI in sports: A comprehensive review of injury risk prediction methods. Diagnostics (Basel) 2024, 14(22), 2516. [Google Scholar] [CrossRef]
- Collier, R. Testing the ethics of genetic testing in sports. CMAJ 2011, 184(1), E45–E46. [Google Scholar] [CrossRef] [PubMed]
- Camporesi, S.; McNamee, M. J. Ethics, genetic testing, and athletic talent: Children’s best interests, and the right to an open (athletic) future. Physiological Genomics 2016, 48(3), 191–195. [Google Scholar] [CrossRef]
- Plaza-Diaz, J.; Izquierdo, D.; Torres-Martos, Á.; Baig, A. T.; Aguilera, C. M.; Ruiz-Ojeda, F. J. Impact of physical activity and exercise on the epigenome in skeletal muscle and effects on systemic metabolism. Biomedicines 2022, 10(1), 126. [Google Scholar] [CrossRef]
- Malsagova, K. A.; Kopylov, A. T.; Stepanov, A. A.; Kulikova, L. I.; Izotov, A. A.; Yurku, K. A.; Balakin, E. I.; Pustovoyt, V. I.; Kaysheva, A. L. Metabolomic and proteomic profiling of athletes performing physical activity under hypoxic conditions. Sports (Basel) 2024, 12(3), 72. [Google Scholar] [CrossRef]
- Vlahovich, N.; Fricker, P. A.; Brown, M. A.; Hughes, D. Ethics of genetic testing and research in sport: A position statement from the Australian Institute of Sport. British Journal of Sports Medicine 2017, 51(1), 5–11. [Google Scholar] [CrossRef]
- Kim, J.-H.; Kim, J.; Kang, H.; Youn, B.-Y. Kim, J.-H., Kim, J., Kang, H., & Youn, B.-Y. (in press). Ethical implications of artificial intelligence in sport: A systematic scoping review. [Journal Name], [Article Number/Page Range]. [CrossRef] [PubMed]
- Wang, P.; Wang, A.; Wang, S. Integrating multimodal AI technologies for sports injury prediction and rehabilitation: Systematic review. Journal of Human Sport and Exercise 2025, *21*(1), 22–37. [Google Scholar] [CrossRef]
- Konopka, M. J.; Sperlich, B.; Rietjens, G.; Zeegers, M. P. Genetics and athletic performance: A systematic SWOT analysis of non-systematic reviews. Frontiers in Genetics 2023, 14, 1232987. [Google Scholar] [CrossRef]
- Musat, C. L.; Mereuta, C.; Nechita, A.; Tutunaru, D.; Voipan, A. E.; Voipan, D.; Mereuta, E.; Gurau, T. V.; Gurău, G.; Nechita, L. C. Diagnostic applications of AI in sports: A comprehensive review of injury risk prediction methods. Diagnostics 2024, *14*(22), 2516. [Google Scholar] [CrossRef]
- Harshman, S. W.; Pitsch, R. L.; Smith, Z. K.; O’Connor, M. L.; Geier, B. A.; Qualley, A. V.; Schaeublin, N. M.; Fischer, M. V.; Eckerle, J. J.; Strang, A. J.; Martin, J. A. The proteomic and metabolomic characterization of exercise-induced sweat for human performance monitoring: A pilot investigation. PLOS ONE 2018, *13*(11), e0203133. [Google Scholar] [CrossRef]
- Bojarczuk, A. Ethical aspects of human genome research in sports—A narrative review. Genes (Basel) 2024, 15(9), 1216. [Google Scholar] [CrossRef] [PubMed]
- Indian Council of Medical Research. (2018). National ethical guidelines for biomedical and health research involving human participants (2017): Handbook. https://naitik.gov.in/DHR/resources/app_srv/DHR/global/pdf/downloads/Handbook_on_ICMR_Ethical_Guidelines.pdf.
