Submitted:
02 February 2026
Posted:
05 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Sample Collection, DNA Extraction, Library Preparation, and Sequencing
2.2. Alignment and Variant Calling
2.3. Functional Annotation and Gene Enrichment Analysis
2.4. Population Structure Analysis
2.5. Selective Sweep Analysis
3. Results
3.1. Read Quality, Mapping, and Depth Coverage
3.2. Variant Calling
3.2.1. Functional Annotation and Gene Enrichment Analysis
| Variant Type | Royal White Indels | White Dorper Indels |
|---|---|---|
| 3’ UTR truncation | 1 | 1 |
| 3’ UTR variant | 67,521 | 55,407 |
| 5’ UTR truncation | 1 | 3 |
| 5’ UTR variant | 21,424 | 17,480 |
| Bidirectional gene fusion | 78 | 68 |
| Conservative inframe deletion | 820 | 617 |
| Conservative inframe insertion | 654 | 479 |
| Disruptive inframe deletion | 1,473 | 1,098 |
| Disruptive inframe insertion | 671 | 509 |
| Downstream gene variant | 386,904 | 318,194 |
| Exon loss variant | 5 | 7 |
| Frameshift variant | 10,427 | 9,510 |
| Gene fusion | 59 | 30 |
| Intergenic region | 1,744,317 | 1,418,580 |
| Intragenic variant | 920 | 793 |
| Intron variant | 4,292,930 | 3,529,191 |
| Non-coding transcript exon variant | 22,460 | 19,116 |
| Non-coding transcript variant | 304 | 235 |
| Splice acceptor variant | 1,339 | 1,106 |
| Splice donor variant | 977 | 759 |
| Splice region variant | 12,640 | 10,121 |
| Start lost | 86 | 54 |
| Start retained variant | 24 | 17 |
| Stop gained | 135 | 127 |
| Stop lost | 64 | 56 |
| Stop retained variant | 10 | 10 |
| Transcript ablation | 1 | 2 |
| Upstream gene variant | 380,196 | 309,740 |
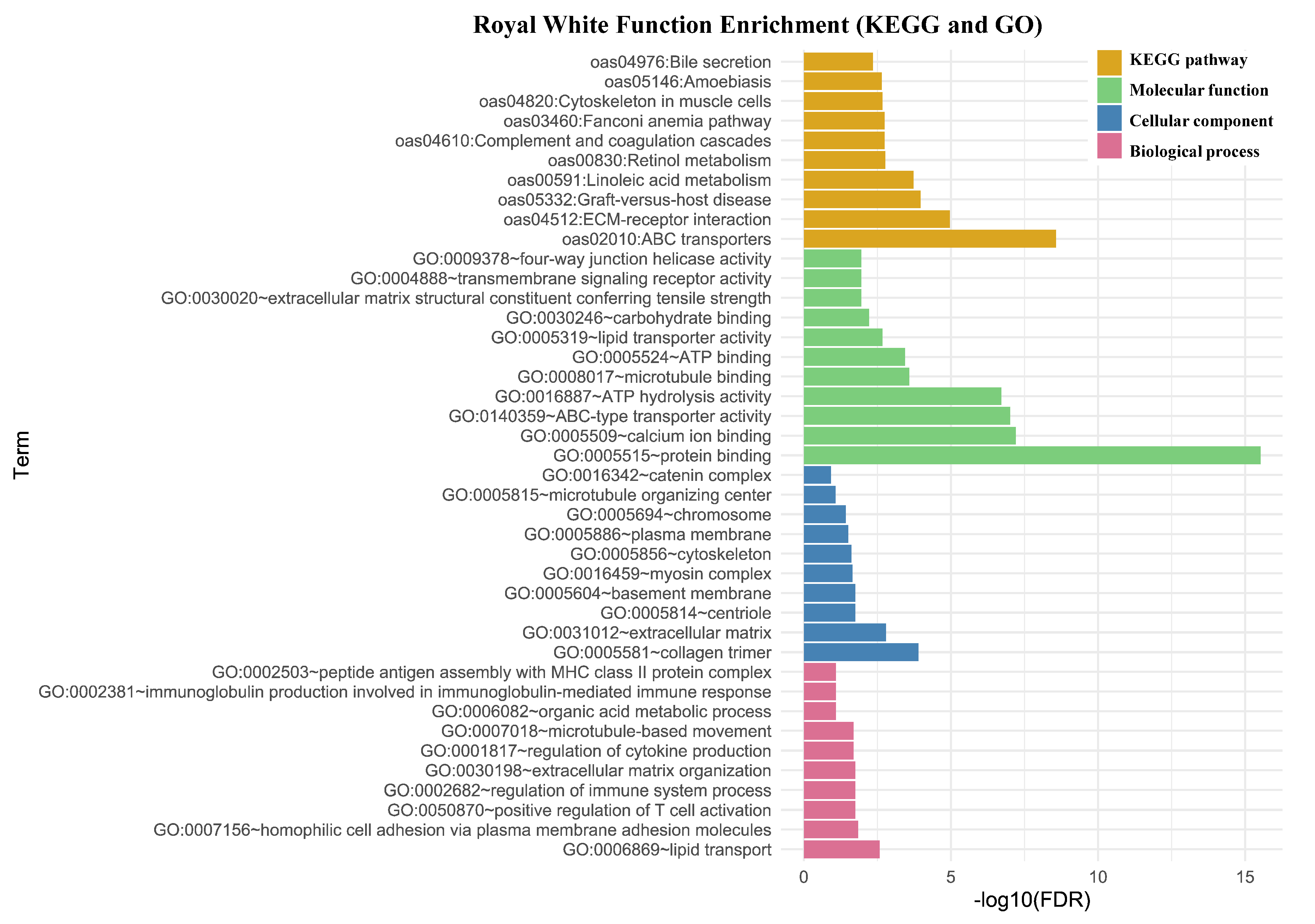
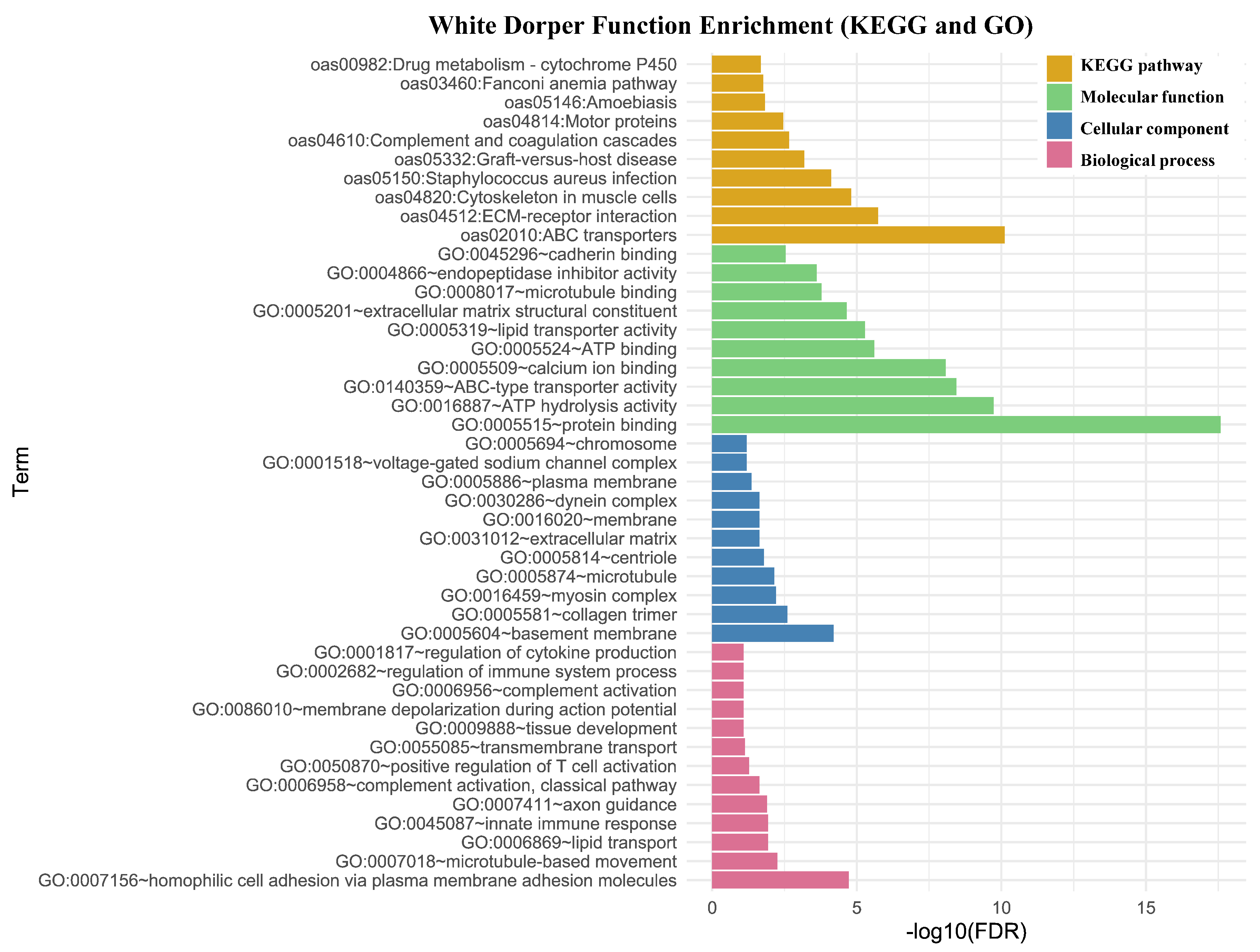
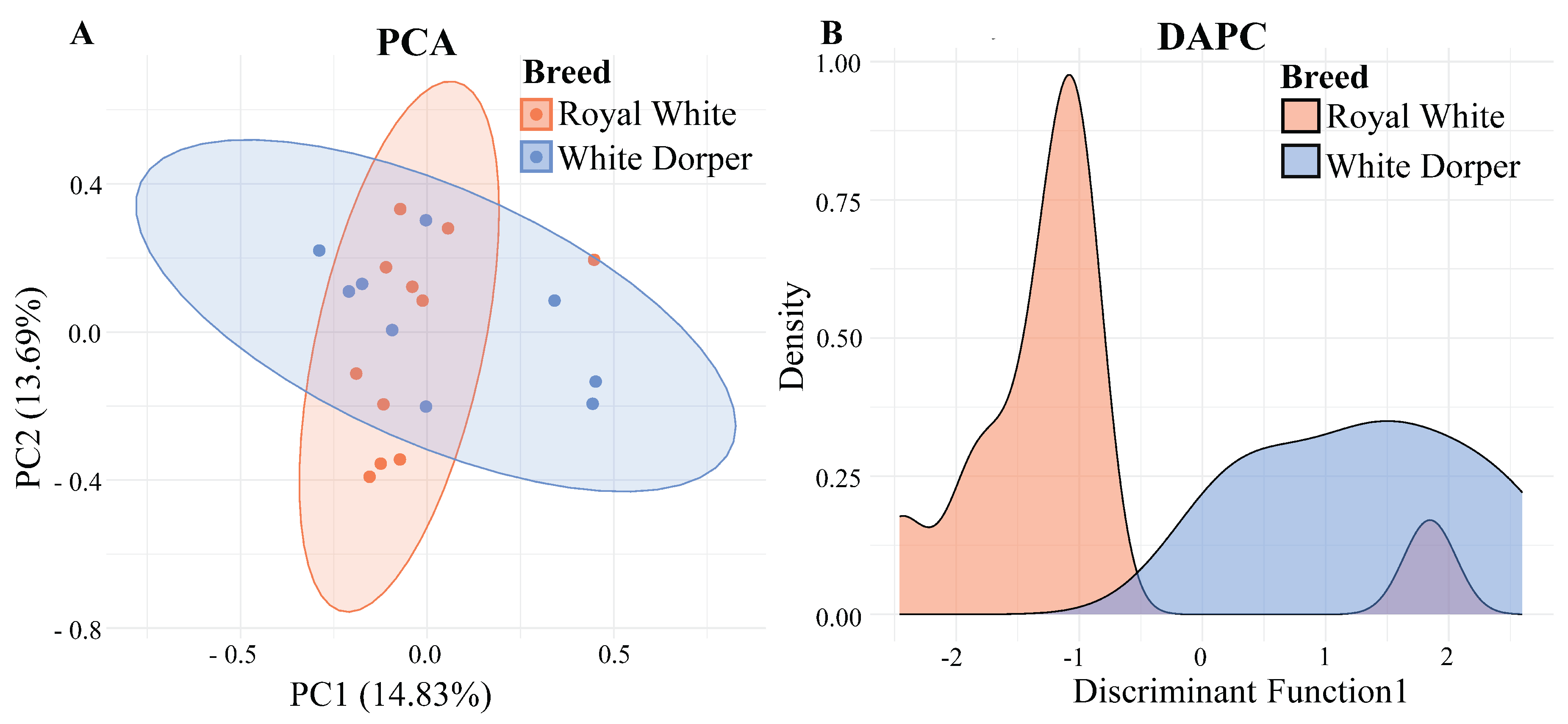
3.3. Population Structure Analysis
3.4. Selective Sweep Analysis
4. Discussion
4.1. Genomic Variant Characteristics
4.2. Functional Annotation and Enrichment
4.3. Candidate Genes Under Selection
4.3.1. Candidate Genes in Royal White Sheep
4.3.2. Candidate Genes in White Dorper Sheep
4.4. Limitations and Future Directions
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Food and Agriculture Organization of the United Nations. FAOSTAT: Live animals – sheep. https://www.fao.org/faostat/en/#data/QA, 2024. Accessed January 31, 2026.
- United States Department of Agriculture, National Agricultural Statistics Service. Sheep and Goats. https://downloads.usda.library.cornell.edu/usda-esmis/files/000000018/zk51xc07n/9593wq66x/shep0125.pdf?utm_source=chatgpt.com, 2025. Released January 2025, Accessed June 19, 2025.
- Royal White® Sheep. Royal White® Sheep – Official Breed Information. https://royalwhitesheep.biz/, 2024. Accessed June 19, 2025.
- Milne, C. The history of the Dorper sheep. Small Ruminant Research 2000, 36, 99–102. [CrossRef]
- Ojango, J.M.; Okpeku, M.; Osei-Amponsah, R.; Kugonza, D.R.; Mwai, O.; Changunda, M.G.; Olori, V.E. Dorper sheep in Africa: a review of their use and performance in different environments. CABI Reviews 2023.
- Wanjala, G.; Astuti, P.K.; Bagi, Z.; Kichamu, N.; Strausz, P.; Kusza, S. Assessing the genomics structure of Dorper and white Dorper variants, and Dorper populations in South Africa and Hungary. Biology 2023, 12, 386. [CrossRef]
- Gavojdian, D.; Budai, C.; Cziszter, L.T.; Csizmar, N.; Jávor, A.; Kusza, S. Reproduction efficiency and health traits in Dorper, White Dorper, and Tsigai sheep breeds under temperate European conditions. Asian-Australasian journal of animal sciences 2015, 28, 599. [CrossRef]
- Daetwyler, H.D.; Capitan, A.; Pausch, H.; Stothard, P.; Van Binsbergen, R.; Brøndum, R.F.; Liao, X.; Djari, A.; Rodriguez, S.C.; Grohs, C.; et al. Whole-genome sequencing of 234 bulls facilitates mapping of monogenic and complex traits in cattle. Nature genetics 2014, 46, 858–865. [CrossRef]
- Hayes, B.J.; Daetwyler, H.D.; Bowman, P.J.; Chamberlian, A.; Vander Jagt, C.; Capitan, A.; Pausch, H.; Stothard, P.; Liao, X.; Schrooten, C.; et al. Genomic prediction from whole genome sequence in livestock: the 1000 bull genomes project. In Proceedings of the 10th World Congress of Genetics Applied to Livestock Production. American Society of Animal Science, 2014.