- Boulygina, E. A.; Borisov, O. V.; Valeeva, E. V.; Semenova, E. A.; Kostryukova, E. S.; Kulemin, N. A.; Larin, A. K.; Nabiullina, R. M.; Mavliev, F. A.; Akhatov, A. M.; Andryushchenko, O. N.; Andryushchenko, L. B.; Zmijewski, P.; Generozov, E. V.; Ahmetov, I. I. Whole genome sequencing of elite athletes. Biology of Sport 2020, 37(3), 295–304. [Google Scholar] [CrossRef] [PubMed]
| Layer | What it measures | What it can tell coaches and parents | Typical sample |
|---|---|---|---|
| Genomics | DNA variants that are inherited and stable | Tendencies for endurance vs power, injury susceptibility, recovery capacity; helps tailor training focus and identify who needs extra joint/tendon protection | Blood, saliva, buccal swab |
| Proteomics | Proteins related to muscle damage, inflammation, and repair | Current muscle stress, level of inflammation, whether recovery is adequate; warns when a child is close to overreaching or overtraining | Blood |
| Metabolomics | Small molecules from energy and nutrient metabolism | How effectively a child uses carbohydrates, fats, and amino acids; hydration status; metabolic fatigue during or after hard blocks | Blood, urine |
| Feature | Whole Exome Sequencing (WES) | Whole Genome Sequencing (WGS) | Genotyping panels (microarray) |
|---|---|---|---|
| Coverage | All protein-coding regions | Almost entire genome | Selected known variants |
| Typical use in sport | Investigation of complex or unexplained cases; advanced profiling | Research and elite-level mechanistic studies | Screening for common performance and injury markers |
| Cost and complexity | Moderate, requires specialist interpretation | Highest, requires advanced infrastructure and ethics oversight | Lower, simpler interpretation |
| Suitability in children | Case-by-case, ideally under research/clinical protocol | Usually research only; not routine for youth athletics | Appropriate with counselling and clear limits on use |
| Gene | Polymorphism | Associated Trait | Scientific Findings (Summary) | Supporting Studies |
|---|---|---|---|---|
| ACE (Angiotensin-Converting Enzyme) | I/D | Endurance (I allele), Power (D allele) | I allele linked to improved oxygen uptake and endurance; D allele associated with power and sprint performance. | Montgomery et al., 1998 (Nature); Nazarov et al., 2001 |
| ACTN3 (Alpha-Actinin-3) | R577X | Sprint ability, fast-twitch muscle performance | R allele associated with power and speed; XX genotype linked to lower fast-twitch fiber efficiency but better endurance adaptation. | Yang et al., 2003 (AJHG); Eynon et al., 2013 |
| COL5A1 | rs12722 (C/T) | Flexibility, reduced injury risk | T allele associated with improved flexibility and lower risk of tendon injuries such as Achilles tendinopathy. | Collins & Raleigh, 2009; September et al., 2009 |
| COL1A1 | Sp1 (G/T) | Ligament strength, injury resistance | T allele associated with increased collagen type I production and reduced ligament injury susceptibility. | Posthumus et al., 2009 |
| PPARGC1A (PGC-1α) | Gly482Ser | Aerobic capacity, mitochondrial efficiency | Gly allele linked to higher VO₂max and improved aerobic metabolism. | Lucia et al., 2005 |
| IL6 | -174 G/C | Inflammation regulation, recovery | C allele associated with lower inflammation and better recovery after intensive training. | Fishman et al., 1998; Yamin et al., 2008 |
| TNF-α | -308 G/A | Muscle damage, recovery rate | An allele linked to elevated inflammatory response and slower muscle recovery. | Saunders et al., 2006 |
| BDKRB2 (Bradykinin Receptor B2) | +9/−9 | Endurance performance | −9 variant associated with improved endurance and vascular adaptation. | Williams et al., 2004 |
| HIF1A | Pro582Ser | Adaptation to hypoxia, anaerobic performance | Ser allele enhances anaerobic power and adaptation to low-oxygen conditions. | Ahmetov et al., 2008 |
| MCT1 | A1470T | Lactate transport, high-intensity performance | T allele linked to improved lactate clearance and better high-intensity performance. | Cupeiro et al., 2012 |
| Marker | What it reflects for coaches | When to be cautious |