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [CrossRef]
- Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv preprint arXiv:1303.3997 2013. [CrossRef]
- Van der Auwera, G.A.; O’Connor, B.D. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Current protocols in bioinformatics 2020, 43, 10–11. [CrossRef]
- Li, H. Statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 2011, 27, 2987–2993. [CrossRef]
- Chen, H.; Boutros, P. VennDiagram: Generate High-Resolution Venn and Euler Plots, 2011. R package version 1.7.3.
- Cingolani, P.; Platts, A.; Wang le, L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 2012, 6, 80–92. [CrossRef]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature protocols 2009, 4, 44–57. [CrossRef]
- Browning, B.L.; Zhou, Y.; Browning, S.R. Improving the accuracy and efficiency of identity-by-descent detection in population data. Genetics 2018, 210, 767–777. [CrossRef]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.; Daly, M.J.; et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. American journal of human genetics 2007, 81, 559–575. [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria, 2023.
- Jombart, T.; Devillard, S.; Balloux, F. Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genetics 2010, 11, 94. [CrossRef]
- Danecek, P.; Auton, A.; Abecasis, G.; Albers, C.A.; Banks, E.; DePristo, M.A.; Handsaker, R.E.; Lunter, G.; Marth, G.T.; Sherry, S.T.; et al. The variant call format and VCFtools. Bioinformatics 2011, 27, 2156–2158. [CrossRef]
- Chen, Z.H.; Xu, Y.X.; Xie, X.L.; Wang, D.F.; Aguilar-Gómez, D.; Liu, G.J.; Li, X.; Esmailizadeh, A.; Rezaei, V.; Kantanen, J.; et al. Whole-genome sequence analysis unveils different origins of European and Asiatic mouflon and domestication-related genes in sheep. Communications Biology 2021, 4, 1307. [CrossRef]
- Tian, D.; Han, B.; Li, X.; Liu, D.; Zhou, B.; Zhao, C.; Zhang, N.; Wang, L.; Pei, Q.; Zhao, K. Genetic diversity and selection of Tibetan sheep breeds revealed by whole-genome resequencing. Animal bioscience 2023, 36, 991. [CrossRef]
- Yao, Y.; Pan, Z.; Di, R.; Liu, Q.; Hu, W.; Guo, X.; He, X.; Gan, S.; Wang, X.; Chu, M. Whole genome sequencing reveals the effects of recent artificial selection on litter size of Bamei mutton sheep. Animals 2021, 11, 157. [CrossRef]
- Fang, C.; Druet, T.; Cao, H.; Liu, W.; Chen, Q.; Farnir, F. Whole genome sequences of 297 Duolang sheep for litter size. Scientific Data 2025, 12, 1086. [CrossRef]
- Ma, R.; Kuang, R.; Zhang, J.; Sun, J.; Xu, Y.; Zhou, X.; Han, Z.; Hu, M.; Wang, D.; Fu, Y.; et al. Annotation and assessment of functional variants in livestock through epigenomic data. Journal of Genetics and Genomics 2025. [CrossRef]
- Wang, Z.H.; Zhu, Q.H.; Li, X.; Zhu, J.W.; Tian, D.M.; Zhang, S.S.; Kang, H.L.; Li, C.P.; Dong, L.L.; Zhao, W.M.; et al. iSheep: an integrated resource for sheep genome, variant and phenotype. Frontiers in Genetics 2021, 12, 714852. [CrossRef]
- Guo, Y.; Liang, J.; Lv, C.; Wang, Y.; Wu, G.; Ding, X.; Quan, G. Sequencing reveals population structure and selection signatures for reproductive traits in Yunnan semi-fine wool sheep (Ovis aries). Frontiers in Genetics 2022, 13, 812753. [CrossRef]
- Kijas, J.W.; Lenstra, J.A.; Hayes, B.; Boitard, S.; Porto Neto, L.R.; San Cristobal, M.; Servin, B.; McCulloch, R.; Whan, V.; Gietzen, K.; et al. Genome-wide analysis of the world’s sheep breeds reveals high levels of historic mixture and strong recent selection. PLoS biology 2012, 10, e1001258. [CrossRef]
- Al-Mamun, H.A.; Kwan, P.; Clark, S.A.; Ferdosi, M.H.; Tellam, R.; Gondro, C. Genome-wide association study of body weight in Australian Merino sheep reveals an orthologous region on OAR6 to human and bovine genomic regions affecting height and weight. Genetics Selection Evolution 2015, 47, 66. [CrossRef]
- Li, H.; Wu, X.L.; Tait Jr, R.; Bauck, S.; Thomas, D.; Murphy, T.; Rosa, G. Genome-wide association study of milk production traits in a crossbred dairy sheep population using three statistical models. Animal Genetics 2020, 51, 624–628. [CrossRef]
- Kaseja, K.; Mucha, S.; Yates, J.; Smith, E.; Banos, G.; Conington, J. Genome-wide association study of health and production traits in meat sheep. animal 2023, 17, 100968. [CrossRef]
- Lopez, B.I.; Santiago, K.G.; Lee, D.; Ha, S.; Seo, K. RNA sequencing (RNA-Seq) based transcriptome analysis in immune response of Holstein cattle to killed vaccine against bovine viral diarrhea virus type I. Animals 2020, 10, 344. [CrossRef]
- Chen, W.; Gu, X.; Lv, X.; Cao, X.; Yuan, Z.; Wang, S.; Sun, W. Non-coding transcriptomic profiles in the sheep mammary gland during different lactation periods. Frontiers in veterinary science 2022, 9, 983562. [CrossRef]
- Lins, J.G.G.; Amarante, A.F. Complement and Coagulation Cascade Activation Regulates the Early Inflammatory Mechanism of Resistance of Suckling Lambs Against Haemonchus contortus. Pathogens 2025, 14, 447. [CrossRef]
- Li, X.; Jiang, S.; Wang, X.; Jia, B. Intestinal transcriptomes in Kazakh sheep with different haplotypes after experimental Echinococcus granulosus infection. Parasite 2021, 28, 14. [CrossRef]
- Wang, M.; Bissonnette, N.; Laterrière, M.; Dudemaine, P.L.; Gagné, D.; Roy, J.P.; Sirard, M.A.; Ibeagha-Awemu, E.M. Gene co-expression in response to Staphylococcus aureus infection reveals networks of genes with specific functions during bovine subclinical mastitis. Journal of Dairy Science 2023, 106, 5517–5536. [CrossRef]
- Lv, X.; Chen, W.; Wang, S.; Cao, X.; Yuan, Z.; Getachew, T.; Mwacharo, J.M.; Haile, A.; Sun, W. Whole-genome resequencing of Dorper and Hu sheep to reveal selection signatures associated with important traits. Animal Biotechnology 2023, 34, 3016–3026. [CrossRef]
- Jin, M.; Wang, H.; Liu, G.; Lu, J.; Yuan, Z.; Li, T.; Liu, E.; Lu, Z.; Du, L.; Wei, C. Whole-genome resequencing of Chinese indigenous sheep provides insight into the genetic basis underlying climate adaptation. Genetics Selection Evolution 2024, 56, 26. [CrossRef]
- Ye, J.; Zhao, X.; Xue, H.; Zou, X.; Liu, G.; Deng, M.; Sun, B.; Guo, Y.; Liu, D.; Li, Y. RNA-Seq reveals miRNA and mRNA co-regulate muscle differentiation in fetal Leizhou goats. Frontiers in Veterinary Science 2022, 9, 829769. [CrossRef]
- How, S.S.; Nathan, S.; Lam, S.D.; Chieng, S. ATP-binding cassette (ABC) transporters: structures and roles in bacterial pathogenesis. Journal of Zhejiang University-Science B 2025, 26, 58–75. [CrossRef]
- Anderson, S.; Côté, S.; Richard, J.; Shafer, A. Genomic architecture of phenotypic extremes in a wild cervid. BMC genomics 2022, 23, 126. [CrossRef]
- Srivastava, D.B.; Ethayathulla, A.S.; Kumar, J.; Somvanshi, R.K.; Sharma, S.; Dey, S.; Singh, T.P. Carbohydrate binding properties and carbohydrate induced conformational switch in sheep secretory glycoprotein (SPS-40): Crystal structures of four complexes of SPS-40 with chitin-like oligosaccharides. Journal of structural biology 2007, 158, 255–266. [CrossRef]
- Boschiero, C.; Gao, Y.; Baldwin, R.L.; Ma, L.; Liu, G.E.; Li, C.J. Characterization of accessible chromatin regions in cattle rumen epithelial tissue during weaning. Genes 2022, 13, 535. [CrossRef]
- Schönemann, A.M.; Beiras, R.; Diz, A.P. Widespread alterations upon exposure to the estrogenic endocrine disruptor ethinyl estradiol in the liver proteome of the marine male fish Cyprinodon variegatus. Aquatic Toxicology 2022, 248, 106189. [CrossRef]
- Hou, X.; Wang, X.; Hou, S.; Dang, J.; Zhang, X.; Tang, J.; Shi, Y.; Ma, S.; Xu, Z. Comparative ultrastructural and transcriptomic profile analysis of skin tissues from indigenous, improved meat, and dairy goat breeds. BMC genomics 2024, 25, 1070. [CrossRef]
- Li, J.; Chen, C.; Zhao, R.; Wu, J.; Li, Z. Transcriptome analysis of mRNAs, lncRNAs, and miRNAs in the skeletal muscle of Tibetan chickens at different developmental stages. Frontiers in Physiology 2023, 14, 1225349. [CrossRef]
- Tsang, H.G.; Clark, E.L.; Markby, G.R.; Bush, S.J.; Hume, D.A.; Corcoran, B.M.; MacRae, V.E.; Summers, K.M. Expression of calcification and extracellular matrix genes in the cardiovascular system of the healthy domestic sheep (Ovis aries). Frontiers in Genetics 2020, 11, 919. [CrossRef]
- Wang, J.; Cui, K.; Hua, G.; Han, D.; Yang, Z.; Li, T.; Yang, X.; Zhang, Y.; Cai, G.; Deng, X.; et al. Skin-specific transgenic overexpression of ovine β-catenin in mice. Frontiers in Genetics 2023, 13, 1059913. [CrossRef]
- Ma, L.; Zhao, X.; Wang, H.; Chen, Z.; Zhang, K.; Xue, J.; Luo, Y.; Liu, H.; Jiang, X.; Wang, J.; et al. DNA Methylation Patterns and Transcriptomic Data Were Integrated to Investigate Candidate Genes Influencing Reproductive Traits in Ovarian Tissue from Sichuan White Geese. International Journal of Molecular Sciences 2025, 26, 3408. [CrossRef]
- Pereira, I.; Hidalgo, C.; Stoore, C.; Baquedano, M.S.; Cabezas, C.; Bastías, M.; Riveros, A.; Meneses, C.; Cancela, M.; Ferreira, H.B.; et al. Transcriptome analysis of Echinococcus granulosus sensu stricto protoscoleces reveals differences in immune modulation gene expression between cysts found in cattle and sheep. Veterinary Research 2022, 53, 8. [CrossRef]
- Varela-Martínez, E.; Abendaño, N.; Asín, J.; Sistiaga-Poveda, M.; Pérez, M.M.; Reina, R.; De Andres, D.; Luján, L.; Jugo, B.M. Molecular signature of aluminum hydroxide adjuvant in ovine PBMCs by integrated mRNA and microRNA transcriptome sequencing. Frontiers in immunology 2018, 9, 2406. [CrossRef]
- Park, M.; de Villavicencio Diaz, T.N.; Lange, V.; Wu, L.; Le Bihan, T.; Ma, B. Exploring the sheep (Ovis aries) immunoglobulin repertoire by next generation sequencing. Molecular Immunology 2023, 156, 20–30. [CrossRef]
- Kumar, P.; Becker, J.C.; Gao, K.; Carney, R.P.; Lankford, L.; Keller, B.A.; Herout, K.; Lam, K.S.; Farmer, D.L.; Wang, A. Neuroprotective effect of placenta-derived mesenchymal stromal cells: role of exosomes. The FASEB Journal 2019, 33, 5836. [CrossRef]
- Ping, X.; Chen, Y.; Wang, H.; Jin, Z.; Duan, Q.; Ren, Z.; Dong, X. Whole-genome sequencing reveals patterns of runs of homozygosity underlying genetic diversity and selection in domestic rabbits. BMC genomics 2025, 26, 425. [CrossRef]
- Guo, Z.; González, J.F.; Hernandez, J.N.; McNeilly, T.N.; Corripio-Miyar, Y.; Frew, D.; Morrison, T.; Yu, P.; Li, R.W. Possible mechanisms of host resistance to Haemonchus contortus infection in sheep breeds native to the Canary Islands. Scientific reports 2016, 6, 26200. [CrossRef]
- Yang, J.; Li, W.R.; Lv, F.H.; He, S.G.; Tian, S.L.; Peng, W.F.; Sun, Y.W.; Zhao, Y.X.; Tu, X.L.; Zhang, M.; et al. Whole-genome sequencing of native sheep provides insights into rapid adaptations to extreme environments. Molecular biology and evolution 2016, 33, 2576–2592. [CrossRef]
- Sun, L.; Yuan, C.; Guo, T.; Zhang, M.; Bai, Y.; Lu, Z.; Liu, J. Resequencing reveals population structure and genetic diversity in Tibetan sheep. BMC genomics 2024, 25, 906. [CrossRef]
- Mei, C.; Gui, L.; Hong, J.; Raza, S.H.A.; Aorigele, C.; Tian, W.; Garcia, M.; Xin, Y.; Yang, W.; Zhang, S.; et al. Insights into adaption and growth evolution: a comparative genomics study on two distinct cattle breeds from Northern and Southern China. Molecular Therapy Nucleic Acids 2021, 23, 959–967. [CrossRef]
- Zhu, S.; Guo, T.; Zhao, H.; Qiao, G.; Han, M.; Liu, J.; Yuan, C.; Wang, T.; Li, F.; Yue, Y.; et al. Genome-wide association study using individual single-nucleotide polymorphisms and haplotypes for erythrocyte traits in Alpine Merino sheep. Frontiers in genetics 2020, 11, 848. [CrossRef]
- Zhao, H.; Guo, T.; Lu, Z.; Liu, J.; Zhu, S.; Qiao, G.; Han, M.; Yuan, C.; Wang, T.; Li, F.; et al. Genome-wide association studies detects candidate genes for wool traits by re-sequencing in Chinese fine-wool sheep. BMC genomics 2021, 22, 127. [CrossRef]
- Al Kalaldeh, M.; Gibson, J.; Lee, S.H.; Gondro, C.; Van Der Werf, J.H. Detection of genomic regions underlying resistance to gastrointestinal parasites in Australian sheep. Genetics Selection Evolution 2019, 51, 37. [CrossRef]
- Yurchenko, A.A.; Deniskova, T.E.; Yudin, N.S.; Dotsev, A.V.; Khamiruev, T.N.; Selionova, M.I.; Egorov, S.V.; Reyer, H.; Wimmers, K.; Brem, G.; et al. High-density genotyping reveals signatures of selection related to acclimation and economically important traits in 15 local sheep breeds from Russia. BMC genomics 2019, 20, 294. [CrossRef]
- Cheng, J.; Cao, X.; Hanif, Q.; Pi, L.; Hu, L.; Huang, Y.; Lan, X.; Lei, C.; Chen, H. Integrating genome-wide CNVs into QTLs and high confidence GWAScore regions identified positional candidates for sheep economic traits. Frontiers in Genetics 2020, 11, 569. [CrossRef]
- Estrada-Reyes, Z.M.; Tsukahara, Y.; Amadeu, R.R.; Goetsch, A.L.; Gipson, T.A.; Sahlu, T.; Puchala, R.; Wang, Z.; Hart, S.P.; Mateescu, R.G. Signatures of selection for resistance to Haemonchus contortus in sheep and goats. BMC genomics 2019, 20, 735. [CrossRef]