|---|---|---|
| Creatine kinase (CK) | Degree of muscle fibre disruption after training | If CK stays elevated across multiple tests, training loads may be too high or recovery insufficient |
| Myoglobin | Acute muscle damage | Very high values after repeated intense sessions warrant rest and technique review |
| IL-6, TNF-α | Systemic inflammation | Persistent elevation suggests inadequate recovery or early overuse risk |
| C-reactive protein (CRP / hsCRP) | General inflammatory load | High baseline levels may signal that the athlete is not fully recovered from previous blocks |
| Testosterone: cortisol ratio |
Balance between anabolic (building) and catabolic (breaking down) processes | A chronically reduced ratio in adolescents can indicate accumulated stress and need for modified training |
| Testing Category | Description | Sample Type | Key Features |
|---|---|---|---|
| Genomic Panels (DNA-based athletic profiling) | Tests for performance-related polymorphisms (e.g., ACTN3, ACE, COL5A1 variants for power/endurance/injury risk) | Saliva, buccal swab, blood | 20-40+ markers; reports on athletic traits, recovery, nutrition guidance |
| Sports Gene Genotyping (Single/Multi-gene) | Focused tests (e.g., ACTN3 R577X for fast-twitch muscle) | Blood, saliva | Single gene or small panels; quick turnaround |
| Integrated Wellness + Sports Genomics | Combines athletic traits with nutrition, injury risk, wellness | Saliva/blood | 40+ traits + nutrigenomics; personalized reports |
| Proteomics (Muscle/Recovery Markers) | Blood panels for CK, LDH, IL-6 (muscle damage, inflammation) | Blood serum/plasma | Clinical-grade; for overtraining assessment |
| Metabolomics (Energy/Nutrition) | Serum/urine analysis of metabolites (lactate, BCAAs) | Blood/urine | Training adaptation, fatigue biomarkers |
| Performance Physiology Labs (VO2max + Biochemical) | VO2max testing + lactate/blood markers | Blood + exercise testing | Integrated with wearables |
| Level | What it is (for coaches) | Main benefit in practice | Who might consider it | Key limitations / points to remember |
|---|---|---|---|---|
| DNA (Genomics) | Relatively fixed blueprint: variants in genes related to muscle type, cardiovascular function, connective tissue, inflammation | Helps tailor training focus (endurance vs power), identify children who may need slower load progression or extra injury-prevention work | Competitive youth athletes and academies seeking long-term planning; usually one-time testing with updates only if needed | Shows tendencies, not guarantees; must not be used to label or exclude; requires robust consent and privacy safeguards |
| RNA (Transcriptomics) | Snapshot of which genes are “switched on” in response to training or recovery | Helps researchers understand how a particular training block is affecting muscle adaptation at a molecular level | Mainly research settings and high-performance centres; can inform future guidelines | Not yet routine for day-to-day coaching; invasive, costlier, and interpretation is complex |
| Protein (Proteomics) | Markers of muscle damage, inflammation, and repair in blood | Indicates whether a training block is within the athlete’s current capacity or causing excessive stress; helps optimize rest days and deload weeks | Health professionals supporting youth teams; intensive training camps; return-to-sport monitoring after injury | Values fluctuate; must always be interpreted with symptoms and training load; over-testing may cause confusion and unnecessary anxiety |
| Integrated panels (multi-omics + wearables) | Combined picture using DNA + proteins + metabolites + heart rate, sleep, workload data | Allows individualized training and recovery plans; supports early detection of overuse risk; can inform nutrition strategies | Elite development programmes; research collaborations; selected high-commitment families | More data ≠ better decisions without expert interpretation; higher cost; strong data protection and ethical oversight required, especially for minors |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