- Rodrigues, J.L.; Braga, L.G.; Watanabe, R.N.; Schenkel, F.S.; Berry, D.P.; Buzanskas, M.E.; Munari, D.P. Genetic diversity and selection signatures in sheep breeds. Journal of Applied Genetics 2025, pp. 1–13. [CrossRef]
- Sallé, G.; Jacquiet, P.; Gruner, L.; Cortet, J.; Sauvé, C.; Prévot, F.; Grisez, C.; Bergeaud, J.P.; Schibler, L.; Tircazes, A.; et al. A genome scan for QTL affecting resistance to Haemonchus contortus in sheep. Journal of animal science 2012, 90, 4690–4705. [CrossRef]
- Horiuchi, S.; Wu, H.; Liu, W.C.; Schmitt, N.; Provot, J.; Liu, Y.; Bentebibel, S.E.; Albrecht, R.A.; Schotsaert, M.; Forst, C.V.; et al. Tox2 is required for the maintenance of GC TFH cells and the generation of memory TFH cells. Science advances 2021, 7, eabj1249. [CrossRef]
- Usai, M.G.; Casu, S.; Sechi, T.; Salaris, S.L.; Miari, S.; Mulas, G.; Cancedda, M.G.; Ligios, C.; Carta, A. Advances in understanding the genetic architecture of antibody response to paratuberculosis in sheep by heritability estimate and LDLA mapping analyses and investigation of candidate regions using sequence-based data. Genetics Selection Evolution 2024, 56, 5. [CrossRef]
- Thompson-Crispi, K.A.; Sargolzaei, M.; Ventura, R.; Abo-Ismail, M.; Miglior, F.; Schenkel, F.; Mallard, B.A. A genome-wide association study of immune response traits in Canadian Holstein cattle. BMC genomics 2014, 15, 559. [CrossRef]
- Hazard, D.; Moreno, C.; Foulquié, D.; Delval, E.; François, D.; Bouix, J.; Sallé, G.; Boissy, A. Identification of QTLs for behavioral reactivity to social separation and humans in sheep using the OvineSNP50 BeadChip. BMC genomics 2014, 15, 778. [CrossRef]
- Moreno García, C.A.; Zhou, H.; Altimira, D.; Dynes, R.; Gregorini, P.; Jayathunga, S.; Maxwell, T.M.; Hickford, J. The glutamate metabotropic receptor 5 (GRM5) gene is associated with beef cattle home range and movement tortuosity. Journal of Animal Science and Biotechnology 2022, 13, 111. [CrossRef]
- Moreno García, C.A.; Perelman, S.B.; Dynes, R.; Maxwell, T.M.; Zhou, H.; Hickford, J. Key Grazing Behaviours of Beef Cattle Identify Specific Genotypes of the Glutamate Metabotropic Receptor 5 Gene (GRM5). Behavior Genetics 2024, 54, 212–229. [CrossRef]
- Hou, Y.; Liu, G.E.; Bickhart, D.M.; Matukumalli, L.K.; Li, C.; Song, J.; Gasbarre, L.C.; Van Tassell, C.P.; Sonstegard, T.S. Genomic regions showing copy number variations associate with resistance or susceptibility to gastrointestinal nematodes in Angus cattle. Functional & integrative genomics 2012, 12, 81–92.
- Moss, D.K.; Bellett, G.; Carter, J.M.; Liovic, M.; Keynton, J.; Prescott, A.R.; Lane, E.B.; Mogensen, M.M. Ninein is released from the centrosome and moves bi-directionally along microtubules. Journal of cell science 2007, 120, 3064–3074. [CrossRef]
- Gholizadeh, M.; Rahimi-Mianji, G.; Nejati-Javaremi, A. Genomewide association study of body weight traits in Baluchi sheep. Journal of Genetics 2015, 94, 143–146. [CrossRef]
- Goyache, F.; Fernández, I.; Tapsoba, A.S.R.; Traoré, A.; Menéndez-Arias, N.A.; Álvarez, I. Functional characterization of Copy Number Variations regions in Djallonké sheep. Journal of Animal Breeding and Genetics 2021, 138, 600–612. [CrossRef]
- Rovadoscki, G.; Pertile, S.; Alvarenga, A.; Cesar, A.; Pértille, F.; Petrini, J.; Franzo, V.; Soares, W.; Morota, G.; Spangler, M.L.; et al. Estimates of genomic heritability and genome-wide association study for fatty acids profile in Santa Inês sheep. BMC genomics 2018, 19, 375. [CrossRef]
- Li, R.; Luo, R.; Luo, Y.; Hou, Y.; Wang, J.; Zhang, Q.; Chen, X.; Hu, L.; Zhou, J. Biological function, mediate cell death pathway and their potential regulated mechanisms for post-mortem muscle tenderization of PARP1: A review. Frontiers in Nutrition 2022, 9, 1093939. [CrossRef]
- Thorne, J.W.; Redden, R.; Bowdridge, S.A.; Becker, G.M.; Stegemiller, M.R.; Murdoch, B.M. Genome-wide analysis of sheep artificially or naturally infected with gastrointestinal nematodes. Genes 2023, 14, 1342. [CrossRef]
- Hai, J.; Zhu, C.Q.; Wang, T.; Organ, S.L.; Shepherd, F.A.; Tsao, M.S. TRIM14 is a putative tumor suppressor and regulator of innate immune response in non-small cell lung cancer. Scientific reports 2017, 7, 39692. [CrossRef]
- Haehling, M.B.; Cruvinel, G.G.; Toscano, J.H.; Giraldelo, L.A.; Santos, I.B.; Esteves, S.N.; Benavides, M.V.; Júnior, W.B.; Niciura, S.C.; Chagas, A.C.S. Four single nucleotide polymorphisms (SNPs) are associated with resistance and resilience to Haemonchus contortus in Brazilian Morada Nova sheep. Veterinary Parasitology 2020, 279, 109053. [CrossRef]
- Poulsen, N.A.; Robinson, R.C.; Barile, D.; Larsen, L.B.; Buitenhuis, B. A genome-wide association study reveals specific transferases as candidate loci for bovine milk oligosaccharides synthesis. BMC genomics 2019, 20, 404. [CrossRef]
- Guo, T.; Li, B.; Kang, Y.; Gu, C.; Fang, F.; Chen, X.; Liu, X.; Lu, G.; Feng, C.; Xu, C. COLGALT2 is overexpressed in ovarian cancer and interacts with PLOD3. Clinical and Translational Medicine 2021, 11, e370. [CrossRef]
- Miles, A.M.; Posbergh, C.J.; Huson, H.J. Direct phenotyping and principal component analysis of type traits implicate novel QTL in bovine mastitis through genome-wide association. Animals 2021, 11, 1147. [CrossRef]
- Atlija, M.; Arranz, J.J.; Martinez-Valladares, M.; Gutiérrez-Gil, B. Detection and replication of QTL underlying resistance to gastrointestinal nematodes in adult sheep using the ovine 50K SNP array. Genetics Selection Evolution 2016, 48, 4. [CrossRef]
- Yang, R.; Han, Z.; Zhou, W.; Li, X.; Zhang, X.; Zhu, L.; Wang, J.; Li, X.; Zhang, C.l.; Han, Y.; et al. Population structure and selective signature of Kirghiz sheep by Illumina Ovine SNP50 BeadChip. PeerJ 2024, 12, e17980. [CrossRef]
- Wang, Z.; Zhang, H.; Yang, H.; Wang, S.; Rong, E.; Pei, W.; Li, H.; Wang, N. Genome-wide association study for wool production traits in a Chinese Merino sheep population. PloS one 2014, 9, e107101. [CrossRef]
- Moaeen-ud Din, M.; Danish Muner, R.; Khan, M.S. Genome wide association study identifies novel candidate genes for growth and body conformation traits in goats. Scientific Reports 2022, 12, 9891. [CrossRef]
- Martinez-Sanchez, N.; Brümmer, A.; Barron, N.J.; Rosoff, D.B.; Liechti, A.; Voronkov, M.; Hayter, E.A.; Chamois, S.; Dreos, R.; Guex, N.; et al. The 18S rRNA methyltransferase, BUD23, is required for appropriate lipid and mitochondrial metabolism. bioRxiv 2025, pp. 2025–05.
- Baxter, M.; Voronkov, M.; Poolman, T.; Galli, G.; Pinali, C.; Goosey, L.; Knight, A.; Krakowiak, K.; Maidstone, R.; Iqbal, M.; et al. Cardiac mitochondrial function depends on BUD23 mediated ribosome programming. Elife 2020, 9, e50705. [CrossRef]
- Gmel, A.I.; Burger, D.; Neuditschko, M. A novel QTL and a candidate gene are associated with the progressive motility of Franches-Montagnes stallion spermatozoa after thaw. Genes 2021, 12, 1501. [CrossRef]
- Marques, D.B.; Bastiaansen, J.W.; Broekhuijse, M.L.; Lopes, M.S.; Knol, E.F.; Harlizius, B.; Guimarães, S.E.; Silva, F.F.; Lopes, P.S. Weighted single-step GWAS and gene network analysis reveal new candidate genes for semen traits in pigs. Genetics Selection Evolution 2018, 50, 40. [CrossRef]
- Pasandideh, M.; Gholizadeh, M.; Rahimi-Mianji, G. A genome-wide association study revealed five SNPs affecting 8-month weight in sheep. Animal genetics 2020, 51, 973–976. [CrossRef]
- Almasi, M.; Zamani, P.; Mirhoseini, S.Z.; Moradi, M.H. Genome-wide association study for postweaning weight traits in Lori-Bakhtiari sheep. Tropical Animal Health and Production 2021, 53, 163. [CrossRef]
- Maddahi, N.; Sadeghi, M.; Miraee Ashtiani, S.R.; Kholghi, M.; Jalil Sarghale, A. Genome-wide association studies and candidate genes networks affecting reproductive traits using Iranian Holstein sequence data. BMC genomics 2025, 26, 656. [CrossRef]
- Ladeira, G.C.; Pilonetto, F.; Fernandes, A.C.; Bóscollo, P.P.; Dauria, B.D.; Titto, C.G.; Coutinho, L.L.; e Silva, F.F.; Pinto, L.F.B.; Mourão, G.B. CNV detection and their association with growth, efficiency and carcass traits in Santa Inês sheep. Journal of Animal Breeding and Genetics 2022, 139, 476–487. [CrossRef]
- Huang, Y.Z.; Qian, L.N.; Wang, J.; Zhang, C.L.; Fang, X.T.; Lei, C.Z.; Lan, X.Y.; Ma, Y.; Bai, Y.Y.; Lin, F.P.; et al. Genetic variants in ADD1 gene and their associations with growth traits in cattle. Animal Biotechnology 2019, 30, 7–12. [CrossRef]
- Li, C.; Pan, Y.; Me, H. Polymorphism of the H-FABP, MC4R and ADD1 genes in the Meishan and four other pig populations in China. South African Journal of Animal Science 2006, 36, 1–6. [CrossRef]
- Gholizadeh, M.; Rahimi-Mianji, G.; Nejati-Javaremi, A.; De Koning, D.J.; Jonas, E. Genomewide association study to detect QTL for twinning rate in Baluchi sheep. Journal of genetics 2014, 93, 489–493. [CrossRef]
- Ocbina, P.J.R.; Tuson, M.; Anderson, K.V. Primary cilia are not required for normal canonical Wnt signaling in the mouse embryo. PloS one 2009, 4, e6839. [CrossRef]
- Ocbina, P.J.R.; Eggenschwiler, J.T.; Moskowitz, I.; Anderson, K.V. Complex interactions between genes controlling trafficking in primary cilia. Nature genetics 2011, 43, 547–553. [CrossRef]
- Salehian-Dehkordi, H.; Xu, Y.X.; Xu, S.S.; Li, X.; Luo, L.Y.; Liu, Y.J.; Wang, D.F.; Cao, Y.H.; Shen, M.; Gao, L.; et al. Genome-wide detection of copy number variations and their association with distinct phenotypes in the world’s sheep. Frontiers in genetics 2021, 12, 670582. [CrossRef]
- Park, S.; Shimada, K.; Fujihara, Y.; Xu, Z.; Shimada, K.; Larasati, T.; Pratiwi, P.; Matzuk, R.M.; Devlin, D.J.; Yu, Z.; et al. CRISPR/Cas9-mediated genome-edited mice reveal 10 testis-enriched genes are dispensable for male fecundity. Biology of Reproduction 2020, 103, 195–204. [CrossRef]
- Chan, C.C.; Yen, T.H.; Tseng, H.C.; Mai, B.; Ho, P.K.; Chou, J.L.; Wu, G.J.; Huang, Y.C. A comprehensive genetic study of microtubule-associated gene clusters for male infertility in a Taiwanese cohort. International Journal of Molecular Sciences 2023, 24, 15363. [CrossRef]
- Wu, C.; Wang, C.; Zhai, B.; Zhao, Y.; Zhao, Z.; Yuan, Z.; Fu, X.; Zhang, M. Study on the region-specific expression of epididymis mRNA in the rams. PLoS One 2021, 16, e0245933. [CrossRef]
- Thimon, V.; Koukoui, O.; Calvo, E.; Sullivan, R. Region-specific gene expression profiling along the human epididymis. Molecular human reproduction 2007, 13, 691–704. [CrossRef]
- Matika, O.; Riggio, V.; Anselme-Moizan, M.; Law, A.S.; Pong-Wong, R.; Archibald, A.L.; Bishop, S.C. Genome-wide association reveals QTL for growth, bone and in vivo carcass traits as assessed by computed tomography in Scottish Blackface lambs. Genetics Selection Evolution 2016, 48, 11. [CrossRef]
- Zhang, L.; Zhang, T.; Yang, Z.; Cai, C.; Hao, S.; Yang, L. Expression of nuclear factor kappa B in ovine maternal inguinal lymph nodes during early pregnancy. BMC Veterinary Research 2022, 18, 266. [CrossRef]
- Kravitz, A.; Liao, M.; Morota, G.; Tyler, R.; Cockrum, R.; Manohar, B.M.; Ronald, B.S.M.; Collins, M.T.; Sriranganathan, N. Retrospective Single Nucleotide Polymorphism Analysis of Host Resistance and Susceptibility to Ovine Johne’s Disease Using Restored FFPE DNA. International Journal of Molecular Sciences 2024, 25, 7748. [CrossRef]
- Kasimanickam, R.; Ferreira, J.C.P.; Kastelic, J.; Kasimanickam, V. Application of genomic selection in beef cattle disease prevention. Animals 2025, 15, 277. [CrossRef]
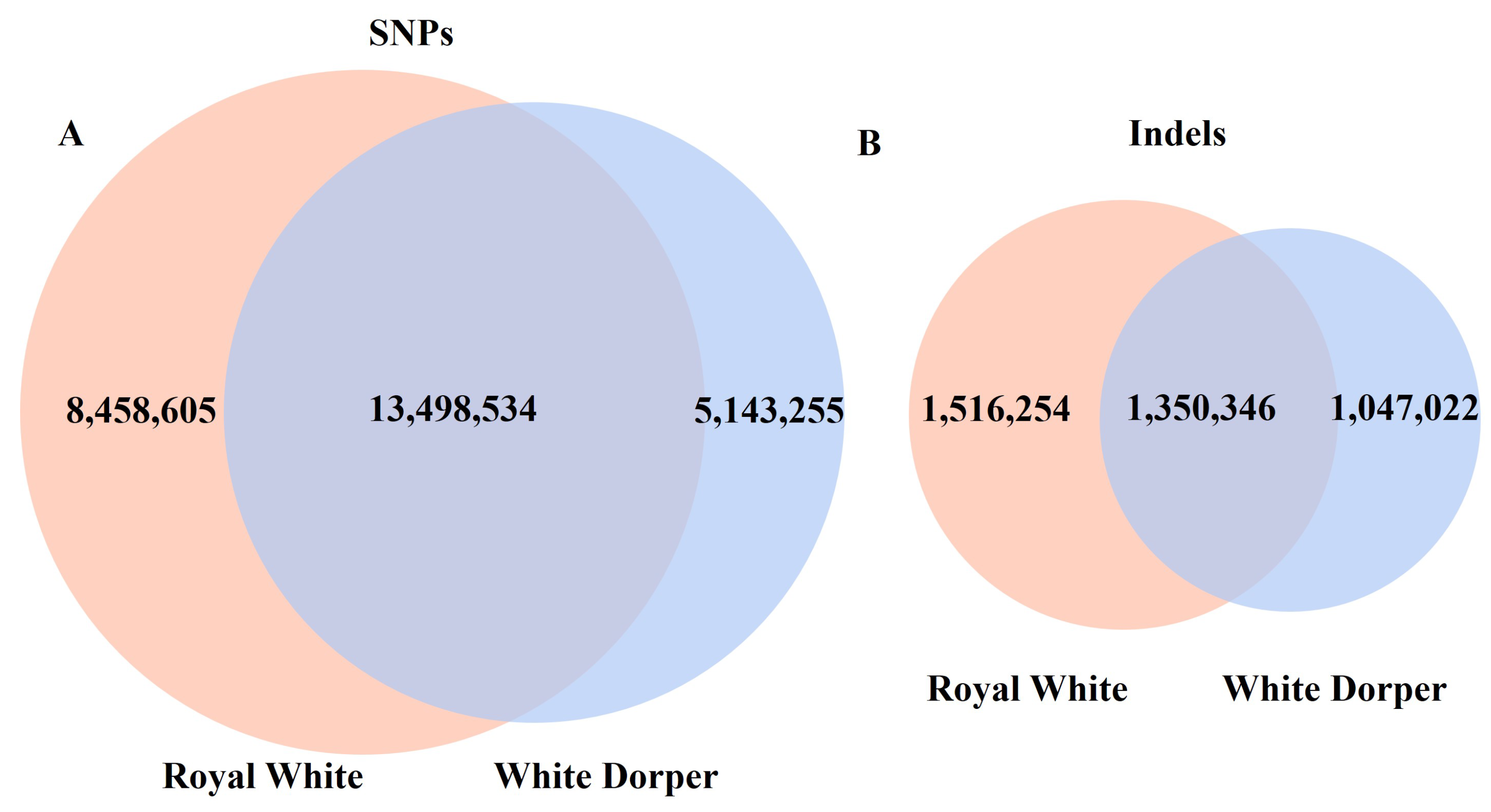
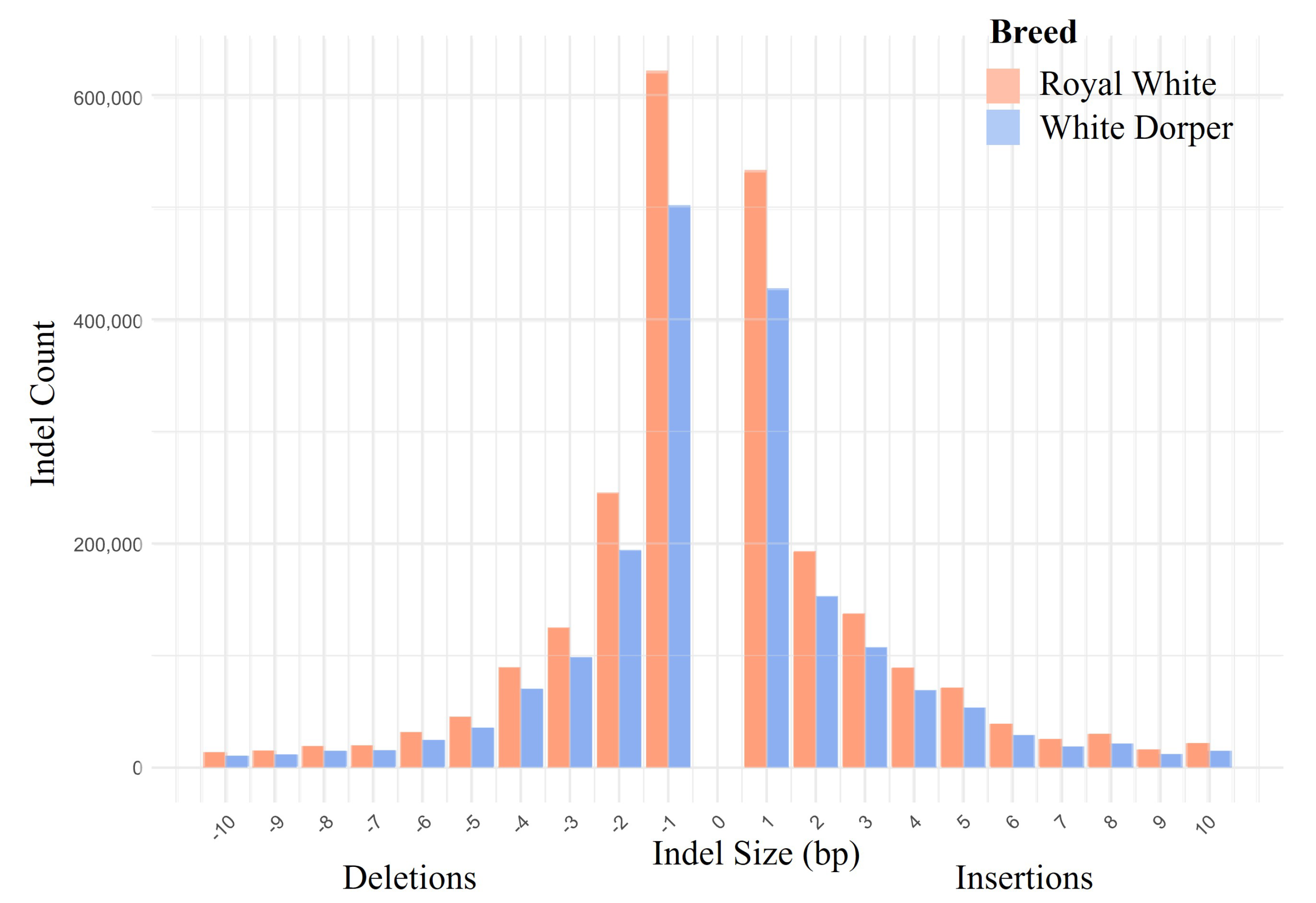
| Breed | Sample | Raw Reads | Cleaned Reads | Cleaned Reads Retained | Mapped Read Rate | Average Depth |
|---|---|---|---|---|---|---|
| Royal White | RW1 | 66,685,965 | 63,899,137 | 95.82% | 99.93% | 6.53× |
| RW2 | 68,121,792 | 65,611,571 | 96.32% | 99.93% | 6.66× | |
| RW3 | 90,611,600 | 87,965,250 | 97.08% | 99.95% | 8.73× | |
| RW4 | 75,592,210 | 71,813,373 | 95.00% | 99.93% | 7.19× | |
| RW5 | 71,073,803 | 68,690,719 | 96.65% | 99.94% | 7.09× | |
| RW6 | 57,832,096 | 53,349,935 | 92.25% | 99.92% | 5.34× | |
| RW7 | 75,326,235 | 70,831,741 | 94.03% | 99.95% | 6.89× | |
| RW8 | 111,436,645 | 105,628,786 | 94.79% | 99.93% | 10.00× | |
| RW9 | 73,792,997 | 68,491,200 | 92.82% | 99.93% | 6.34× | |
| RW10 | 69,186,458 | 65,441,733 | 94.59% | 99.93% | 6.44× | |
| RW11 | 83,568,451 | 79,745,350 | 95.43% | 99.92% | 7.89× | |
| White Dorper | WD1 | 81,859,706 | 76,544,352 | 93.51% | 99.94% | 7.21× |
| WD2 | 75,404,617 | 69,620,505 | 92.32% | 99.94% | 6.52× | |
| WD3 | 65,262,440 | 60,110,417 | 92.11% | 99.95% | 5.66× | |
| WD4 | 63,321,027 | 54,063,159 | 85.38% | 99.91% | 5.19× | |
| WD5 | 50,539,051 | 46,329,622 | 91.67% | 99.90% | 5.01× | |
| WD6 | 71,909,006 | 68,680,645 | 95.51% | 99.94% | 6.86× | |
| WD7 | 78,854,393 | 75,289,750 | 95.48% | 99.93% | 7.41× | |
| WD8 | 62,284,501 | 59,082,446 | 94.86% | 99.92% | 5.99× | |
| WD9 | 75,691,236 | 72,451,313 | 95.72% | 99.93% | 7.11× |
| Metric | Royal White | White Dorper |
|---|---|---|
| SNP | 21,957,139 | 18,641,789 |
| Ts/Tv Ratio | 2.30 | 2.16 |
| Het/Hom (SNP) | 0.999 | 0.998 |
| Known SNP (%) | 16,958,892 (77.24%) | 14,460,461 (77.57%) |
| Novel SNP (%) | 4,998,247 (22.76%) | 4,181,328 (22.43%) |
| Indels | 2,866,600 | 2,397,368 |
| Het/Hom (Indels) | 0.998 | 0.992 |
| Known Indels (%) | 2,240,722 (78.17%) | 1,873,106 (78.13%) |
| Novel Indels (%) | 625,878 (21.83%) | 524,262 (21.87%) |
| Variant Type | Royal White SNP | White Dorper SNP |
|---|---|---|
| 3’ UTR variant | 335,924 | 286,072 |
| 5’ UTR premature start codon gain | 24,360 | 20,620 |
| 5’ UTR variant | 144,911 | 122,548 |
| Downstream gene variant | 2,330,242 | 1,975,745 |
| Initiator codon variant | 47 | 44 |
| Intergenic region | 11,967,899 | 10,138,113 |
| Intragenic variant | 6,681 | 6,640 |
| Intron variant | 27,465,429 | 23,392,133 |
| Missense variant | 142,906 | 124,264 |
| Non-coding transcript exon variant | 166,581 | 143,434 |
| Splice acceptor variant | 970 | 897 |
| Splice donor variant | 974 | 981 |
| Splice region variant | 42,203 | 36,165 |
| Start lost | 418 | 295 |
| Start retained variant | 1 | 0 |
| Stop gained | 3,525 | 3,886 |
| Stop lost | 304 | 237 |
| Stop retained variant | 185 | 149 |
| Synonymous variant | 195,280 | 165,971 |
| Upstream gene variant | 2,308,159 | 1,951,646 |
| Genes1 | Chr | FST2 | SNP IDs3 | QTL Traits4 | Category |
|---|---|---|---|---|---|
| GRM5 | 21 | 0.17 | rs424837012 | Vocalization during arena test | Behavior |
| MAGI2 | 4 | 0.18 | rs429561404 | Locomotion during arena test | Behavior |
| GRM5 | 21 | 0.39 | rs424244818 | Locomotion during isolation box test | Behavior |
| JADE2 | 5 | 0.19 | rs413619557 | Body weight (body weight at 6 months) | Growth |
| ALDH5A1 | 20 | 0.11 | rs421181203 | Mycobacterium avium subsp. paratuberculosis susceptibility (infection status and antibody titer) | Health |
| TGFB2 | 12 | 0.11 | rs160759291 | Gastrointestinal nematode resistance (Haemonchus contortus) | Health |
| TGFB2 | 12 | 0.11 | rs162057314 | Gastrointestinal nematode resistance (Haemonchus contortus) | Health |
| TOX2 | 13 | 0.13 | rs423531735 | Fecal egg count (Haemonchus contortus FEC2) | Health |
| HERC6 | 6 | 0.16 | rs424266480 | Fecal egg count | Health |
| NRXN1 | 3 | 0.24 | rs409057468 | Red blood cell distribution width | Health |
| PARP8 | 16 | 0.17 | rs416975775 | Meat omega-6 to omega-3 fatty acid ratio | Meat |
| NIN | 7 | 0.12 | rs410734119 | Milk yield | Milk |
| NRXN1 | 3 | 0.30 | rs429232758 | Fiber diameter coefficient of variance | Wool |
| Genes1 | Chr | FST2 | SNP IDs3 | QTL Traits4 | Category |
|---|---|---|---|---|---|
| PLXDC2 | 13 | 0.14 | rs401963094 | Body weight (body weight at 9 months) | Growth |
| COLGALT2 | 12 | 0.14 | rs402132699 | Average daily gain (daily weight gain after nematode challenge) | Growth |
| HYDIN | 14 | 0.29 | rs410323459 | Body weight (body weight at 8 months) | Growth |
| LAMC1 | 12 | 0.13 | rs596561468 | Gastrointestinal nematode resistance (Haemonchus contortus resistance) | Health |
| COLGALT2 | 12 | 0.14 | rs402132699 | Fecal egg count | Health |
| COLGALT2 | 12 | 0.14 | rs402132699 | Fecal egg count (fecal egg count after nematode challenge) | Health |
| COLGALT2 | 12 | 0.14 | rs402132699 | Hematocrit (packed cell volume after nematode challenge) | Health |
| TRIM14 | 2 | 0.18 | rs422296454 | Change in hematocrit (packed cell volume change) | Health |
| EPHA5 | 6 | 0.27 | rs426828157 | Fecal egg count | Health |
| ADD2 | 3 | 0.14 | rs417859328 | Dressing percentage | Meat |
| TENM2 | 5 | 0.11 | rs409487914 | Milk fat yield (180-day milk fat yield) | Milk |
| BUD23 | 24 | 0.18 | rs430795622 | Milk fat yield (180-day milk fat yield) | Milk |
| SCN8A | 3 | 0.40 | rs419496265 | Milk fat percentage | Milk |
| LCN8 | 3 | 0.12 | rs415039972 | Horn number | Morphology |
| NFKB1 | 6 | 0.17 | rs404225841 | Bone area | Morphology |
| STPG3 | 3 | 0.16 | rs430682724 | Offspring number (litter size) | Reproduction |
| DYNC2H1 | 15 | 0.24 | rs413723884 | Offspring number (total number of lambs across first four parities) | Reproduction |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




